AI-Powered Sperm Morphology Analysis: A Technical Guide for Biomedical Research and Development
This article provides a comprehensive technical review of artificial intelligence applications in automated sperm morphology analysis, a critical yet subjective component of male fertility assessment.
AI-Powered Sperm Morphology Analysis: A Technical Guide for Biomedical Research and Development
Abstract
This article provides a comprehensive technical review of artificial intelligence applications in automated sperm morphology analysis, a critical yet subjective component of male fertility assessment. Tailored for researchers, scientists, and drug development professionals, we detail the foundational challenges in conventional analysis that drive AI adoption, explore the technical architecture of deep learning models—from convolutional neural networks (CNNs) to transformers—and their implementation for segmenting and classifying sperm components. The scope extends to troubleshooting key development hurdles, including dataset limitations and model generalizability, and concludes with a rigorous validation and comparative analysis of AI systems against established diagnostic methods and their emerging role in predicting functional outcomes like DNA fragmentation.
The Diagnostic Imperative: Why AI is Revolutionizing Sperm Morphology Analysis
The Global Burden of Male Infertility and the Central Role of Sperm Morphology
Male infertility constitutes a significant and growing global public health challenge, with male factors contributing to approximately 50% of all infertility cases among couples [1] [2]. The comprehensive assessment of male fertility potential has traditionally relied on semen analysis, among which sperm morphology—evaluating the size, shape, and structural integrity of spermatozoa—represents a crucial diagnostic parameter [3] [4]. Historically, semen analysis has been hampered by subjectivity and variability, but the emergence of artificial intelligence (AI) is revolutionizing this field through automated, objective, and high-throughput evaluations [5] [6]. This whitepaper examines the global burden of male infertility through recent epidemiological data, details the central role of sperm morphology in clinical assessment, and explores how AI technologies are transforming diagnostic protocols and research methodologies for scientists and drug development professionals.
The Global Burden of Male Infertility: Epidemiological Landscape
Recent data from the Global Burden of Disease (GBD) studies reveals a substantial and increasing burden of male infertility worldwide, with notable disparities across geographical regions and socio-economic groupings [7] [8] [9].
Global Prevalence and Temporal Trends
The GBD 2019 study provides comprehensive estimates of male infertility prevalence and associated disability. The data demonstrates a alarming increase in the global burden over nearly three decades.
Table 1: Global Burden of Male Infertility (1990-2019)
| Metric | 1990 Estimate | 2019 Estimate | Absolute Change | Percentage Change |
|---|---|---|---|---|
| Global Prevalence | 31,941.9 thousand | 56,530.4 thousand | +24,588.5 thousand | +76.9% |
| Age-Standardized Prevalence Rate (ASPR) | 1,179.07 per 100,000 | 1,402.98 per 100,000 | +223.91 per 100,000 | +19.0% |
| Age-Standardized YLD Rate (ASYR) | Not specified | Equivalent to DALY rate | Not applicable | Not applicable |
Analysis of more recent data from GBD 2021 indicates these trends are continuing, showing a 74.66% global increase in the number of cases of male infertility among reproductive-aged men (15-49 years) between 1990 and 2021 [9]. This period also saw a 74.64% increase in Disability-Adjusted Life Years (DALYs), highlighting the significant health impact of this condition [9].
Regional and Socio-Demographic Variations
The burden of male infertility is not uniformly distributed globally. Specific regions and socio-demographic index (SDI) groupings bear a disproportionately high burden.
Table 2: Regional and SDI-Based Distribution of Male Infertility Burden (2019)
| Region/SDI Group | ASPR/ASYR Status | Noteworthy Observations |
|---|---|---|
| Western Sub-Saharan Africa | Highest ASPR/ASYR | Among the regions with the greatest burden |
| Eastern Europe | Highest ASPR/ASYR | Among the regions with the greatest burden |
| East Asia | Highest ASPR/ASYR | Among the regions with the greatest burden |
| High-middle SDI | Exceeds global average | Burden far exceeds the global average |
| Middle SDI | Exceeds global average | Burden far exceeds the global average; recorded highest number of cases in 2021 [9] |
| Low & Middle-low SDI | Notable upward trend since 2010 | Indicates a shifting burden |
Furthermore, the burden of male infertility demonstrates a negative correlation with national SDI levels, meaning countries with lower socio-demographic development often experience a greater relative burden [9]. From an age distribution perspective, the peak prevalence and years lived with disability (YLDs) occur in the 30-34 year age group globally, with the 35-39 year age group also reporting the highest number of cases in 2021, underscoring the impact on men in their prime reproductive years [7] [9].
Sperm Morphology: Fundamentals and Clinical Assessment
Defining Sperm Morphology and Its Clinical Relevance
Sperm morphology refers to the size, shape, and structural integrity of spermatozoa, evaluated based on strict criteria established by the World Health Organization (WHO) [3] [2]. A normal spermatozoon features a smooth, oval head (approximately 5-6 micrometers long and 2.5-3.5 micrometers wide), an intact acrosome covering 40-70% of the head, a well-defined midpiece, and a single, unbranched tail that is approximately 45 micrometers long [3] [4]. Cytoplasmic droplets should not exceed one-third of the sperm head size [1].
The clinical value of sperm morphology as a standalone prognostic factor is debated. While it is integrated into a broader diagnostic picture, its predictive power for natural conception or assisted reproductive technology (ART) outcomes is limited [3] [10]. The reference value for "normal" morphology has been progressively tightened in successive WHO manuals, from ≥80.5% in the first edition to a current threshold of ≥4% [4]. It is common for even fertile men to have a high percentage (90-96%) of abnormally shaped sperm in their ejaculate [3]. The French BLEFCO Group's 2025 guidelines explicitly recommend against using the percentage of normal-form sperm as a prognostic criterion before IUI, IVF, or ICSI, or for selecting the ART procedure [10].
Methodologies for Morphology Assessment
Assessment methods range from basic light microscopy to advanced electron microscopy, each with distinct applications and limitations.
- Conventional Semen Analysis (CSA): This is the traditional method involving the visual examination of stained sperm smears under a light microscope at 100x magnification. It is the clinical standard but is prone to significant inter-laboratory and inter-technician subjectivity [1] [2].
- Computer-Aided Sperm Analysis (CASA): These systems use digital imaging and basic algorithms to provide a more objective assessment of sperm concentration, motility, and, in advanced versions, morphology. They reduce but do not eliminate subjectivity, as they often rely on pre-set thresholds and staining [6].
- Transmission Electron Microscopy (TEM): TEM allows for ultrastructural analysis at high magnifications, enabling the detailed evaluation of internal sperm organelles. It is crucial for identifying and classifying specific defects, such as systematic sperm defects (e.g., globozoospermia, macrozoospermia, dysplasia of the fibrous sheath), and for research applications [4]. TEM analysis has facilitated the development of a "Fertility Index" to quantify sperm without ultrastructural defects [4].
The AI Revolution in Sperm Morphology Analysis
Artificial intelligence is addressing the critical limitations of traditional morphology assessment by introducing automation, objectivity, and the ability to discern subtle, predictive patterns.
From Conventional Machine Learning to Deep Learning
The evolution of AI in this field has progressed through distinct phases:
- Conventional Machine Learning (ML): Early approaches relied on handcrafted feature extraction. Algorithms like Support Vector Machines (SVM), K-means clustering, and decision trees were used to classify sperm based on manually engineered features such as shape descriptors (Hu moments, Zernike moments), texture, and grayscale intensity [2]. While a step forward, these models were limited by their dependence on human expertise for feature selection and often showed poor generalizability across different datasets [2].
- Deep Learning (DL): Deep learning, a subset of AI using multi-layered neural networks, has marked a significant breakthrough. DL models, particularly Convolutional Neural Networks (CNNs), can automatically learn hierarchical features directly from raw pixel data without manual intervention [5] [2]. This allows them to capture complex and subcellular morphological patterns that are imperceptible to the human eye [1]. Models like ResNet50 have been successfully adapted for sperm classification, demonstrating superior accuracy and robustness compared to conventional methods [1].
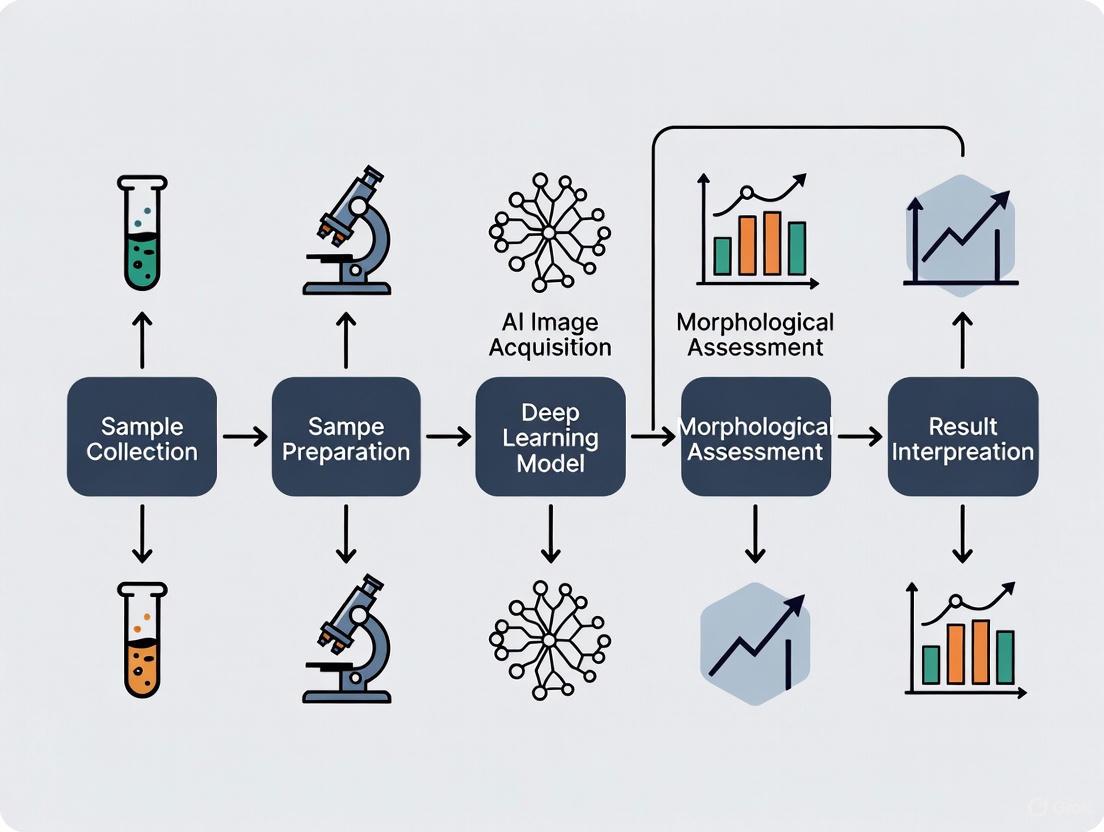
Experimental Protocol for AI-Based Morphology Assessment
A landmark 2025 study detailed an experimental protocol for an AI model that assesses unstained, live sperm morphology using confocal laser scanning microscopy [1]. This protocol is a template for robust AI development in this domain.
1. Sample Collection and Preparation:
- Participants: 30 healthy male volunteers (aged 18-40) were enrolled with 2-7 days of sexual abstinence [1].
- Sample Processing: Semen samples were collected via masturbation and allowed to liquefy. Each sample was divided into three aliquots for parallel analysis by the in-house AI model, CASA, and CSA [1].
2. Image Acquisition and Dataset Creation:
- Microscopy: A 6 μL droplet of sample was placed on a chamber slide. Images were captured using a confocal laser scanning microscope (LSM 800) at 40x magnification in Z-stack mode (interval: 0.5 μm, total range: 2 μm) [1].
- Annotation: Embryologists and researchers manually annotated well-focused sperm images using the LabelImg program, categorizing sperm into nine datasets based on strict WHO criteria for normal and abnormal features (head, neck, tail) [1]. The inter-observer correlation coefficient for normal sperm detection was 0.95 [1].
- Dataset: The final dataset contained 21,600 images, with 12,683 annotated sperm images. The model was trained on a subset of 9,000 images (4,500 normal, 4,500 abnormal) [1].
3. AI Model Development and Training:
- Model Architecture: The ResNet50, a pre-trained deep learning model, was selected and fine-tuned using a transfer learning approach [1].
- Training: The model was trained to minimize the difference between its predictions and the manual annotations. It achieved a test accuracy of 93% after 150 epochs, with a precision of 0.95 and recall of 0.91 for abnormal sperm, and 0.91 precision and 0.95 recall for normal sperm [1].
- Performance: The AI model's performance was evaluated against CASA and CSA. It showed a strong correlation with CASA (r=0.88) and CSA (r=0.76). The correlation between CASA and CSA was weaker (r=0.57) [1].
4. Key Advantage: A critical outcome of this study was the model's ability to accurately analyze unstained, live sperm. This is a significant advancement because it allows for the selection of viable sperm for use in Assisted Reproductive Technology (ART) immediately after assessment, preserving sperm viability [1].
The Scientist's Toolkit: Essential Research Reagents and Materials
For researchers aiming to replicate or build upon these advanced experiments, the following tools and reagents are essential.
Table 3: Research Reagent Solutions for AI-Based Sperm Morphology Analysis
| Item | Function/Application | Example from Literature |
|---|---|---|
| Confocal Laser Scanning Microscope | High-resolution, Z-stack imaging of live, unstained sperm, enabling 3D structural analysis. | LSM 800 [1] |
| Standardized Chamber Slides | Provides a consistent depth for sample preparation, ensuring uniform imaging conditions. | Leja standard two-chamber slide (20 μm depth) [1] |
| Annotation Software | Allows for precise manual labeling of sperm images to create ground-truth datasets for AI training. | LabelImg program [1] |
| Deep Learning Framework | Provides the programming environment for building, training, and validating neural network models. | ResNet50 model [1] |
| High-Performance Computing Unit | Processes large image datasets and performs the computationally intensive training of deep learning models. | GPU-accelerated computing [6] |
| Public & Proprietary Datasets | Serves as a benchmark for training and validating new models. | HSMA-DS, MHSMA, SVIA datasets [2] |
Visualization of Sperm Morphology Assessment Criteria
The following diagram synthesizes WHO criteria and AI classification logic for evaluating normal versus abnormal sperm morphology, providing a clear decision framework.
The global burden of male infertility is substantial and rising, necessitating advanced and standardized diagnostic approaches. Sperm morphology remains a central, though complex, component of male fertility assessment. The integration of artificial intelligence, particularly deep learning, is poised to fundamentally reshape this field. AI technologies offer a path toward fully automated, highly objective, and more prognostically powerful sperm morphology analysis. For researchers and drug developers, mastering these AI-driven tools and methodologies is critical for advancing both our understanding of male infertility and the development of novel therapeutic interventions. Future work must focus on creating larger, standardized datasets, improving model interpretability, and conducting rigorous clinical trials to validate AI systems' impact on live birth outcomes.
Sperm morphology analysis, the process of evaluating the shape and size of sperm cells, is a cornerstone of male fertility assessment. It provides critical insights into reproductive potential, as normal sperm morphology is associated with intact DNA and favorable clinical outcomes in assisted reproductive technology (ART) [1]. According to the World Health Organization (WHO) guidelines, this analysis requires the classification of at least 200 spermatozoa into categories such as normal, head defects, neck/midpiece defects, tail defects, and excess residual cytoplasm [11]. This evaluation offers diagnosticians valuable information on male testicular and epididymal function, helping to predict natural pregnancy outcomes and inform treatment strategies [2].
Despite its clinical importance, the conventional methodology for sperm morphology assessment has remained largely unchanged for decades, relying on trained technicians to visually evaluate sperm cells under a microscope after staining. This manual process is characterized by fundamental limitations that compromise its diagnostic reliability. As stated in a 2025 expert review from the French BLEFCO Group, "There is a huge variability in the performance and interpretation of this test," challenging its clinical value in infertility workups [10]. This technical guide examines the core limitations of conventional manual analysis—subjectivity, variability, and excessive workload—framed within the context of how artificial intelligence (AI) research is pioneering solutions to these long-standing challenges.
Core Limitations of Conventional Manual Analysis
The traditional approach to sperm morphology assessment faces three interconnected critical limitations that affect its analytical reliability and clinical utility.
Subjectivity in Morphological Interpretation
The classification of sperm as "normal" or "abnormal" relies heavily on the visual interpretation of complex, often subtle, morphological criteria by human observers. This introduces significant subjectivity into the diagnostic process.
- Complex Classification Standards: According to WHO standards, sperm morphology is divided into head, neck, and tail components, with 26 distinct types of abnormal morphology recognized [2]. Technicians must mentally process these complex criteria while evaluating each cell, leading to cognitive overload and inconsistent application of standards.
- Lack of Detailed Analysis Consensus: The French BLEFCO Group's 2025 guidelines explicitly recommend against "systematic detailed analysis of abnormalities (or groups of abnormalities) during sperm morphology assessment" due to the inherent subjectivity involved [10]. This recommendation from a leading expert group underscores the recognition that detailed manual classification is unreliable.
- Challenging Visual Assessments: Manual analysis struggles with accurately distinguishing subtle features such as vacuoles in the sperm head or slight irregularities in the acrosome, neck, and tail structures [2]. These visual challenges are compounded when sperm appear intertwined in images or when only partial structures are visible at the edges of the microscopic field [2].
High Inter-Observer and Intra-Observer Variability
The subjective nature of manual morphology assessment directly translates to substantial variability in results, both between different technicians and when the same technician repeats the analysis.
- Reproducibility Challenges: A 2025 review highlighted that morphological evaluation of sperm "faces considerable limitations in reproducibility and objectivity" [2]. This lack of reproducibility undermines the test's reliability for tracking changes in sperm quality over time or comparing results between different clinics.
- Expert Disagreement: Research has confirmed a "high degree of inter-expert variability in the SMA" [2]. When different highly-trained embryologists evaluate the same sample, they frequently produce differing classifications and percentages of normal forms, calling into question the objective basis for critical clinical decisions.
- Technical Source of Variability: This variability stems not only from differences in visual interpretation but also from inconsistencies in sample preparation, including staining techniques and slide preparation, which can alter the appearance of sperm cells [2].
Excessive Workload and Operational Inefficiency
The manual process of sperm morphology analysis is exceptionally time-consuming and labor-intensive, creating practical barriers to consistent, high-quality assessment.
- Labor-Intensive Process: The requirement to evaluate a minimum of 200 spermatozoa per sample across multiple microscopic fields represents a "substantial workload" for technicians [2]. This process becomes particularly burdensome in high-volume clinical settings.
- Time-Consuming Procedures: Advanced manual techniques like the morphological examination of multiple sperm organelles for intracytoplasmic morphologically selected sperm injection (IMSI) "necessitate magnifications of >600× and are time-intensive procedures" [1]. This limits their practical application in routine clinical practice.
- Impact on Data Management: Many medical institutions using conventional methods fail to systematically save valuable image data, "leading to data loss" [2]. This represents both an operational inefficiency and a lost opportunity for building datasets needed to train and validate AI systems.
Table 1: Quantitative Evidence of Conventional Analysis Limitations
| Limitation Category | Evidence from Literature | Impact on Clinical Practice |
|---|---|---|
| Subjectivity | 26 types of abnormal morphology to classify visually [2] | Inconsistent application of diagnostic criteria |
| Inter-Observer Variability | "High degree of inter-expert variability" confirmed [2] | Reduced reliability for treatment decisions and longitudinal tracking |
| Operational Inefficiency | Analysis of ≥200 sperm per sample creates "substantial workload" [2] | Limited throughput and high labor costs in clinical settings |
| Data Management | Valuable image data often lost due to manual methods [2] | Lost opportunity for research and model development |
The AI Revolution: Addressing Conventional Limitations
Artificial intelligence research is directly targeting each of the fundamental limitations of conventional manual analysis through automated, data-driven approaches.
Enhanced Objectivity Through Automated Classification
AI systems replace subjective human judgment with consistent, algorithm-driven classification based on learned patterns from large datasets.
- Standardized Quantitative Assessment: Deep learning models apply the same mathematical criteria to every sperm cell analyzed, eliminating the cognitive biases and fatigue that affect human observers. A 2025 study demonstrated this enhanced objectivity, with an AI model achieving a precision of 0.95 and recall of 0.91 for detecting abnormal sperm morphology, and 0.91 precision with 0.95 recall for normal morphology [1].
- Detection of Subtle Features: AI models can be trained to identify specific subcellular features with high accuracy. One approach achieved F0.5 scores of 84.74% for acrosome abnormalities, 83.86% for head defects, and 94.65% for vacuole detection [11]. This level of consistent precision in detecting subtle features exceeds human capabilities.
- Objective Unstained Analysis: AI enables the assessment of unstained, live sperm morphology using confocal laser scanning microscopy at low magnification [1]. This breakthrough maintains sperm viability for subsequent use in ART procedures, overcoming a critical limitation of conventional staining methods that render sperm unusable.
Improved Consistency and Reduced Variability
By applying uniform classification standards, AI systems dramatically reduce both inter-observer and intra-observer variability.
- Superior Correlation with Standards: A 2025 experimental study comparing an in-house AI model to conventional methods found the AI model showed the strongest correlation with computer-aided semen analysis (r=0.88), followed by conventional semen analysis (r=0.76) [1]. The weaker correlation between conventional and computer-aided methods (r=0.57) highlights the variability inherent in manual assessment.
- High Inter-Operator Reliability: When urology residents were trained on an AI-based system, they achieved excellent inter-operator reliability (ICC=0.89) and intra-operator repeatability (ICC=0.92) for assessing progressive motility [12]. This demonstrates how AI systems can standardize assessments even across operators with different experience levels.
- Consistent Performance Metrics: In bovine sperm analysis, a YOLOv7-based deep learning system achieved a global mAP@50 of 0.73, precision of 0.75, and recall of 0.71 across multiple morphological categories [11]. These consistent performance metrics demonstrate the reduced variability compared to manual methods.
Dramatic Efficiency Improvements
AI automation addresses the workload burden through rapid, high-throughput analysis capabilities.
- Rapid Processing Speeds: One AI model demonstrated a processing time of approximately 139.7 seconds for 25,000 images, equating to an average prediction time of about 0.0056 seconds per image [1]. This represents a speed improvement of several orders of magnitude compared to manual evaluation.
- Streamlined Clinical Workflows: An AI-enabled commercial analyzer (LensHooke X1 PRO) provided results "approximately 1 minute after complete semen liquefaction" [12]. This rapid turnaround enables quicker clinical decision-making and enhances laboratory efficiency.
- High-Throughput Capability: The STAR (Sperm Tracking and Recovery) system for severe male factor infertility can scan through a semen sample, taking "more than 8 million images in under an hour" to identify rare viable sperm [13]. This capability is simply unattainable through manual search methods.
Table 2: AI Performance Metrics in Addressing Conventional Limitations
| AI Solution | Technical Approach | Performance Metrics |
|---|---|---|
| In-house AI Model for Unstained Sperm [1] | Deep learning with ResNet50 transfer learning on confocal microscopy images | Test accuracy: 0.93; Precision: 0.95 (abnormal), 0.91 (normal); Processing speed: 0.0056 s/image |
| Bovine Sperm Analysis System [11] | YOLOv7 object detection framework | mAP@50: 0.73; Precision: 0.75; Recall: 0.71 |
| STAR System for Severe Male Infertility [13] | High-powered imaging with AI identification and robotic capture | Capable of identifying viable sperm from 8+ million images in <1 hour |
| AI-Based Commercial Analyzer [12] | AI algorithms with autofocus optical technology | Results in ~1 minute post-liquefaction; Inter-operator ICC=0.89 |
Experimental Protocols in AI-Based Sperm Morphology Research
Protocol: AI Assessment of Unstained Live Sperm Morphology
A landmark 2025 study developed a novel methodology for assessing unstained live sperm using AI, providing a template for automated viability-preserving analysis [1].
Sample Preparation and Image Acquisition:
- Participants: 30 healthy male volunteers aged 18-40 years with 2-7 days of sexual abstinence [1].
- Sample Preparation: A 6 μL semen droplet was dispensed onto a standard two-chamber slide with a depth of 20μm (Leja) [1].
- Image Acquisition: Sperm images were captured using a confocal laser scanning microscope (LSM 800) at 40× magnification in confocal mode (LSM, Z-stack). The Z-stack interval was 0.5 μm, covering a total range of 2 μm. Five slides were generated per sample, each with a frame time of 633.03 ms and size of 512 × 512 pixels [1].
- Data Scale: At least 200 sperm images were collected per sample, with each capture containing 2-3 sperm cells [1].
Annotation and Model Development:
- Manual Annotation: Embryologists and researchers manually annotated well-focused sperm images using the LabelImg program, achieving a coefficient of correlation of 0.95 for normal sperm detection and 1.0 for abnormal sperm detection [1].
- Classification Criteria: Sperm were categorized into nine datasets based on WHO sixth edition criteria, including normal sperm with smooth oval head, length-to-width ratio of 1.5-2, no vacuoles, slender regular neck, uniform tail calibre, and cytoplasmic droplets less than one-third of the sperm head [1].
- Model Architecture: The study employed a ResNet50 transfer learning model, a deep neural network designed for image classification tasks [1].
- Training Parameters: The model was trained on a dataset of 21,600 images with 12,683 annotated sperm. The training used a subset of 9,000 images (4,500 normal morphology, 4,500 abnormal morphology) and was tested on 900 batches of previously unseen images [1].
Protocol: Deep Learning-Based Bovine Sperm Morphology Analysis
A 2025 veterinary study implemented a YOLOv7-based system for automated bovine sperm analysis, demonstrating the transferability of AI approaches across species [11].
Sample Collection and Processing:
- Subjects: Sexually active Brahman bulls over 24 months of age with at least 32 cm scrotal circumference [11].
- Semen Collection: Electroejaculation technique using a transrectal probe with preprogrammed electrical stimulus cycles. Semen was collected directly from the penis using a sterile collection bag [11].
- Sample Processing: Semen was diluted in a 1:1 ratio (v/v) with Optixcell extender in Eppendorf tubes prewarmed at 37°C to avoid temperature shock. Further dilution at 1:20 ratio achieved concentration of 17.5–27.5 (×10⁶/mL) [11].
Morphology Analysis and Image Capture:
- Slide Preparation: 10 μL from each diluted sample was placed on a slide, topped with a coverslip, and placed in the Trumorph system for fixation through brief exposure to 60°C and pressure of 6 kp [11].
- Microscopy: Fixed samples were evaluated under a B-383Phi microscope with 1× eyepiece and 40× negative phase contrast objective [11].
- Image Capture: Images were captured with the PROVIEW application and stored in JPG format [11].
- Classification Categories: Six morphological categories were used: (1) Normal, (2) Agglutination, (3) Dirt particles, (4) Folded tail, (5) Loose head, and (6) Loose tail [11].
Deep Learning Framework:
- Model: YOLOv7 object detection framework trained on a dataset of 277 annotated images containing six morphological categories [11].
- Performance: The system achieved global mAP@50 of 0.73, precision of 0.75, and recall of 0.71, demonstrating a balanced tradeoff between accuracy and efficiency [11].
Essential Research Reagent Solutions and Materials
Table 3: Key Research Reagents and Materials for AI-Based Sperm Morphology Analysis
| Item Name | Specification/Model | Research Function |
|---|---|---|
| Confocal Laser Scanning Microscope [1] | LSM 800 | High-resolution imaging of unstained live sperm at 40x magnification with Z-stack capability |
| Computer-Assisted Semen Analyzer [12] | LensHooke X1 PRO | AI-enabled portable analyzer for rapid assessment of concentration, motility, and morphology |
| Deep Learning Model [1] | ResNet50 transfer learning | Image classification architecture for distinguishing normal vs. abnormal sperm morphology |
| Object Detection Framework [11] | YOLOv7 | Real-time detection and classification of sperm abnormalities in microscopic images |
| Sperm Fixation System [11] | Trumorph system | Dye-free fixation through controlled pressure (6 kp) and temperature (60°C) |
| Microscope for Veterinary Use [11] | Optika B-383Phi | Bright-field microscopy with negative phase contrast for sperm morphology evaluation |
| Annotation Software [1] | LabelImg program | Manual annotation of sperm images for training dataset creation |
| Staining Method [1] | Diff-Quik stain (Romanowsky variant) | Conventional staining for comparative analysis in method validation studies |
The limitations of conventional manual sperm morphology analysis—subjectivity, variability, and excessive workload—represent fundamental challenges that have persisted despite technological advancements in other areas of laboratory medicine. The subjective interpretation of complex morphological criteria, combined with the labor-intensive nature of the process, has resulted in a test with acknowledged reliability issues that affect its clinical utility for infertility workups and treatment planning [10].
Artificial intelligence research is systematically addressing each of these limitations through automated, data-driven approaches. Deep learning models provide standardized quantitative assessment that eliminates human subjectivity, with studies demonstrating superior correlation with established methods and exceptional classification accuracy [1]. AI systems dramatically reduce inter-observer variability while processing images at speeds unattainable through manual methods, thereby addressing both reliability and efficiency concerns [1] [11].
The experimental protocols and technical approaches detailed in this review provide a roadmap for researchers and clinicians seeking to implement AI solutions in reproductive medicine. As these technologies continue to evolve, with growing adoption documented in global surveys of fertility specialists [14], they promise to transform sperm morphology analysis from a subjective, variable assessment into a precise, standardized component of male fertility evaluation. This transformation aligns with the broader movement toward data-driven, objective diagnostic methodologies across medicine, potentially leading to more accurate prognostication and improved outcomes in assisted reproduction.
The morphological evaluation of human spermatozoa remains a cornerstone of male fertility assessment, establishing a critical structure-function relationship that informs clinical diagnosis. This complex morphogenetic process during spermiogenesis produces highly differentiated cells designed to transport genetic material to the oocyte. The clinical examination of sperm morphology essentially represents a pathological assessment, where the presence of "ideal" spermatozoa suggests optimal fertilizing potential [15]. The World Health Organization (WHO) has systematically refined the standards for this assessment across multiple editions of its laboratory manual, creating a foundational framework for predicting conception potential based on semen quality parameters [6]. This technical guide explores the precise definitions of normal and abnormal morphological features across sperm compartments—head, neck, and tail—within the context of WHO standards, while framing this clinical target within the rapidly evolving field of artificial intelligence (AI) research in reproductive medicine.
The inherent challenge in sperm morphology analysis lies in the remarkable morphological heterogeneity of human sperm compared to other mammalian species. Even in fertile men, spermatozoa that are morphologically 'unfinished,' 'immature,' or malformed significantly outnumber those with "ideal" morphology [15]. This biological reality complicates clinical assessment and underscores the importance of precise, standardized classification systems. Furthermore, the selection process occurring naturally in the female genital tract filters out many abnormal forms, meaning the sperm population reaching the oocyte demonstrates markedly improved morphology compared to native semen samples [15]. This physiological selection process conceptually underpins the development of strict morphological criteria for clinical assessment.
WHO Standards and Compartment-Specific Morphological Definitions
Evolution of WHO Guidelines and the "Strict" Criteria
The WHO laboratory manual for semen analysis has undergone significant evolution, with successive editions published in 1980, 1987, 1992, 1999, 2010, and 2021 progressively refining the criteria for sperm morphology assessment [6]. The most transformative development was the introduction and consolidation of the "strict" morphology criteria, which fundamentally shifted assessment paradigms. Before strict criteria were implemented, classification methods often used vague definitions or no definitions at all, resulting in highly inconsistent results between observers, with reported percentages of normal spermatozoa as high as 80% and inter-observer differences exceeding 30% [15]. The strict method established rigorous, standardized definitions for morphologically normal spermatozoa based on the microscopic characteristics of well-proportioned spermatozoa recovered from the female genital tract [15].
According to the Kruger strict criteria, a spermatozoon is classified as normal only when it possesses a smooth, oval head with a well-defined acrosome covering 40-70% of the head area, no neck/midpiece or tail defects, and no cytoplasmic droplets of more than half the sperm head size [3] [16]. The current WHO threshold establishes that a sample with less than 4% normal forms is classified as teratozoospermia [16]. This strict classification system has dramatically improved inter-laboratory consistency but has also revealed that most sperm in even fertile men's samples don't meet these ideal standards, with typical values ranging from 4% to 10% normal forms [3].
Detailed Compartmental Analysis: Head, Neck, and Tail Defects
Sperm Head Abnormalities: The sperm head contains the highly condensed nucleus and acrosomal enzymes essential for oocyte penetration. Normal head dimensions are approximately 4.0-5.0 μm in length and 2.5-3.5 μm in width, with a smooth, oval configuration [15]. Head abnormalities represent the most clinically significant defects due to their direct impact on genetic material delivery and fertilization competence. Common head anomalies include:
- Macrocephalic and Microcephalic Sperm: Abnormally large or small heads often associated with chromosomal abnormalities.
- Pyriform Heads: Tapered, pear-shaped heads that impair hydrodynamic efficiency.
- Amorphous Heads: Irregularly shaped heads without defined structure.
- Vacuolated Heads: Presence of multiple or large vacuoles in the nuclear region.
- Acrosomal Abnormalities: Absent, small, or large acrosomes affecting fertilization capability.
The French BLEFCO Group specifically recommends that laboratories implement qualitative or quantitative methods for detecting monomorphic abnormalities like globozoospermia (round-headed sperm without acrosomes) and macrocephalic spermatozoa syndrome, as these conditions have profound implications for fertilization potential [10].
Neck and Midpiece Abnormalities: The neck region connects the sperm head to the tail and contains the centrioles, while the midpiece houses the mitochondria responsible for energy production. Common abnormalities include:
- Bent Necks: Sharp angulation at the head-neck junction.
- Asymmetric Midpiece Insertion: Off-center attachment of the midpiece to the head.
- Abnormally Thick or Thin Midpieces: Disrupted mitochondrial sheath organization.
- Cytoplasmic Droplets: Residual cytoplasm retained around the midpiece, considered normal if less than half the sperm head size.
Tail Abnormalities: The sperm tail (flagellum) provides motility through its complex axonemal structure. Critical tail defects include:
- Coiled Tails: Flagellum tightly coiled around itself.
- Bent Tails: Sharp angulations along the tail length.
- Short Tails: Abnormally truncated flagellum.
- Multiple Tails: Duplication of flagellar structures.
- Absent Tails: Complete lack of flagellum (rare).
Table 1: Comprehensive Classification of Sperm Morphological Abnormalities Based on WHO Standards
| Sperm Compartment | Abnormality Type | Morphological Description | Clinical Significance |
|---|---|---|---|
| Head | Macrocephalic | Abnormally large head, often with multiple flagella | Associated with genetic abnormalities; poor fertilization potential |
| Microcephalic | Abnormally small head | Often indicates chromosomal abnormalities | |
| Pyriform | Tapered, pear-shaped head | Altered hydrodynamics; reduced motility | |
| Amorphous | Irregular shape with undefined structure | Impaired zona pellucida binding | |
| Vacuolated | Large vacuoles in nuclear region | Potential DNA fragmentation concern | |
| Neck/Midpiece | Bent Neck | Sharp angulation at head-neck junction | Compromised energy transmission to tail |
| Asymmetric Insertion | Off-center midpiece attachment | Aberrant motility patterns | |
| Cytoplasmic Droplet | Residual cytoplasm >50% head size | Indicator of sperm immaturity | |
| Tail | Coiled | Flagellum tightly coiled around itself | Severely impaired or absent motility |
| Bent | Sharp angulation along tail length | Non-progressive motility | |
| Multiple | Two or more tail structures | Complete dysfunction | |
| Absent | Lack of flagellum | Non-motile |
The Analytical Challenge: Variability and Standardization in Morphology Assessment
Subjectivity and Training Deficiencies
Sperm morphology assessment remains one of the most challenging and variable tests in andrology laboratories, primarily due to its subjective nature and lack of standardized training protocols. Unlike sperm concentration and motility, which can be objectively measured with computer-assisted systems, morphology assessment relies heavily on technician expertise and judgment [17]. This subjectivity introduces significant variability, with studies showing that even expert morphologists only agreed on normal/abnormal classification for 73% of sperm images when using a simple binary system [17]. The problem compounds with more complex classification systems; untrained users achieved only 53% accuracy when using a detailed 25-category classification system compared to 81% accuracy with a simple 2-category (normal/abnormal) system [17].
The variability stems from multiple factors, including differences in staining techniques, microscope optics, individual interpretation of criteria, and the inherent difficulty of classifying complex morphological anomalies. Recent research has demonstrated that without standardized training, novice morphologists show high variation (coefficient of variation = 0.28) and widely ranging accuracy scores from 19% to 77% [17]. This alarming variability has serious clinical implications, as morphology assessment directly influences treatment decisions, including the selection of appropriate assisted reproductive technologies.
Quality Control and Standardization Efforts
Efforts to standardize sperm morphology assessment have focused on both analytical protocols and training methodologies. External quality control programs such as the German QuaDeGA and UK NEQAS provide limited proficiency testing, but these are often implemented infrequently due to expense and availability constraints [17]. When morphologists fail quality control assessments, recommended re-training typically involves side-by-side assessment with a senior morphologist, introducing potential bias from the trainer's own subjective interpretations [17].
The emergence of standardized training tools based on machine learning principles represents a significant advancement. These tools utilize "ground truth" datasets established through expert consensus, similar to the methodology used for training AI models. Studies have demonstrated that structured training using these tools can dramatically improve accuracy, with novice morphologists achieving final accuracy rates of 98% (2-category), 97% (5-category), 96% (8-category), and 90% (25-category) across different classification systems [17]. Furthermore, training significantly reduces assessment time, from 7.0±0.4 seconds to 4.9±0.3 seconds per image, enhancing laboratory efficiency [17].
Table 2: Impact of Training on Morphology Assessment Accuracy Across Classification Systems
| Classification System | Number of Categories | Untrained Accuracy | Trained Accuracy | Improvement |
|---|---|---|---|---|
| Binary | 2 (Normal/Abnormal) | 81.0% ± 2.5% | 98.0% ± 0.43% | +17.0% |
| Location-Based | 5 (Head, Midpiece, Tail, Cytoplasmic Droplet, Normal) | 68.0% ± 3.59% | 97.0% ± 0.58% | +29.0% |
| Extended Bovine | 8 (Various specific defects) | 64.0% ± 3.5% | 96.0% ± 0.81% | +32.0% |
| Comprehensive | 25 (All defects defined individually) | 53.0% ± 3.69% | 90.0% ± 1.38% | +37.0% |
Artificial Intelligence in Sperm Morphology Analysis
From Traditional Machine Learning to Deep Learning Approaches
Artificial intelligence is revolutionizing sperm morphology analysis by introducing objectivity, standardization, and high-throughput capabilities to a traditionally subjective domain. AI applications in this field have evolved from conventional machine learning (ML) approaches to sophisticated deep learning (DL) algorithms capable of extracting intricate features directly from sperm images [6]. Conventional ML techniques, including K-means clustering, support vector machines (SVM), and decision trees, initially demonstrated promising results but were fundamentally limited by their reliance on manually engineered features (e.g., grayscale intensity, edge detection, contour analysis) and non-hierarchical structures [18]. For instance, early Bayesian Density Estimation models achieved approximately 90% accuracy in classifying sperm heads into four morphological categories, but their performance was constrained by focusing exclusively on shape-based features [18].
The paradigm shift toward deep learning has addressed many of these limitations through automated feature extraction and enhanced pattern recognition capabilities. Deep learning, characterized by neural networks with multiple hidden layers (typically more than three layers including inputs and outputs), excels at processing complex image data without requiring manual feature specification [5] [18]. These algorithms automatically learn hierarchical representations from raw pixel data, enabling them to detect subtle morphological patterns often imperceptible to human observers. The distinguishing advantage of DL is its scalability—as larger and more diverse datasets become available, model performance continues to improve without architectural changes, earning it the designation of "scalable machine learning" [5].
Technical Architectures and Implementation Frameworks
Deep learning applications in sperm morphology analysis primarily utilize convolutional neural networks (CNNs) optimized for image segmentation and classification tasks. The technical pipeline typically involves two critical stages: accurate automated segmentation of sperm morphological structures (head, neck, and tail), followed by efficient classification of normal and abnormal forms [18]. Advanced architectures like U-Net and Mask R-CNN have demonstrated particular efficacy in sperm segmentation tasks, achieving precise delineation of sperm components even in challenging imaging conditions [18].
More sophisticated approaches integrate multiple neural networks in ensemble methods or leverage transfer learning to adapt models pre-trained on large natural image datasets (e.g., ImageNet) to the specialized domain of sperm morphology [6]. The emergence of transformer architectures and vision-language models represents the cutting edge, potentially enabling more contextual understanding of morphological features and their clinical correlations. These technical advancements directly address the core challenges of traditional morphology assessment by providing consistent, quantitative, and high-throughput analysis capabilities essential for both clinical diagnostics and research applications.
Experimental Protocols and Research Methodologies
Traditional Morphology Assessment Protocol
The conventional sperm morphology assessment protocol follows standardized methodology outlined in the WHO laboratory manual. The essential steps include:
Sample Preparation: Semen samples are collected after 2-7 days of sexual abstinence and allowed to liquefy for 15-30 minutes at 37°C. A standardized smear is prepared using 5-10μL of well-mixed semen spread evenly across a clean glass slide.
Staining Procedure: Slides are air-dried and fixed using methanol for 5-15 minutes. Various staining techniques can be employed, including:
- Diff-Quik Staining: Fixed slides are dipped in solution I (eosin) for 10-15 seconds, solution II (methylene blue) for 10-15 seconds, then rinsed gently with distilled water and air-dried.
- Papanicolaou Staining: A more complex protocol involving sequential immersion in Harris hematoxylin, Orange G, and EA-50 solutions with multiple rinsing steps.
- Other Methods: Shorr staining, Bryan-Leishman staining, or rapid staining methods may be used depending on laboratory preferences.
Microscopic Evaluation: Stained slides are examined under oil immersion at 1000x magnification. A minimum of 200 spermatozoa are systematically evaluated across multiple microscopic fields. Each spermatozoon is classified according to strict criteria, noting specific abnormalities in the head, neck, and tail compartments.
Quality Control: Regular participation in external quality assurance programs and internal validation procedures ensures ongoing accuracy and consistency. Laboratories should maintain inter-technician variability of less than 5-10% for morphology assessments.
Advanced Imaging and AI Integration Protocols
Emerging technologies have introduced sophisticated protocols that enhance traditional morphology assessment:
Digital Holographic Microscopy (DHM) Protocol: DHM enables non-invasive, label-free morphological assessment of live spermatozoa in three dimensions, bypassing artifacts introduced by staining and fixation procedures [16]. The experimental workflow involves:
- Sample Preparation: Fresh semen samples are allowed to liquefy completely. A small droplet (5-10μL) is placed on a microscope slide without fixation or staining.
- Hologram Acquisition: The sample is placed on the DHM stage and illuminated with a coherent laser light source. Interference patterns between the object beam (transmitted through the sample) and reference beam are recorded by a CCD camera as digital holograms.
- Numerical Reconstruction: Recorded holograms are processed through numerical back-propagation algorithms to reconstruct quantitative phase images of individual spermatozoa.
- 3D Parameter Extraction: The methodology extracts novel three-dimensional morphological parameters, including head height, acrosome/nucleus height, and head/midpiece height, which show less variability in fertile men compared to infertile patients [16].
- Motility Integration: DHM simultaneously assesses sperm motility parameters, providing correlated morphological and functional data from the same cells.
AI-Based Morphology Analysis Protocol: The integration of artificial intelligence follows a structured pipeline:
- Dataset Curation: Collect and annotate large datasets of sperm images using expert consensus to establish "ground truth" labels. Public datasets include HSMA-DS, MHSMA, VISEM-Tracking, and SVIA, though limitations in size and quality persist [18].
- Model Selection and Training: Choose appropriate deep learning architectures (e.g., CNN, U-Net) and train models using the annotated datasets. Implement data augmentation techniques to enhance model robustness and generalization.
- Validation and Testing: Evaluate model performance using separate validation and test datasets not seen during training. Assess metrics including accuracy, precision, recall, F1-score, and area under the ROC curve.
- Clinical Implementation: Deploy validated models in clinical settings, either as standalone systems or as decision-support tools alongside manual assessment. Ensure continuous monitoring and model updating as new data becomes available.
Table 3: Research Reagent Solutions for Sperm Morphology Analysis
| Reagent/Equipment | Application Purpose | Technical Specifications | Experimental Considerations |
|---|---|---|---|
| Methanol | Slide fixation | Analytical grade, 100% concentration | Fix for 5-15 minutes; ensures cellular preservation |
| Diff-Quik Stain | Sperm staining | Commercial staining kit | Rapid staining (30 seconds total); consistent results |
| Percoll Gradient | Sperm selection | 90% and 45% layers | Selects morphologically normal sperm for ART |
| Digital Holographic Microscope | Live sperm imaging | Laser source, CCD camera, reconstruction software | Enables 3D morphological analysis without staining |
| Phase Contrast Optics | Unstained sperm viewing | 1000x magnification with oil immersion | Reduces staining artifacts in assessment |
| AI Training Datasets | Model development | SVIA: 125,000 annotated instances | Quality annotation is critical for model accuracy |
| Computer-Assisted Semen Analysis (CASA) | Automated assessment | Integrated optics and analysis software | Must be validated against manual methods |
Data Integration and Clinical Correlation
Quantitative Morphological Parameters and Fertility Correlations
The clinical value of sperm morphology assessment lies in its correlation with fertility outcomes, though this relationship is complex and multifactorial. Traditional 2D morphological parameters include head length (4.0-5.0μm), head width (2.5-3.5μm), midpiece length (3.0-5.0μm), and tail length (approximately 45μm) [15] [16]. Advanced 3D parameters obtained through digital holographic microscopy reveal additional discriminatory power, with studies showing reduced variability in parameters like head height, acrosome/nucleus height, and head/midpiece height in fertile men compared to infertile patients [16].
The teratozoospermic index (TZI) and other multiple anomaly indices (sperm deformity index - SDI, multiple anomalies index - MAI) provide composite scores that quantify the average number of defects per abnormal spermatozoon. Research indicates mean TZI values of approximately 1.31±0.17 in fertile men compared to 1.45±0.12 in infertile patients, though statistical significance between groups is not always achieved [16]. The French BLEFCO Group's recent guidelines, however, question the clinical utility of these indices, stating there is "insufficient evidence to demonstrate the clinical value of indexes of multiple sperm defects (TZI, SDI, MAI) in investigation of infertility and before ART" [10].
The most significant clinical correlation exists between specific monomorphic abnormalities and fertilization failure. Conditions like globozoospermia (round-headed acrosomeless sperm) and macrocephalic spermatozoa syndrome demonstrate virtually zero fertilization potential without technological intervention, highlighting the critical importance of detecting these specific morphological patterns [10].
AI-Enhanced Predictive Modeling
Artificial intelligence enables more sophisticated predictive modeling by integrating morphological data with clinical outcomes. Machine learning algorithms can identify complex, non-linear relationships between specific morphological patterns and reproductive success that escape conventional statistical analysis. Supervised learning approaches have been applied to:
- Predict fertilization success in IVF cycles based on sperm morphological patterns
- Identify patients with specific genetic conditions (e.g., Klinefelter syndrome) from azoospermic samples
- Forecast improvement in semen parameters following medical or surgical interventions (e.g., varicocelectomy)
- Select optimal sperm for intracytoplasmic sperm injection (ICSI) based on subtle morphological features
Random forest models have demonstrated superior performance compared to traditional logistic regression in predicting post-varicocelectomy sperm analysis improvement, highlighting the power of ensemble ML methods in andrological applications [5]. Furthermore, deep learning systems can process the continuum of sperm biometrics rather than relying on binary classifications, potentially uncovering novel morphological biomarkers of fertility potential.
Future Directions and Research Implications
Technological Advancements and Methodological Innovations
The future of sperm morphology analysis lies in the continued integration of advanced technologies that enhance objectivity, throughput, and predictive value. Several promising directions are emerging:
Multi-Modal Data Integration: Next-generation systems will combine morphological data with proteomic, genomic, and metabolomic profiles to create comprehensive sperm quality assessments. The correlation between specific morphological defects and molecular abnormalities will enable more precise diagnosis of infertility etiology and targeted therapeutic interventions.
Advanced Imaging Technologies: Techniques like digital holographic microscopy and inferometric phase microscopy will continue to evolve, providing label-free, quantitative 3D morphological data from live spermatozoa without processing artifacts [16]. These technologies enable longitudinal studies of the same sperm cells, potentially revealing dynamic morphological changes associated with capacitation and other functional processes.
Explainable AI in Morphology Assessment: As AI systems become more complex, research focus will shift toward developing explainable AI that provides transparent rationale for morphological classifications. This will enhance clinical trust and potentially reveal novel morphological biomarkers not previously recognized by human experts.
Standardized Dataset Development: A critical priority is the creation of large, diverse, and high-quality annotated datasets to support robust AI model development. Current public datasets (HSMA-DS, MHSMA, VISEM-Tracking, SVIA) suffer from limitations in sample size, image quality, and annotation consistency [18]. International collaborative efforts to establish standardized datasets with expert-validated "ground truth" annotations will significantly advance the field.
Clinical Translation and Personalized Medicine
The ultimate goal of technological advancement in sperm morphology analysis is improved patient care through personalized treatment strategies. Future clinical applications may include:
Precision ART Selection: AI-based morphology analysis will provide more accurate predictions of which assisted reproductive technology (IUI, IVF, or ICSI) is most appropriate for individual couples based on specific morphological patterns. The French BLEFCO Group currently recommends against using normal morphology percentage alone for ART selection [10], but more sophisticated multidimensional assessments may restore the prognostic value of morphology.
Sperm Selection Algorithms: Real-time AI systems may guide embryologists in selecting individual spermatozoa for ICSI based on comprehensive morphological analysis correlated with clinical outcomes. This would extend beyond current IMSI (intracytoplasmic morphologically selected sperm injection) practices by incorporating subtle features detectable only through computational analysis.
Therapeutic Monitoring: Advanced morphology assessment will enable more precise monitoring of medical or surgical interventions for male infertility, providing objective metrics of treatment response and guiding therapeutic adjustments.
Public Health Applications: Large-scale morphology screening coupled with AI analysis could identify environmental or occupational factors affecting sperm health, contributing to public health initiatives aimed at addressing declining semen quality trends observed in various populations [18].
As these technologies mature, the field must simultaneously develop appropriate regulatory frameworks, validation standards, and ethical guidelines to ensure their responsible implementation in clinical practice. The integration of artificial intelligence with established WHO standards represents not a replacement of traditional methods, but rather an enhancement that preserves clinical wisdom while augmenting it with computational power and objectivity.
The assessment of sperm quality represents a cornerstone in the evaluation of male fertility, with sperm morphology analysis serving as a critical predictor of reproductive success. Traditional manual semen analysis has long been plagued by subjectivity, inter-observer variability, and labor-intensive processes, limiting its reproducibility and clinical utility [2]. The emergence of Computer-Aided Sperm Analysis (CASA) systems initially promised to overcome these limitations through automation and standardization. However, early CASA systems demonstrated significant limitations in analyzing complex parameters like sperm morphology, particularly in distinguishing subtle defects across the head, neck, and tail compartments [19]. The integration of artificial intelligence (AI), particularly deep learning algorithms, has catalyzed a revolutionary shift from automated measurement to intelligent diagnostic interpretation, enabling unprecedented accuracy in sperm quality assessment while revealing novel biomarkers predictive of fertility outcomes [6] [2].
This evolution mirrors broader trends in biomedical imaging, where AI has demonstrated transformative potential in applications ranging from synthetic contrast generation in radiology to embryo selection in assisted reproduction [20]. The convergence of advanced imaging technologies with sophisticated machine learning algorithms has created a new paradigm in which sperm analysis transcends traditional morphological assessment to encompass functional evaluation, including DNA integrity and kinematic patterns [21] [6]. This technical review examines the architectural foundations, methodological frameworks, and clinical validation of AI-driven sperm analysis systems within the context of a broader thesis on how sperm morphology analysis operates within contemporary AI research, providing researchers, scientists, and drug development professionals with a comprehensive understanding of this rapidly evolving field.
The Technical Evolution: From Conventional CASA to AI-Driven Architectures
Limitations of Conventional CASA Systems
First-generation CASA systems established the fundamental principle of automated sperm analysis through computer vision techniques, but their architectural constraints limited their diagnostic accuracy and clinical utility. These systems primarily relied on threshold-based image processing and manual feature engineering, extracting basic parameters such as sperm concentration, motility, and elementary morphology [6] [19]. Performance evaluations revealed critical vulnerabilities, particularly with challenging samples; the coefficient of variation (CV) for sperm concentration and progressive motility (PR) significantly increased with decreasing sperm concentration (r = -0.561, p = 0.001) and PR values (r = -0.621, p < 0.001), rendering them unreliable for severe oligozoospermia and asthenozoospermia cases [19].
The technical limitations extended to morphological assessment, where conventional CASA systems demonstrated limited capability in segmenting complete sperm structures. These systems typically achieved high coincidence rates for overall sperm morphology (99.40%) and head morphology (99.67%) when compared to manual methods, but this apparent accuracy masked fundamental deficiencies in detecting midpiece and tail abnormalities [19]. The reliance on handcrafted features (e.g., grayscale intensity, edge detection, contour analysis) made these systems susceptible to over-segmentation or under-segmentation artifacts, particularly with overlapping sperm or debris-rich samples [2]. The algorithmic constraints manifested in classification inaccuracies, with some conventional machine learning approaches achieving only 49% accuracy for non-normal sperm head classification, significantly below clinical requirements [2].
The AI Revolution: Machine Learning to Deep Learning
The integration of artificial intelligence represents a architectural paradigm shift from programmed algorithms to learned feature representation. This transition encompasses both conventional machine learning and deep learning approaches, each with distinct methodological frameworks and performance characteristics, as detailed in Table 1.
Table 1: Evolution of Algorithmic Approaches in Sperm Morphology Analysis
| Algorithm Type | Key Examples | Technical Approach | Performance Characteristics | Primary Limitations |
|---|---|---|---|---|
| Conventional Machine Learning | Support Vector Machines (SVM), K-means clustering, Bayesian Density Estimation | Manual feature extraction (Hu moments, Zernike moments, Fourier descriptors) combined with classifiers | Accuracy: 49-90% depending on feature set; SVM achieved AUC-ROC of 88.59% for head classification [2] | Limited to pre-defined features; poor generalization; inability to detect complete sperm structures |
| Deep Learning | CNN (ResNet50), U-Net, GANs, Transformer networks (GC-ViT) | Automated feature extraction from raw pixel data; hierarchical representation learning | Test accuracy: 93%; precision: 0.95 for abnormal morphology; processing speed: 0.0056s/image [1] | Requires large annotated datasets; computational intensity; "black box" interpretation challenges |
Conventional machine learning approaches established the foundation for automated sperm analysis but faced fundamental constraints. Techniques such as Bayesian Density Estimation achieved 90% accuracy in classifying sperm heads into four morphological categories, while SVM classifiers demonstrated strong discriminatory power with 88.59% area under the receiver operating characteristic curve (AUC-ROC) and precision rates above 90% [2]. However, these systems required explicit programming of feature extraction algorithms, limiting their adaptability to the complex, high-dimensional patterns in sperm morphology.
Deep learning architectures overcome these limitations through hierarchical feature learning, enabling the automatic discovery of relevant morphological patterns from raw image data. The ResNet50 transfer learning model, trained on confocal laser scanning microscopy images, exemplifies this approach, achieving a test accuracy of 0.93 after 150 epochs with precision of 0.95 and recall of 0.91 for detecting abnormal sperm morphology [1]. Ensemble methods that combine multiple architectures, such as morphology-assisted AI models incorporating transformer-based GC-ViT, have demonstrated capability in predicting DNA fragmentation from phase contrast images with 60% sensitivity and 75% specificity, establishing correlations between morphological features and functional fertility parameters [21].
Experimental Frameworks and Methodological Standards
Dataset Development and Annotation Protocols
The performance of AI-based sperm morphology analysis is intrinsically linked to the quality, diversity, and scale of training datasets. Significant research efforts have focused on addressing the historical limitations in sperm image data collection through standardized acquisition protocols and annotation frameworks. The development of high-resolution, low-magnification datasets using confocal laser scanning microscopy (LSM 800) at 40× magnification with Z-stack intervals of 0.5μm represents a methodological advancement, capturing detailed morphological information across a total range of 2μm [1]. This approach generates comprehensive image data with frame times of 633.03ms and image sizes of 512×512 pixels, covering a physical area of 159.7×159.7μm per slide.
The annotation process establishes the ground truth for model training, typically involving manual bounding box placement around well-focused sperm using programs such as LabelImg. Expert embryologists and researchers achieve high inter-annotator reliability, with correlation coefficients of 0.95 for normal sperm morphology detection and 1.0 for abnormal morphology detection [1]. Categorization follows WHO sixth edition guidelines, classifying sperm into nine distinct datasets based on comprehensive morphological criteria including smooth oval head appearance, length-to-width ratio of 1.5-2, absence of vacuoles, slender and regular neck structure, uniform tail calibre, and cytoplasmic droplets less than one-third of the sperm head size [1]. Contemporary datasets have dramatically expanded in scale and annotation depth, with the SVIA (Sperm Videos and Images Analysis) dataset comprising 125,000 annotated instances for object detection, 26,000 segmentation masks, and 125,880 cropped image objects for classification tasks [2].
AI Model Development and Validation Workflows
The implementation of AI models for sperm analysis follows structured computational workflows encompassing data preprocessing, architecture selection, training, and validation. The following DOT language visualization illustrates a standardized pipeline for developing deep learning models in sperm morphology analysis:
Diagram 1: AI Model Development Workflow for Sperm Morphology Analysis
The experimental workflow implements rigorous validation protocols to ensure model robustness and generalizability. Internal validation during training continuously assesses performance on holdout data not used in training, typically reporting metrics such as precision (0.95 for abnormal sperm), recall (0.91 for abnormal sperm), and overall test accuracy (0.93) [1]. External validation represents a critical step, evaluating model performance on completely separate datasets from different clinical environments, with correlation analyses comparing AI results with established reference methods including CASA and conventional semen analysis (CSA) [1] [20]. This multi-stage validation framework ensures that reported performance metrics reflect real-world clinical utility rather than optimized performance on training data.
Research Reagent Solutions and Essential Materials
The experimental implementation of AI-enhanced sperm analysis requires specific technical resources and reagent systems. The following table catalogues essential research solutions and their functions within the methodological framework:
Table 2: Essential Research Reagent Solutions for AI-Based Sperm Morphology Analysis
| Resource Category | Specific Examples | Technical Function | Implementation Context |
|---|---|---|---|
| Imaging Systems | Confocal Laser Scanning Microscope (LSM 800), NIKON Eclipse Ci with phase contrast, IP103100A digital camera | High-resolution image acquisition; Z-stack capability for 3D reconstruction; phase contrast for unstained samples | Unstained live sperm imaging; dataset development [1] [19] |
| Staining Reagents | Diff-Quik stain (Romanowsky variant), Papanicolaou staining solutions | Cellular contrast enhancement; nuclear and acrosomal detail differentiation | Conventional morphology reference standard; fixed sperm analysis [19] |
| Analysis Platforms | GSA-810 system, LensHooke X1 PRO, IVOS II, Sperm Class Analyzer (SCA) | Automated sperm tracking; parameter quantification; AI algorithm integration | Clinical validation; performance benchmarking [22] [19] |
| Quality Control Materials | Latex bead suspensions (high: 80±8×10⁶/mL; low: 15±1.5×10⁶/mL) | Accuracy verification; precision monitoring; system calibration | Daily quality assurance; method validation [19] |
| Annotation Software | LabelImg program, Custom annotation interfaces | Bounding box placement; morphological classification; dataset labeling | Ground truth establishment; training data preparation [1] |
| Deep Learning Frameworks | TensorFlow, PyTorch, Custom implementations (ResNet50, U-Net, GANs) | Model architecture implementation; transfer learning; performance optimization | Algorithm development; experimental validation [1] [2] |
The integration of these resources enables the comprehensive implementation of AI-enhanced sperm analysis, from initial image acquisition through final clinical validation. The LensHooke X1 PRO exemplifies the convergence of these technologies, combining AI algorithms with autofocus optical technology to assess semen parameters with a 40× objective (numerical aperture 0.65), frame rate of 60 fps, and field of view of 500×500μm, while tracking sperm trajectories over ≥30 consecutive frames [22]. This technological integration facilitates rapid analysis, with results available approximately one minute after complete semen liquefaction, representing a significant advancement over traditional manual methods [22].
Performance Benchmarking and Clinical Validation
Quantitative Performance Metrics
The transition from conventional CASA to AI-enhanced systems has demonstrated measurable improvements in analytical performance across multiple parameters. Validation studies employing standardized quality control materials, such as latex bead suspensions with nominal values of (80.00 ± 8.0) × 10⁶/mL and (15.00 ± 1.5) × 10⁶/mL, confirm the analytical accuracy of modern systems, with detection values consistently within target ranges [19]. The quantitative advancement is particularly evident in morphological assessment, where AI-based systems achieve correlation coefficients of 0.88 with computer-aided semen analysis and 0.76 with conventional semen analysis, exceeding the correlation between CASA and conventional methods (r = 0.57) [1].
The performance characteristics of AI systems extend beyond correlation metrics to encompass diagnostic precision and operational efficiency. Deep learning models demonstrate precision of 0.95 and recall of 0.91 for detecting abnormal sperm morphology, with complementary performance for normal sperm morphology (precision: 0.91, recall: 0.95) [1]. Computational efficiency enables high-throughput analysis, with processing times of approximately 139.7 seconds for 25,000 images, corresponding to an average prediction time of 0.0056 seconds per image [1]. This combination of diagnostic accuracy and operational efficiency represents a significant advancement over conventional CASA systems, which exhibited poor repeatability for oligozoospermia and asthenozoospermia samples, limiting their clinical utility for severe male factor infertility [19].
Clinical Workflow Integration and Validation
The implementation of AI-based sperm analysis within clinical environments requires validation of both analytical performance and operational integration. Prospective studies evaluating AI systems operated by urology residents demonstrate the clinical translation of these technologies, with structured training protocols (8-hour didactic modules plus 10 hours of supervised hands-on sessions) yielding technical competency evidenced by inter-operator variability for progressive motility of ICC = 0.89 and intra-operator repeatability of ICC = 0.92 [22]. This operational reliability enables the detection of clinically significant improvements following therapeutic interventions, with AI-CASA systems documenting statistically significant postoperative enhancements across multiple conventional and nonconventional sperm parameters in patients undergoing varicocelectomy [22].
The following DOT language visualization illustrates the experimental framework for clinical validation of AI-based sperm analysis systems:
Diagram 2: Clinical Validation Framework for AI-Based Sperm Analysis
The clinical validation of AI systems extends beyond analytical performance to encompass practical utility in therapeutic decision-making. The ability to detect subtle improvements in sperm parameters following medical interventions provides clinicians with objective metrics for evaluating treatment efficacy [22]. Furthermore, the correlation between AI-derived morphological assessments and functional parameters such as DNA fragmentation establishes a foundation for predictive models that transcend traditional morphology-function relationships [21]. This evolution from descriptive morphology to predictive analytics represents the culmination of the transition from conventional CASA to AI-driven objective assessment, positioning sperm morphology analysis as a cornerstone of personalized fertility care.
The evolution from Computer-Aided Semen Analysis to artificial intelligence represents a fundamental transformation in objective sperm assessment, transitioning from automated measurement systems to intelligent diagnostic platforms. This paradigm shift encompasses technological advancements in imaging systems, algorithmic innovations in deep learning architectures, and methodological refinements in validation protocols, collectively enabling unprecedented accuracy in sperm morphology classification, segmentation, and functional prediction. The integration of convolutional neural networks, generative adversarial networks, and transformer-based models has addressed historical limitations in conventional CASA systems, particularly in analyzing complex morphological patterns and correlating structural features with functional fertility parameters.
Future research directions will likely focus on several critical areas, including the development of standardized, multi-center datasets to enhance model generalizability, the integration of multi-modal data streams encompassing morphological, kinematic, and molecular parameters, and the implementation of explainable AI techniques to address the "black box" limitations of complex deep learning models [6] [2]. Additionally, the correlation between AI-derived morphological assessments and clinical outcomes such as fertilization rates, embryo quality, and live birth rates will establish the ultimate validation of these technologies in reproductive medicine [22] [6]. As AI-based sperm analysis continues to evolve, its integration with emerging technologies in genomics, proteomics, and metabolomics will further advance personalized fertility diagnostics, ultimately transforming the evaluation and treatment of male factor infertility through data-driven, objective assessment methodologies.
Architectures in Action: Deep Learning Models and Workflows for Automated Morphology Classification
The accurate assessment of sperm morphology is a critical determinant in the diagnosis of male infertility. Traditional methods, which rely on manual visual inspection under a microscope, are inherently subjective, time-consuming, and prone to inter-observer variability [23] [2]. The World Health Organization (WHO) recommends the examination of at least 200 sperm per patient for a reliable diagnosis, a process that is often impractical in routine clinical settings, leading to compromises in diagnostic consistency [23]. While Computer-Aided Sperm Analysis (CASA) systems offered improvements, their adoption has been limited by high costs and operational complexities [23]. These challenges have created a critical gap in reproductive medicine, fueling the pursuit of fully automated, objective, and highly accurate analytical systems. Artificial intelligence (AI), particularly deep learning, has emerged as a transformative solution. This whitepaper explores the evolution and application of core AI model architectures—from established Convolutional Neural Networks (CNNs) like ResNet to the emerging paradigm of Vision Transformers (ViTs)—in advancing the field of automated sperm morphology analysis for researchers and drug development professionals.
Comparative Analysis of Deep Learning Architectures
The journey towards automation in sperm morphology analysis has been driven by successive generations of deep learning architectures, each offering distinct advantages and limitations.
2.1 Convolutional Neural Networks (CNNs) and ResNet CNNs have been the workhorse of medical image analysis, including sperm morphology classification. Their design, featuring convolutional layers that act as learnable filters, is inherently well-suited for identifying local spatial features such as edges, textures, and shapes in sperm images (e.g., head contour, acrosome presence, tail structure) [23]. Transfer learning, where a pre-trained network like VGG16 or GoogleNet is fine-tuned on sperm datasets, has been a common and effective strategy [23].
ResNet (Residual Network) advanced CNN capabilities by introducing residual connections, or "skip connections," that mitigate the vanishing gradient problem. This innovation enabled the training of much deeper networks, leading to more powerful feature representations. In sperm analysis, an ensemble of six custom CNNs achieved accuracies of 85.18% on the HuSHeM dataset and 90.73% on the SMIDS dataset [23]. Another study utilizing a two-stage fine-tuning strategy with VGG-16 and GoogleNet reported accuracies of 92.1% on HuSHeM and 90.87% on SMIDS, demonstrating the effectiveness of sophisticated CNN-based approaches [23].
Table 1: Performance of CNN-Based Models on Benchmark Sperm Morphology Datasets
| Model Architecture | Dataset | Key Methodology | Reported Accuracy | Reference |
|---|---|---|---|---|
| Ensemble of 6 CNNs | HuSHeM | Hard & Soft Voting | 85.18% | [23] |
| Ensemble of 6 CNNs | SMIDS | Hard & Soft Voting | 90.73% | [23] |
| VGG-16 & GoogleNet | HuSHeM | Two-Stage Fine-Tuning | 92.1% | [23] |
| VGG-16 & GoogleNet | SMIDS | Two-Stage Fine-Tuning | 90.87% | [23] |
| Custom CNN | SMD/MSS | Data Augmentation | 55% - 92% | [24] |
2.2 YOLO (You Only Look Once) While not directly featured in the provided literature for classification, the YOLO architecture is highly relevant for a complete automated sperm analysis pipeline. Before morphology can be classified, individual sperm must be located and segmented within a larger microscopic field of view that may contain debris and other cells. YOLO is a state-of-the-art single-shot object detection algorithm that performs both localization (drawing bounding boxes) and classification in one pass of the network, making it extremely fast and efficient. Its potential application lies in the initial "sperm detection" stage, identifying and cropping individual sperm cells for subsequent detailed morphological analysis by a CNN or ViT classifier [2].
2.3 Emerging Vision Transformers (ViTs) Vision Transformers represent a paradigm shift from the inductive biases of CNNs. Originally developed for natural language processing, ViTs process images by dividing them into a sequence of patches, which are then linearly embedded and processed by a transformer encoder with a self-attention mechanism [23]. This allows the model to capture global dependencies and long-range interactions between different parts of an image from the very first layer.
In sperm morphology analysis, this capability translates to a more holistic understanding of the entire sperm structure—for instance, simultaneously relating the shape of the head to the integrity of the midpiece and tail. A seminal 2025 study demonstrated that pure ViT architectures consistently outperform traditional CNN-based methods [23] [25]. After extensive hyperparameter optimization, the BEiT_Base model achieved state-of-the-art accuracies of 93.52% on the HuSHeM dataset and 92.5% on the SMIDS dataset, surpassing the previous best CNN-based approaches by 1.42% and 1.63%, respectively [23]. These improvements were statistically significant (p < 0.05, t-test). Visualization techniques like Attention Maps and Grad-CAM confirmed ViTs' superior ability to focus on discriminative morphological features, validating their clinical relevance [23].
Table 2: Vision Transformer vs. CNN Performance Benchmarking
| Model Type | Specific Model | HuSHeM Accuracy | SMIDS Accuracy | Key Advantage |
|---|---|---|---|---|
| Vision Transformer | BEiT_Base | 93.52% | 92.5% | Captures global context & long-range dependencies |
| CNN (Previous SOTA) | VGG-16/GoogleNet | 92.1% | 90.87% | Strong local feature extraction |
| Performance Delta | +1.42% | +1.63% | Statistically significant improvement (p<0.05) |
Experimental Protocols for Model Evaluation
Robust experimental design is paramount for validating the performance of AI models in a clinical context. The following protocol, derived from recent comparative studies, outlines a standardized methodology.
3.1 Dataset Preparation and Curation
- Datasets: Use publicly available, benchmark datasets such as the Human Sperm Head Morphology (HuSHeM) dataset and the Sperm Morphology Image Data Set (SMIDS) [23]. HuSHeM contains 216 images across four classes (normal, pyriform, tapered, amorphous), while SMIDS contains approximately 3,000 images across three classes (normal, abnormal, non-sperm).
- Pre-processing: For a fully automated pipeline, use raw images without manual cropping or rotation. Standardize image resolution through resizing.
- Data Augmentation: To address limited data and improve model generalization, apply extensive augmentation techniques including random rotation, flipping, scaling, brightness/contrast adjustments, and elastic deformations [23] [24]. A study on the SMD/MSS dataset successfully expanded 1,000 images to 6,035 samples via augmentation [24].
- Data Splitting: Partition data into training, validation, and test sets using an 80/10/10 or 70/15/15 split. Ensure stratification to maintain class distribution across splits. Crucially, perform splitting at the patient level to prevent data leakage.
3.2 Model Training and Hyperparameter Optimization
- Hyperparameter Tuning: Conduct an extensive search for optimal learning rates, optimization algorithms (e.g., Adam, SGD), batch sizes, and data augmentation scales [23].
- Training Regime: For ViTs, leverage transfer learning by pre-training on large-scale image datasets like ImageNet. Follow this with supervised fine-tuning on the target sperm morphology dataset.
- Comparative Analysis: Train and evaluate multiple architectures under identical conditions, including pure CNNs (e.g., ResNet), hybrid CNN-Transformer models, and pure ViTs (e.g., BEiT, DeiT) [23].
3.3 Model Validation and Interpretation
- Evaluation Metrics: Report standard metrics including accuracy, precision, recall, F1-score, and area under the receiver operating characteristic curve (AUC-ROC).
- Statistical Testing: Perform statistical significance tests (e.g., paired t-test) to confirm that performance improvements are not due to chance [23].
- Model Interpretability: Employ visualization techniques such as Attention Maps and Grad-CAM to understand the model's decision-making process. This validates that the model focuses on clinically relevant morphological features (e.g., head shape, vacuoles, tail structure) rather than artifacts [23].
Visualization of Analytical Workflows
The following diagrams, generated using Graphviz, illustrate the core logical workflows and architectural concepts in AI-based sperm morphology analysis.
Diagram 1: AI Sperm Analysis Pipeline. This workflow integrates object detection (YOLO) for localization, CNNs for local feature extraction, and Vision Transformers for global context modeling.
Diagram 2: CNN vs. ViT Architectural Focus. This diagram contrasts the local feature extraction hierarchy of CNNs with the global context modeling of Vision Transformers via self-attention.
The Scientist's Toolkit: Essential Research Reagents and Materials
The development and validation of AI models for sperm morphology analysis rely on a foundation of high-quality data and computational resources.
Table 3: Essential Research Reagents and Materials for AI-Driven Sperm Analysis
| Item Name | Type | Function in Research | Example / Specification |
|---|---|---|---|
| Annotated Sperm Datasets | Data | Training and benchmarking AI models. | HuSHeM [23], SMIDS [23], SVIA [2], SMD/MSS [24] |
| High-Throughput Microscopy | Equipment | Acquiring high-resolution, consistent sperm images for model input. | MMC CASA System [24] |
| Staining Reagents | Wet Lab | Enhancing contrast and visualizing morphological details (acrosome, nucleus). | Diff-Quik, Papanicolaou stain [2] |
| GPU Computing Cluster | Hardware | Accelerating model training and hyperparameter optimization. | NVIDIA GPUs (e.g., A100, V100) |
| Deep Learning Frameworks | Software | Implementing, training, and deploying CNN and Transformer models. | TensorFlow, PyTorch, Hugging Face Transformers |
| Data Augmentation Tools | Software | Artificially expanding dataset size and diversity to improve model robustness. | Albumentations, Torchvision Transforms [23] [24] |
The integration of AI into sperm morphology analysis marks a significant leap toward standardizing and improving the diagnosis of male infertility. While CNNs and architectures like ResNet have laid a strong foundation, providing robust and interpretable results, emerging Vision Transformers have demonstrated a clear potential for superior performance by capturing a more holistic view of sperm morphology. The state-of-the-art accuracy achieved by models like BEiT_Base underscores a meaningful advance in diagnostic capability [23].
The future trajectory of this field points toward multi-modal AI systems that integrate morphology analysis with other semen parameters (e.g., motility) and patient clinical data to provide a comprehensive fertility assessment [26]. Furthermore, the establishment of larger, more diverse, and meticulously annotated public datasets will be crucial for developing models that generalize across different populations and clinical settings [2]. As these technologies mature, the focus will shift to seamless integration into clinical workflows, requiring rigorous validation through prospective trials and addressing challenges related to model transparency, ethical implementation, and regulatory approval [27]. For researchers and drug development professionals, mastering these core architectures is no longer a niche skill but a fundamental component of driving innovation in reproductive medicine.
The integration of artificial intelligence into the field of andrology is transforming the fundamental approach to sperm morphology analysis, a critical component in diagnosing male factor infertility. Traditional manual semen analysis, while foundational, is plagued by substantial inter-observer variability and subjectivity, hindering its reproducibility and diagnostic power [28] [2]. This technical guide examines the pivotal pipeline of data acquisition and annotation, which serves as the bedrock for developing robust AI models. By exploring the transition from conventional stained smears to the dynamic analysis of unstained live sperm, we frame this technical workflow within the broader thesis that high-quality, standardized data is the essential prerequisite for AI to revolutionize male fertility assessment, enabling more objective, efficient, and predictive diagnostics [29] [6].
Data Acquisition Methodologies
The process of acquiring sperm images for AI model training is a critical first step, with the chosen methodology directly influencing the type and quality of morphological data that can be extracted.
Analysis of Stained Sperm Smears
Stained smears represent the traditional and most established method for detailed morphological assessment. This process involves creating a thin film of semen on a glass slide, which is then fixed and stained to enhance the contrast of cellular structures under a microscope.
Key Staining and Imaging Protocols:
- Slide Preparation and Staining: Samples are prepared using stains like Papanicolaou, Diff-Quik, or Spermac to differentially color sperm components [30]. The specific protocol, including fixation time and staining duration, must be rigorously standardized to minimize technical variation.
- Image Capture: Imaging is typically performed using high-resolution brightfield microscopes. For AI model training, it is crucial to capture images at a standardized magnification (commonly 100x oil immersion) and with consistent lighting conditions to ensure uniformity across the dataset [2].
Analysis of Unstained Live Sperm
The analysis of unstained, motile sperm represents a significant advancement, moving from static morphology to dynamic assessment without the potential artifacts introduced by chemical staining.
Key Live Imaging Protocols:
- Motility and Morphology Coupling: This approach utilizes advanced microscopy techniques, such as phase-contrast or differential interference contrast (DIC) microscopy, to visualize live sperm without fixation or staining. This allows for the simultaneous analysis of sperm morphology and motility parameters from video data [28] [6].
- Video Data Acquisition: Instead of static images, short video clips are captured. This requires high-frame-rate cameras and stable environmental chambers to maintain sperm viability during imaging. The resulting videos are decomposed into frames, which are then used for annotation and model training [6].
Table 1: Comparison of Data Acquisition Modalities for Sperm Morphology Analysis
| Feature | Stained Smears | Unstained Live Sperm |
|---|---|---|
| Sample State | Fixed, non-viable | Live, motile |
| Primary Imaging | Brightfield microscopy | Phase-contrast, DIC microscopy |
| Data Output | High-contrast static images | Video sequences (frame stacks) |
| Key Advantage | Detailed structural clarity | Combined motility & morphology analysis |
| Main Limitation | Potential staining artifacts | Lower contrast for some defects |
| AI Application | Classification of head, midpiece, and tail defects | Dynamic selection and holistic health assessment |
Data Annotation and Dataset Curation
The creation of high-quality, annotated datasets is the most significant bottleneck and critical success factor in developing accurate AI models for sperm morphology analysis [2].
Annotation Standards and Criteria
Annotation must adhere to strict, internationally recognized criteria to ensure biological relevance and model generalizability.
- WHO 6th Edition Standards: The current gold standard defines a morphologically normal sperm as having a smooth, oval head with a well-defined acrosome covering 40-70% of the head area, a midpiece that is slender and aligned with the head axis, and a tail of uniform caliber that is approximately ten times the head length without sharp angulations [30]. Any deviation from these criteria in the head, neck, midpiece, or tail is considered abnormal.
- Strict Kruger Criteria: This is a more rigorous application of the WHO standards, where borderline forms are classified as abnormal. It is frequently used in the context of assisted reproductive technologies (ART) [31].
The Annotation Workflow and Its Challenges
The practical workflow for creating a labeled dataset involves several complex steps fraught with challenges.
- Expert-Dependent Labeling: Annotation must be performed by trained embryologists or andrologists. This process is time-consuming and suffers from inherent inter- and intra-observer variability, which can introduce noise into the training labels [28] [2].
- Handling Complex Scenarios: A significant challenge in annotation includes correctly identifying and labeling sperm that are clumped together, partially overlapping, or only partially visible at the image borders [2]. Furthermore, the presence of non-sperm cells and debris in the semen sample must be correctly identified and excluded from the sperm morphology dataset.
Publicly Available Datasets
Several research groups have created and made public datasets to foster innovation in the field. The table below summarizes some key examples.
Table 2: Key Public Datasets for Sperm Morphology AI Research
| Dataset Name | Key Characteristics | Content and Annotations | Primary Use Case |
|---|---|---|---|
| VISEM-Tracking [2] | Video dataset of motile sperm | 125,000 annotated instances for detection; 26,000 segmentation masks | Sperm tracking and motility analysis |
| SVIA Dataset [2] | Comprehensive video and image collection | Object detection, segmentation masks, and cropped images for classification | Multi-task model development (detection, classification) |
| MHSMA [2] | Focus on stained morphology | 1,540 sperm images with features like acrosome and vacuoles | Sperm head classification |
Sperm Data Annotation Workflow
Integration with AI Model Development
The curated and annotated datasets form the foundation upon which machine learning and deep learning models are built to automate sperm morphology analysis.
From Conventional ML to Deep Learning
The evolution of AI in this field mirrors trends in other areas of computer vision.
- Conventional Machine Learning: Early approaches relied on handcrafted features. For sperm head analysis, this included shape-based descriptors (e.g., Hu moments, Zernike moments, Fourier descriptors), texture, and grayscale features [2]. These features were then fed into classifiers like Support Vector Machines (SVM) or decision trees. While achieving accuracies up to 90% for head classification, these models were limited by their dependence on manual feature engineering and often failed to generalize across different datasets [2].
- Deep Learning (DL) Models: Modern systems employ DL architectures, such as Convolutional Neural Networks (CNNs), which automatically learn hierarchical features directly from the raw pixel data [29] [6]. This approach is more powerful and can be trained to perform not only classification but also object detection (locating all sperm in an image) and semantic segmentation (precisely outlining the head, midpiece, and tail of each sperm) [2]. One study using a CNN for sperm motility analysis reported a strong correlation with manual analysis (r=0.90, p<0.001) [28].
Table 3: Performance of Different AI Models in Sperm Analysis
| Algorithm/Model | Task | Reported Performance | Key Strengths/Limitations |
|---|---|---|---|
| Support Vector Machine (SVM) [2] | Sperm head classification | AUC-ROC: 88.59%, Precision: >90% | Good with handcrafted features, limited generalization |
| Bayesian Density Estimation [2] | Sperm head classification | Accuracy: 90% | Effective for specific morphological categories |
| Fourier Descriptor + SVM [2] | Non-normal head classification | Accuracy: 49% | Highlights variability and challenge of some tasks |
| Artificial Neural Network (ANN) [28] | Sperm concentration prediction | Accuracy: 93%, Sensitivity: 95.45% | Good for parameter prediction from spectral data |
| Convolutional Neural Network (CNN) [28] | Sperm motility prediction | Correlation with manual: r=0.90 | Automates feature extraction from raw video/image data |
End-to-End AI Pipeline
The integration of data and models creates a comprehensive automated system. The workflow begins with the input of either a stained image or a live video. A detection model first localizes all individual sperm cells. For live videos, a tracking algorithm links sperm across frames to analyze motility. Each detected sperm is then passed to a segmentation model that delineates its key morphological components—head, midpiece, and tail. Finally, these segmented regions are processed by a classification model that identifies specific defects based on the learned annotation criteria, outputting a detailed morphological report [2] [6].
AI Morphology Analysis Pipeline
The Scientist's Toolkit: Essential Research Reagents and Materials
The following table details key reagents, materials, and tools essential for conducting research in AI-based sperm morphology analysis.
Table 4: Essential Research Reagent Solutions and Materials
| Item | Function/Application |
|---|---|
| Sperm Staining Kits (e.g., Papanicolaou, Diff-Quik) | Provides differential staining for detailed structural analysis of fixed sperm on smears [30]. |
| Makler Counting Chamber | A specialized chamber for standardized manual semen analysis and validation of automated systems [32]. |
| Phase-Contrast Microscope | Enables high-contrast imaging of unstained, live sperm for motility and concurrent morphology studies [28]. |
| Computer-Assisted Semen Analysis (CASA) System | A commercial automated system for semen analysis; serves as a benchmark and data source for AI model development [28] [6]. |
| Public Datasets (e.g., VISEM, SVIA) | Provides pre-collected, annotated image and video data for training and validating new AI algorithms [2]. |
| Deep Learning Frameworks (e.g., TensorFlow, PyTorch) | Software libraries used to build, train, and deploy convolutional neural networks for image analysis [29] [6]. |
The integration of artificial intelligence into reproductive medicine has revolutionized the assessment of sperm morphology, a critical parameter in male fertility diagnosis. Traditional manual analysis is inherently subjective, time-consuming, and suffers from significant inter-observer variability, with reported disagreement rates of up to 40% between expert embryologists [2] [33]. This technical guide delineates the comprehensive analysis pipeline for AI-driven sperm morphology assessment, framing it within the broader thesis that automated, objective, and precise evaluation is paramount for advancing fertility research and treatment outcomes. The pipeline transforms raw microscopic images into quantifiable morphological features through a sequence of sophisticated computational stages, enabling the detection of subtle patterns indistinguishable to the human eye. By examining current methodologies, performance metrics, and experimental protocols, this review provides researchers, scientists, and drug development professionals with a foundational understanding of the technical workflow that underpins modern computer-aided sperm analysis (CASA) systems and their application in clinical and research settings [6].
Image Acquisition and Pre-processing
Image Acquisition Modalities
The initial stage of the AI analysis pipeline involves acquiring high-quality sperm images, with the chosen modality significantly impacting subsequent processing stages. Current research utilizes both stained and unstained sperm imaging, each presenting distinct advantages and challenges. Stained images, typically using Diff-Quik or other Romanowsky stain variants, provide enhanced contrast that facilitates the distinction of sperm structures [34]. However, staining procedures render sperm non-viable for further clinical use and may introduce morphological alterations [1] [34]. Consequently, there is a growing research focus on analyzing unstained, live sperm, which preserves cell viability for use in assisted reproductive technology (ART) procedures like intracytoplasmic sperm injection (ICSI) [1]. Unstained analysis necessitates more advanced imaging and processing techniques due to lower signal-to-noise ratios and indistinct structural boundaries [34].
Advanced microscopy techniques are employed to overcome these limitations. Confocal laser scanning microscopy at 40x magnification has been used to create novel datasets of live sperm, capturing Z-stack images at 0.5 μm intervals to generate high-resolution, three-dimensional structural information [1]. Super-resolution techniques, including Structured Illumination Microscopy (SIM) and Airyscan, achieve resolutions of approximately 100-140 nm, enabling detailed visualization of nanoscopic structures critical for accurate morphological assessment [35]. These technological advancements provide the high-fidelity input data required for robust AI model development.
Image Pre-processing Techniques
Image pre-processing is crucial for enhancing image quality and preparing data for subsequent analysis stages. Deep learning-based methods have increasingly supplanted traditional techniques for tasks such as denoising, deblurring, and resolution enhancement [36]. Traditional spatial domain operations include high-pass and low-pass filters, median filters for noise reduction, and deconvolution algorithms like Richardson-Lucy for image restoration [36]. Transfer domain methods, such as Fourier and Wavelet transforms, are also employed for noise reduction and edge detection [36].
Deep learning architectures have demonstrated superior performance in microscopy image enhancement. As summarized in Table 1, various network architectures have been applied to critical pre-processing tasks. For super-resolution, Generative Adversarial Networks (GANs) like Real-ESRGAN and IIM-GAN have achieved Peak Signal-to-Noise Ratio (PSNR) values of 37.84 and Structural Similarity Index (SSIM) of 0.99 [36]. U-Net architectures have been widely adopted for image restoration and denoising tasks, with DnCNN achieving PSNR of 37.01 for denoising [36]. These enhanced images provide cleaner inputs for subsequent detection and segmentation stages, significantly improving overall pipeline performance.
Table 1: Deep Learning Models for Microscopy Image Enhancement
| Network | Year | Task | Architecture | Key Results |
|---|---|---|---|---|
| IIM-GAN | 2021 | Super-resolution | GAN | PSNR=37.84, SSIM=0.99 |
| Real-ESRGAN | 2023 | Super-resolution | GAN | - |
| SF-SIM | 2022 | Super-resolution | CNN + Attention | PSNR=31.19, SSIM=0.732 |
| U-Net | 2020 | Super-resolution | U-Net | PSNR=20.32, SSIM=0.40 |
| RedrawNet | 2023 | Restoration | U-Net | Accuracy: 0.9086 |
| DnCNN | 2022 | Denoising | TL | PSNR=37.01, SSIM=0.924 |
| BoostNET | 2021 | Denoising | DCNN | PSNR=35.62, SSIM=0.9129 |
| IRUNET | 2021 | Denoising | Encoder/Decoder | PSNR=38.38, SSIM=0.98 |
Figure 1: Image Pre-processing Workflow
Sperm Detection and Tracking
Deep Learning-Based Detection Models
Sperm detection constitutes the foundational step of identifying and localizing individual sperm within microscopic images or video sequences. Traditional computer vision approaches relied on handcrafted features and classical algorithms like K-means clustering for sperm head detection [2] [18]. However, contemporary research has shifted toward deep learning-based object detection models that automatically learn discriminative features from data.
The YOLO (You Only Look Once) family of architectures has demonstrated remarkable efficacy in real-time sperm detection. Recent research has introduced DP-YOLOv8n, a specialized deep sperm recognition model that incorporates the GSConv module, SE attention mechanism, and an additional small target detection layer to improve detection accuracy and real-time performance [37]. On the VISEM-1 dataset, this model achieved a mean Average Precision ([email protected]) of 86.8%, representing a 3.4% improvement over the baseline YOLOv8n, while maintaining a detection speed of 38.875 frames per second [37]. This balance between accuracy and speed is crucial for clinical applications requiring high-throughput analysis.
Other advanced architectures include Mask R-CNN, which has shown superior performance in segmenting smaller and more regular sperm structures like heads and nuclei [34]. The evolution from traditional machine learning to deep learning represents a paradigm shift in sperm detection capabilities, enabling more robust performance across varying image qualities and sperm densities.
Multi-Sperm Tracking Algorithms
Sperm motility analysis requires robust multi-object tracking to monitor individual sperm movement across video frames—a challenging task due to frequent occlusions, collisions, and high sperm density in samples. Traditional tracking algorithms like the Joint Probabilistic Data Association Filter (JPDAF) and Multiple Hypothesis Tracker (MHT) struggle with real-time performance due to their computational complexity [37].
The Interacting Multiple Model (IMM) architecture represents a significant advancement in sperm tracking technology. Recent research has proposed IMM-ByteTrack, which integrates Singer and Constant Turn (CT) motion models to better capture the complex movement patterns of motile sperm [37]. This algorithm combines dynamic model switching with interactive filtering mechanisms to improve tracking accuracy in challenging clinical scenarios featuring overlap and occlusion. On benchmark datasets VISEM-1 and LCH-SD, IMM-ByteTrack achieved Multiple Object Tracking Accuracy (MOTA) metrics of 70.51% and 75.13% respectively, outperforming baseline algorithms by 2.95% and 4.03% [37].
Table 2: Performance Comparison of Sperm Detection and Tracking Algorithms
| Algorithm | Task | Dataset | Key Metric | Performance |
|---|---|---|---|---|
| DP-YOLOv8n | Detection | VISEM-1 | [email protected] | 86.8% |
| DP-YOLOv8n | Detection | VISEM-1 | FPS | 38.875 |
| IMM-ByteTrack | Tracking | VISEM-1 | MOTA | 70.51% |
| IMM-ByteTrack | Tracking | LCH-SD | MOTA | 75.13% |
| YOLOv8n (Baseline) | Detection | VISEM-1 | [email protected] | 83.4% |
Figure 2: Sperm Detection and Tracking Pipeline
Sperm Segmentation
Multi-Part Segmentation Architectures
Segmentation represents a critical phase in the analysis pipeline, partitioning detected sperm into distinct morphological components—head, acrosome, nucleus, neck, and tail—for detailed structural analysis. Accurate segmentation is prerequisite for precise morphological characterization and abnormality detection. Recent research has systematically evaluated multiple deep learning architectures for this task, with each demonstrating distinct advantages for different sperm components [34].
Mask R-CNN, a two-stage instance segmentation architecture, has shown exceptional performance in segmenting smaller and more regular structures like sperm heads, nuclei, and acrosomes [34]. Its region proposal network effectively localizes these components before detailed mask prediction, yielding high precision for well-defined structures. For morphologically complex components like sperm tails, U-Net achieves superior performance, leveraging its encoder-decoder structure with skip connections to capture multi-scale contextual information essential for segmenting elongated, thin structures [34]. Single-stage detectors like YOLOv8 and YOLO11 have also demonstrated competitive performance, particularly for the neck region, offering an optimal balance between accuracy and computational efficiency [34].
Comparative Performance Analysis
Quantitative evaluation of segmentation models employs multiple metrics, including Intersection over Union (IoU), Dice coefficient, Precision, Recall, and F1 Score. Table 3 summarizes the comparative performance of various architectures across different sperm components, based on evaluation using live, unstained human sperm datasets [34].
The segmentation performance varies significantly across sperm components, reflecting their distinct morphological challenges. Smaller, well-defined structures like heads and nuclei generally achieve higher IoU scores (≥0.85 with Mask R-CNN), while complex structures like tails present greater challenges, with U-Net achieving the highest performance (IoU: 0.82) [34]. This component-specific performance variation underscores the potential advantage of ensemble approaches that leverage multiple architectures optimized for different morphological structures.
Table 3: Segmentation Performance Across Sperm Components and Models
| Sperm Component | Best Model | IoU | Dice | Precision | Recall | F1 Score |
|---|---|---|---|---|---|---|
| Head | Mask R-CNN | 0.87 | 0.93 | 0.94 | 0.92 | 0.93 |
| Acrosome | Mask R-CNN | 0.85 | 0.92 | 0.93 | 0.91 | 0.92 |
| Nucleus | Mask R-CNN | 0.86 | 0.92 | 0.93 | 0.91 | 0.92 |
| Neck | YOLOv8 | 0.83 | 0.90 | 0.91 | 0.89 | 0.90 |
| Tail | U-Net | 0.82 | 0.90 | 0.91 | 0.89 | 0.90 |
Feature Extraction and Classification
Deep Feature Engineering
Following segmentation, feature extraction transforms the partitioned sperm components into quantifiable descriptors that capture clinically relevant morphological characteristics. Traditional machine learning approaches relied on handcrafted features such as shape descriptors (Hu moments, Zernike moments, Fourier descriptors), texture features, and grayscale statistics [2] [18]. However, these manual feature engineering approaches often failed to capture the subtle morphological patterns indicative of sperm quality.
Deep feature engineering represents a paradigm shift, combining the representational power of deep neural networks with classical feature selection techniques. Recent research has proposed a hybrid architecture integrating ResNet50 with Convolutional Block Attention Module (CBAM), enhanced by a comprehensive feature engineering pipeline [38] [33]. This framework extracts features from multiple layers (CBAM, Global Average Pooling, Global Max Pooling, pre-final) and combines them with feature selection methods including Principal Component Analysis (PCA), Chi-square test, Random Forest importance, and variance thresholding [33]. This approach achieved exceptional test accuracies of 96.08% ± 1.2% on the SMIDS dataset and 96.77% ± 0.8% on the HuSHeM dataset, representing improvements of 8.08% and 10.41% respectively over baseline CNN performance [38] [33].
The attention mechanisms in CBAM enable the network to focus on semantically salient regions like head shape, acrosome integrity, and tail structure, while suppressing irrelevant background information [33]. This targeted feature extraction significantly enhances the discriminative power of the resulting feature representations for subsequent classification tasks.
Classification Methodologies
The final pipeline stage utilizes the extracted features for sperm morphology classification. While end-to-end deep learning approaches directly output classification results, hybrid approaches that decouple feature extraction and classification often achieve superior performance. Support Vector Machines (SVM) with Radial Basis Function (RBF) kernels have demonstrated particular efficacy when applied to deep feature embeddings, effectively mapping the features to morphological classes [33].
The optimal reported configuration (GAP + PCA + SVM RBF) significantly outperformed recent Vision Transformer and ensemble methods [33]. This hybrid leverage strategy combines the representational power of deep networks with the classification efficiency of traditional machine learning algorithms. Classification typically follows WHO guidelines, categorizing sperm into normal and abnormal morphological classes, with further subclassification of abnormalities based on affected components (head, neck, tail) [1] [2].
Figure 3: Feature Extraction and Classification Workflow
Experimental Protocols and Research Reagents
Detailed Methodologies for Key Experiments
Standardized experimental protocols are essential for reproducible sperm morphology analysis research. A representative protocol for developing an AI model for unstained live sperm assessment involves specific methodologies [1]:
Sample Collection and Preparation: Semen samples are collected from healthy volunteers (typically aged 18-40) following 2-7 days of sexual abstinence. Samples are collected via masturbation into sterile containers, with liquefaction checked within 30 minutes of ejaculation. Specimens are maintained at 37°C during analysis, and each sample is divided into three aliquots for comparative analysis [1].
Image Acquisition Protocol: For unstained live sperm imaging, a 6 μL droplet is dispensed onto a standard two-chamber slide with 20 μm depth. Images are captured using confocal laser scanning microscopy at 40x magnification in confocal mode (LSM, Z-stack). The Z-stack interval is typically set at 0.5 μm, covering a total range of 2 μm. Each image capture uses a frame time of approximately 633.03 ms with an image size of 512 × 512 pixels, corresponding to 159.7 × 159.7 μm per slide [1].
Dataset Annotation: Embryologists and researchers manually annotate well-focused sperm images using bounding boxes in programs like LabelImg. Annotation quality is ensured through inter-observer correlation metrics, with target correlation coefficients of 0.95 for normal sperm morphology detection and 1.0 for abnormal morphology detection [1].
AI Model Training: The ResNet50 transfer learning model is trained using a dataset of annotated sperm images. A typical training regimen might utilize 9,000 images (4,500 normal, 4,500 abnormal) derived from 32 pattern samples, with training conducted over 150 epochs. Performance is evaluated on a separate test set of 900 batches of previously unseen images [1].
Research Reagent Solutions
Table 4: Essential Research Reagents and Materials for Sperm Morphology Analysis
| Reagent/Material | Function/Application | Specifications |
|---|---|---|
| Diff-Quik Stain | Sperm staining for conventional morphology analysis | Romanowsky stain variant for fixed sperm |
| Leja Slides | Sample preparation for microscopy | Standard two-chamber slides, 20 μm depth |
| MemBright Probes | Membrane staining for enhanced visualization | Lipophilic fluorescent dyes for live/fixed samples |
| Fluorescent Phalloidin | Actin staining for spine morphology | Binds F-actin in fixed samples |
| Wheat Germ Agglutinin | Membrane staining | Lectin binding to surface carbohydrates |
| Antibody Markers | Specific protein detection (e.g., tubulin) | Immunofluorescence for structural components |
The integrated pipeline for AI-driven sperm morphology analysis—encompassing image pre-processing, detection, segmentation, and feature extraction—represents a transformative advancement in male fertility assessment. This technical overview demonstrates how contemporary deep learning architectures have surpassed traditional methods in accuracy, efficiency, and clinical utility. The emergence of standardized, high-quality datasets, coupled with sophisticated models like CBAM-enhanced ResNet50 for classification and Mask R-CNN/U-Net for segmentation, has enabled unprecedented precision in morphological evaluation. Nevertheless, challenges persist in model interpretability, generalizability across diverse populations, and seamless integration into clinical workflows. Future research directions should focus on developing more explainable AI systems, establishing robust validation frameworks across multiple clinical sites, and creating standardized benchmarking datasets. As these computational methodologies continue to evolve, they hold significant promise for delivering objective, reproducible, and clinically actionable sperm morphology assessments that can enhance diagnostic accuracy and personalize treatment strategies in reproductive medicine.
Sperm morphology, the study of the size and shape of sperm, is a cornerstone of male fertility assessment [3]. The analysis is critical because the sperm's physical structure directly influences its ability to penetrate and fertilize an oocyte [39]. Traditional manual assessment, however, is inherently subjective, labor-intensive, and prone to significant inter-technologist variability, making it a challenging parameter to standardize [40] [2] [41]. Artificial intelligence (AI), particularly deep learning, is poised to revolutionize this field by introducing automation, standardization, and enhanced accuracy to sperm morphology evaluation [40] [6] [2]. This technical guide details the classification of sperm defects and explores how AI research is overcoming the limitations of conventional analysis, thereby providing researchers and drug development professionals with a framework for advanced diagnostic and therapeutic development.
Comprehensive Classification of Morphological Defects
Sperm morphology is systematically categorized based on the anatomical region of the defect. The following sections and corresponding tables provide a detailed breakdown of anomalies affecting the sperm head, midpiece, and tail, synthesized from clinical and research classifications including the modified David classification and WHO criteria [40] [42] [39].
Head Anomalies
The sperm head contains the genetic material and enzymes necessary for egg penetration, making its integrity crucial for fertilization. Head defects are the most prevalent type of morphological abnormality [42]. These anomalies can indicate disrupted spermatogenesis, genetic traits, or external factors such as increased testicular temperature or exposure to toxic chemicals [39].
Table 1: Classification and Functional Implications of Sperm Head Anomalies
| Anomaly Type | Key Morphological Description | Reported Prevalence | Functional Implications & Associated Factors |
|---|---|---|---|
| Macrocephaly [40] [39] | Giant head, often containing extra chromosomes [39]. | A specific subtype of head defect [40]. | Linked to homozygous mutation of the aurora kinase C gene; impaired fertilization potential [39]. |
| Microcephaly [40] [39] | Smaller than normal head; also called small-head sperm [39]. | A specific subtype of head defect [40]. | Often associated with a defective acrosome or reduced genetic material [39]. |
| Pinhead [39] | Head appears as a pin with minimal to no paternal DNA. | A variation of microcephaly [39]. | May indicate a diabetic condition [39]. |
| Tapered Head [40] [39] | "Cigar-shaped" or elongated head [40] [39]. | One of 7 classified head defects [40]. | Suggests varicocele or scrotal heat exposure; often contains abnormal chromatin/DNA packaging and aneuploidy [39]. |
| Thin/Narrow Head [40] [39] | Extreme variation of the tapered head [39]. | One of 7 classified head defects [40]. | Associated with broken DNA, varicocele, or disrupted head formation [39]. |
| Globozoospermia [39] | Round-headed sperm with an absent acrosome. | A distinct abnormality [39]. | Missing enzymes to penetrate the egg; inability to activate the egg post-fertilization [39]. |
| Abnormal Acrosome [40] | Malformed or missing acrosomal cap. | One of 7 classified head defects [40]. | Directly impairs the sperm's ability to digest and penetrate the egg's outer layers [40] [39]. |
| Nuclear Vacuoles [39] | Two or more large vacuoles or multiple small vacuoles in the sperm head. | Visible under high magnification [39]. | Studies conflict on fertilization potential; some show low potential, others show no effect [39]. |
| Multiple Heads [40] [39] | Sperm with two or more heads [40]. | One of 7 classified head defects [40]. | Linked to exposure to toxic chemicals, heavy metals, smoke, or high prolactin hormone [39]. |
Midpiece Defects
The midpiece, or neck, houses the mitochondria that provide energy for sperm motility. Defects in this region are primarily associated with impairments in sperm movement and energy metabolism [42] [39].
Table 2: Classification and Functional Implications of Sperm Midpiece Defects
| Defect Type | Key Morphological Description | Reported Prevalence | Functional Implications & Associated Factors |
|---|---|---|---|
| Bent Neck [40] [42] | Sharp angular bend at the sperm neck [40]. | A specific midpiece defect [40] [42]. | Strongly associated with impairments in progressive and rapid progressive motility [42]. |
| Cytoplasmic Droplet [40] [39] | A persistent droplet of cytoplasm located along the midpiece. | A specific midpiece defect [40]. | Indicates immature sperm; may be related to defective mitochondria or missing/broken centrioles [39]. |
| Large Swollen Midpiece [39] | Abnormally thick and swollen neck region. | A noted midpiece defect [39]. | Suggests defective mitochondria or issues with the centrioles, which guide chromosome movement [39]. |
Tail Abnormalities
The tail is essential for propulsion. Abnormalities in tail structure directly compromise motility, preventing sperm from navigating the female reproductive tract to reach the oocyte [42] [39].
Table 3: Classification and Functional Implications of Sperm Tail Abnormalities
| Abnormality Type | Key Morphological Description | Reported Prevalence | Functional Implications & Associated Factors |
|---|---|---|---|
| Coiled Tail [40] [39] | Tail is coiled upon itself [40]. | A specific tail defect [40] [42]. | Sperm cannot swim; linked to incorrect seminal fluid conditions, bacterial presence, or heavy smoking [39]. |
| Short Tail [40] [39] | Abnormally short tail, also known as stump tail [40]. | A specific tail defect [40] [42]. | Very low or no motility; caused by Dysplasia of the Fibrous Sheath (DFS), an autosomal recessive genetic disease; associated with chronic respiratory disease and a higher rate of sperm aneuploidy [39]. |
| Multiple Tails [40] [39] | Presence of two or more tails [40]. | A specific tail defect [40]. | Similar to multiple heads, associated with exposure to toxins [39]. |
| Bent Tail [39] | A crooked or angled tail. | A noted tail abnormality [39]. | Impedes straight, progressive movement. |
| Tail-less (Acaudate) [39] | Complete absence of a tail. | A noted tail abnormality [39]. | Sperm is immotile; often seen during cellular necrosis [39]. |
Experimental Protocols for AI-Based Morphology Analysis
The development of robust AI models for sperm morphology analysis relies on rigorous experimental protocols encompassing data curation, model training, and validation. The following workflow details a representative methodology from a recent deep learning study.
Diagram 1: Experimental workflow for AI-based sperm morphology analysis, based on the SMD/MSS study [40].
Data Acquisition and Annotation Protocol
A critical first step is the creation of a high-quality, annotated dataset [40] [2].
- Sample Preparation and Inclusion Criteria: In a prospective study design, semen samples are obtained from patients after informed consent. To ensure image quality and avoid overlap, samples with a sperm concentration of at least 5 million/mL are typically included, while those with very high concentrations (>200 million/mL) are excluded. Smears are prepared according to WHO manual guidelines and stained using a standardized kit (e.g., RAL Diagnostics) [40].
- Image Acquisition: Images of individual spermatozoa are captured using a Computer-Assisted Semen Analysis (CASA) system, such as the MMC CASA system. Acquisition is performed using bright-field microscopy with a high-resolution camera and an oil immersion 100x objective to ensure sufficient detail for morphological assessment [40].
- Expert Annotation and Ground Truth Establishment: Each sperm image is independently classified by three experienced experts to establish a reliable ground truth. The classification should follow a standardized system, such as the modified David classification, which defines 12 classes of defects across the head, midpiece, and tail [40]. An Excel spreadsheet or dedicated database is used to compile the annotations from all experts, along with morphometric data (e.g., head length, tail length). The final ground truth file includes the image name, expert classifications, and detailed annotations for sperm with associated anomalies [40].
Data Pre-processing and Augmentation Protocol
Raw sperm images require pre-processing to be suitable for AI model training [40].
- Image Pre-processing: This step aims to denoise images and standardize their format. Techniques include handling missing values or outliers, and normalization. A common approach is to resize all images to a uniform dimension (e.g., 80x80 pixels) and convert them to grayscale to reduce computational complexity [40].
- Data Augmentation: To address the common problem of limited and imbalanced datasets, augmentation techniques are employed to artificially expand the dataset and balance the representation of different morphological classes. This can include random rotations, flips, and color variations. For example, an initial dataset of 1,000 images can be augmented to over 6,000 images, significantly improving model robustness [40].
AI Model Training and Evaluation Protocol
This phase involves building and validating the predictive model [40].
- Dataset Partitioning: The entire augmented dataset is randomly partitioned into two subsets: a training set (typically 80% of the data) used to teach the model, and a testing set (the remaining 20%) reserved for final evaluation on unseen data. A portion of the training set may also be used for validation during training [40].
- Model Training: A Convolutional Neural Network (CNN) architecture is implemented in a programming environment like Python 3.8. The CNN learns hierarchical features directly from the pre-processed sperm images. The model is trained on the training subset, with its parameters iteratively adjusted to minimize classification error [40].
- Model Evaluation: The trained model's performance is quantitatively evaluated using the held-out test set. Key metrics include classification accuracy, which has been reported in recent studies to range from 55% to 92%, reflecting the varying complexity of different morphological classes and the degree of expert agreement [40].
Deep Learning Architectures for Defect Classification
AI research in sperm morphology has evolved from conventional machine learning to deep learning models, with the latter demonstrating superior performance by automatically learning relevant features from raw image data.
From Conventional Machine Learning to Deep Learning
Conventional machine learning (ML) approaches, such as Support Vector Machines (SVM), K-means clustering, and decision trees, have been applied to sperm morphology analysis [2]. These models typically rely on a two-stage pipeline: first, manual extraction of image features (e.g., shape descriptors like Hu moments, Zernike moments, Fourier descriptors, texture, and grayscale intensity), and second, feeding these handcrafted features into a classifier [2]. While these methods achieved accuracies as high as 90% for specific tasks like head shape classification, they are fundamentally limited. They are cumbersome, time-consuming, and their performance is highly dependent on the quality of the manual feature engineering, often leading to poor generalization on new datasets [2]. A significant shortcoming is that most conventional studies focus only on the sperm head, failing to provide a complete structural analysis of the midpiece and tail [2].
Convolutional Neural Networks (CNNs) for End-to-End Learning
Deep learning, specifically Convolutional Neural Networks (CNNs), has emerged as the state-of-the-art solution, enabling end-to-end learning from raw pixels to classification output [40] [2].
Diagram 2: Simplified architecture of a CNN for sperm morphology classification [40].
- Model Architecture and Workflow: A typical CNN for sperm morphology classification begins with an input layer that receives pre-processed grayscale images. This is followed by a series of convolutional and pooling layers that act as hierarchical feature extractors, automatically learning to detect edges, shapes, and complex patterns specific to different sperm defects. The final feature maps are then flattened and passed through one or more fully connected (dense) layers, which perform the final classification into the predefined morphological classes (e.g., the 12 classes of the modified David classification) [40].
- Impact and Performance: This approach has demonstrated "satisfactory" to "promising" results, with accuracy figures competitive with expert judgment [40]. The key advantage is the system's ability to standardize and accelerate semen analysis, reducing reliance on subjective human assessment and enabling high-throughput evaluation [40] [6]. Furthermore, AI systems like the STAR (Sperm Tracking and Recovery) platform have shown clinical utility by successfully identifying viable sperm in severely oligospermic samples, leading to successful pregnancies where traditional methods failed [13].
The Scientist's Toolkit: Essential Research Reagents and Materials
The following table catalogs key reagents, technologies, and computational tools essential for conducting research in AI-based sperm morphology analysis.
Table 4: Key Reagents and Solutions for AI-Driven Sperm Morphology Research
| Tool/Reagent | Specification/Example | Primary Function in Research |
|---|---|---|
| Staining Kits | RAL Diagnostics staining kit [40]. | Provides contrast for microscopic visualization of sperm structures (head, midpiece, tail) for image acquisition. |
| CASA Systems | MMC CASA system [40]. | Automated platform for acquiring and storing high-resolution digital images of individual spermatozoa from smears. |
| Reference Datasets | SMD/MSS Dataset [40], SVIA Dataset [2]. | Provides large volumes of annotated sperm images for training, validating, and benchmarking deep learning models. |
| Programming Environments | Python 3.8 [40]. | Core programming language for implementing deep learning algorithms, data pre-processing, and analysis pipelines. |
| Deep Learning Frameworks | TensorFlow, PyTorch (inferred from context). | Provides libraries and tools for building, training, and deploying convolutional neural network (CNN) models. |
| Data Augmentation Tools | Integrated Python libraries (e.g., TensorFlow's ImageDataGenerator) [40]. | Algorithmically generates variations of original images (rotations, flips) to expand and balance training datasets. |
| High-Performance Computing | GPUs (Graphics Processing Units) [6]. | Accelerates the computationally intensive process of training complex deep learning models on large image datasets. |
The precise classification of sperm morphological defects into head, midpiece, and tail abnormalities provides a critical foundation for understanding male infertility. The integration of AI, particularly through deep learning models, is transforming this field from a subjective, manual exercise into an objective, automated, and data-driven science. While challenges remain—including the need for larger, more diverse datasets and the resolution of the "black-box" nature of complex algorithms—the trajectory is clear [6] [2]. AI-powered morphology analysis is poised to enhance diagnostic accuracy, personalize fertility treatments, and improve success rates in assisted reproduction, ultimately offering new hope to couples worldwide [40] [13] [41]. For researchers and drug developers, these advancements open new avenues for creating sophisticated diagnostic tools and targeted therapeutic interventions aimed at the underlying causes of defective spermatogenesis.
The analysis of sperm quality is a cornerstone of male fertility assessment, with sperm DNA fragmentation (SDF) representing a crucial parameter beyond conventional morphology. Elevated SDF levels are strongly associated with reduced fertilization rates, impaired embryo development, and increased miscarriage rates. Traditionally, assessing DNA fragmentation requires specialized, invasive assays that compromise sample viability.
This technical guide explores an emerging paradigm: the use of artificial intelligence (AI) to predict DNA fragmentation status directly from non-invasive, label-free phase-contrast images. This approach is framed within the broader thesis of how AI is revolutionizing sperm morphology analysis by moving beyond static, human-visible features to decode subtle, sub-visual biomarkers correlated with cellular function and integrity. By leveraging deep learning, researchers can extract patterns from phase-contrast images that are imperceptible to the human eye, potentially related to changes in refractive index and cellular density that accompany nuclear damage [43]. This methodology promises to transform diagnostic workflows in reproductive medicine and drug development by enabling high-throughput, non-destructive SDF screening.
Technical Foundations
Phase-Contrast Microscopy in Biology
Phase-contrast microscopy is a contrast-enhancing optical technique that allows for the visualization of transparent and colorless specimens, such as living cells, without the need for killing, fixing, or staining [44]. It works by translating small changes in the phase of light, caused by interactions with the specimen, into corresponding changes in amplitude (brightness), which are then seen as differences in image contrast [45].
Key Advantages for Live Cell Analysis:
- Non-destructive Observation: Enables the study of living cells in their natural state, preserving biological activity [44].
- Real-time Monitoring: Facilitates the observation of dynamic biological processes [44].
- Label-free: Avoids the use of fluorescent dyes or stains, which can be cytotoxic, expensive, and introduce experimental complexity [44].
AI and Deep Learning in Reproductive Medicine
Artificial intelligence, particularly deep learning (a subset of machine learning), has experienced rapid growth in its application to reproductive medicine. Deep learning models can automatically learn hierarchical features from large, complex datasets, such as medical images, and use these patterns to make predictions or classifications [46]. In the context of sperm analysis, AI is being applied to tasks such as sperm selection, embryo selection, and morphology analysis, with the goal of improving objectivity, standardization, and success rates of assisted reproductive technologies (ART) [29] [46] [14].
Conventional machine learning models for sperm morphology analysis often relied on manually engineered features (e.g., shape, texture) and showed limited performance, particularly in segmenting complete sperm structures and generalizing across datasets [2]. Deep learning models overcome these limitations by automatically learning relevant features directly from the image data, leading to substantial improvements in the efficiency and accuracy of sperm morphology analysis [29] [2].
Methodological Framework: An AI Workflow for Predicting DNA Fragmentation
The following section outlines a detailed experimental protocol for developing an AI model to classify sperm DNA fragmentation status using phase-contrast images, based on methodologies demonstrated in analogous cell studies [43].
Experimental Workflow
The end-to-end process, from sample preparation to model prediction, is visualized in the following workflow diagram.
Detailed Experimental Protocols
1. Sample Preparation and Induction of DNA Fragmentation:
- Cell Model: While the referenced study used K562 cells (a human leukemia cell line) [43], the protocol can be adapted for human sperm cells. Fresh semen samples are collected and processed using standard density gradient centrifugation or swim-up techniques to isolate motile, morphologically normal sperm.
- Induction of DNA Fragmentation (Optional): To create a robust training dataset with a wide range of SDF levels, researchers may optionally induce DNA fragmentation in a subset of the sample. This can be achieved through methods such as exposure to hydrogen peroxide, irradiation, or cryopreservation-induced stress. This ensures the model is trained on both normal and high SDF examples.
2. Parallel Staining and Image Acquisition:
- This is a critical step for generating the "ground truth" data needed to supervise the AI model.
- Fluorescence Staining: Process the sperm samples to detect key biomarkers of apoptosis and DNA fragmentation. As per the protocol by Kikuchi et al. [43], this includes:
- Caspase Activity Staining: Use a fluorescently-labeled caspase inhibitor (e.g., FITC-DEVD-FMK) to identify cells undergoing apoptosis.
- DNA Fragmentation Staining: Use a terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) assay or similar (e.g., propidium iodide) to directly label fragmented DNA.
- Microscopy: For the same field of view, acquire two sets of images:
- Phase-Contrast Images: First, capture high-resolution phase-contrast images of the unstained, living sperm cells.
- Fluorescence Images: Immediately afterward, without moving the sample, capture the corresponding fluorescence images showing the caspase activity and DNA fragmentation signals.
3. AI Model Training and Validation:
- Dataset Curation and Labeling: Use image registration software to align the phase-contrast images with their corresponding fluorescence images. Based on the fluorescence signals, assign each cell in the phase-contrast image to a class. A typical classification schema is shown in the table below [43].
- Model Selection and Training: A common and effective approach is to use a pre-trained convolutional neural network (CNN) like ResNet50, a model with 50 layers known for its performance in image classification [43]. The model is trained by feeding it the phase-contrast images (input) and the corresponding class labels derived from fluorescence (output). The model learns to associate subtle visual features in the phase-contrast images with the DNA fragmentation status.
- Validation: Employ a robust validation method such as five-fold cross-validation [43]. The dataset is split into five parts. The model is trained on four parts and validated on the fifth. This process is repeated five times, with each part used exactly once as the validation data. This provides a reliable estimate of the model's performance and guards against overfitting.
Table 1: Cell Classification Schema Based on Fluorescence Staining
| Class | Caspase Activity | DNA Fragmentation | Interpretation |
|---|---|---|---|
| Class 1 | Negative | Negative | Viable cell, intact DNA |
| Class 2 | Positive | Negative | Early apoptosis, DNA largely intact |
| Class 3 | Positive | Positive | Late apoptosis, significant DNA fragmentation |
Quantitative Data and Model Performance
The following tables summarize hypothetical quantitative data and performance metrics based on the established methodology [43]. In the cited study, AI models successfully classified cells into three apoptosis-related groups using only phase-contrast images.
Table 2: Example AI Model Performance Metrics (5-Fold Cross-Validation)
| AI Model | Accuracy (%) | Precision (%) | Recall (%) | F-Score |
|---|---|---|---|---|
| ResNet50 (Server-based) | 94.5 | 95.1 | 93.8 | 0.944 |
| Lobe | 91.2 | 92.3 | 90.5 | 0.914 |
Interpretation of Metrics:
- Accuracy: The overall proportion of correct predictions.
- Precision: The proportion of cells predicted as a specific class (e.g., high SDF) that actually belong to that class. High precision means fewer false positives.
- Recall (Sensitivity): The proportion of cells that actually belong to a specific class that were correctly predicted. High recall means fewer false negatives.
- F-Score: The harmonic mean of precision and recall, providing a single metric that balances both.
Table 3: Research Reagent Solutions for DNA Fragmentation Analysis
| Reagent / Material | Function / Application | Experimental Role |
|---|---|---|
| FITC-DEVD-FMK | Fluorescent inhibitor of caspase activity | Serves as ground truth for detecting early apoptotic events in cells [43]. |
| TUNEL Assay Kit | Fluorescently labels fragmented DNA | Provides the definitive ground truth measurement for DNA fragmentation [43]. |
| Phase Contrast Microscope | High-resolution imaging of unstained cells | Generates the input data (images) for the AI model [44]. |
| Fluorescence Microscope | Detection of specific fluorescent signals | Used to acquire ground truth labels for model training [43]. |
| ResNet50 Model | Deep convolutional neural network architecture | The AI engine that learns to map phase-contrast features to fragmentation classes [43]. |
Discussion and Future Directions
The ability to predict DNA fragmentation from phase-contrast images represents a significant leap forward. The underlying hypothesis is that the biochemical and structural alterations in the sperm nucleus during DNA fragmentation induce subtle, sub-resolution changes in the cell's refractive index and mass-density distribution. These changes, while invisible to a human observer, are captured as complex patterns in the phase-contrast image and can be decoded by a sufficiently powerful deep learning model [43].
Integration with Broader AI Trends in Sperm Morphology
This specific application is a powerful example of a broader trend in AI-driven sperm morphology analysis, which is evolving from simple classification of head shape towards holistic, functional assessment. The field is moving beyond conventional machine learning, which was limited by manual feature extraction and often focused only on the sperm head [2]. Deep learning enables the segmentation and analysis of the complete sperm structure (head, neck, and tail) and the discovery of novel, non-intuitive biomarkers of health and function [29] [2].
Challenges and Clinical Translation
Despite its promise, several challenges remain. A major hurdle is the lack of large, standardized, and high-quality annotated datasets required to train robust and generalizable models [29] [2] [27]. Furthermore, the "black box" nature of some AI systems can limit clinical trust and adoption. Key barriers to adoption in clinical practice include high implementation costs and a lack of training for embryologists [14]. Finally, rigorous external validation through large-scale, multi-center randomized controlled trials is needed to prove that AI predictions truly improve clinical outcomes, such as live birth rates [46] [27].
Future directions will likely involve the integration of multi-modal data (e.g., combining phase-contrast images with motility parameters from time-lapse imaging) and the development of more transparent, explainable AI systems. As these technologies mature, they hold the potential to become an indispensable tool in the reproductive clinic and drug discovery pipeline, enabling non-invasive, high-throughput, and highly accurate assessment of sperm quality.
Navigating Development Challenges: Data, Generalization, and Computational Efficiency
The application of artificial intelligence (AI) in male fertility research, particularly in sperm morphology analysis, represents a paradigm shift in diagnostic precision and standardization. However, the performance of these AI models is fundamentally constrained by the quality and scale of the annotated datasets used for their training. Manual sperm morphology assessment is recognized as a challenging parameter to standardize due to its subjective nature, often reliant on the operator's expertise [40]. This variability in manual analysis creates a critical bottleneck in developing robust AI systems that can achieve clinical-grade reliability. The inherent complexity of sperm morphology, characterized by numerous possible defects across the head, midpiece, and tail, necessitates exceptionally well-annotated datasets to train models effectively [2]. The "black-box" nature of many complex deep learning algorithms further underscores the necessity for meticulously curated training data, as model decisions must be traceable to biologically grounded features [6]. This technical guide outlines comprehensive strategies for creating high-quality, annotated datasets specifically tailored for AI-based sperm morphology research, addressing the fundamental data challenges that currently limit widespread clinical adoption.
The Sperm Morphology Analysis Challenge
Sperm morphology analysis (SMA) is a crucial laboratory test in male fertility assessment, where clinicians evaluate sperm quality by analyzing the proportion of abnormal morphology in a fixed number of sperms (typically over 200) and identifying specific types of defects [2]. According to classification standards established by the World Health Organization (WHO), sperm morphology is divided into the head, neck, and tail, with 26 types of abnormal morphology recognized [2]. The manual assessment process faces significant reproducibility challenges due to its reliance on human expertise and subjective interpretation.
Table 1: Key Challenges in Manual Sperm Morphology Analysis
| Challenge Category | Specific Limitations | Impact on AI Development |
|---|---|---|
| Subjectivity & Standardization | High inter- and intra-observer variability; difficult to teach and standardize [40] [2] | Creates inconsistent ground truth for model training |
| Workload Intensity | Requires analysis of >200 sperm per sample; substantial manual effort [2] | Limits the scale of datasets that can be feasibly annotated |
| Morphological Complexity | 26 types of abnormalities across head, midpiece, and tail compartments [2] | Demands fine-grained annotation schema with expert knowledge |
| Image Quality Issues | Sperm may appear intertwined, partially displayed, or with overlapping debris [2] | Complicates automated segmentation and classification |
Existing Computer-Assisted Semen Analysis (CASA) systems only partially address these challenges due to their limited ability to accurately distinguish between spermatozoa and cellular debris and to classify midpiece and tail abnormalities [40]. The limited quality of captured microscopic images often leads to unsatisfactory results, creating an urgent need for more sophisticated AI solutions built upon superior data foundations [40].
Foundational Principles for Dataset Creation
Defining Clear Annotation Objectives
The initial phase of dataset creation requires precise definition of annotation objectives aligned with the clinical and research goals. For sperm morphology analysis, this involves determining the appropriate classification system (e.g., WHO, Kruger, or David's modified classification) and the granularity of defect categorization [40] [2]. Each annotation task must be designed to capture biologically relevant features that contribute to diagnostic validity. Establishing these objectives upfront guides all subsequent decisions regarding data collection, annotation taxonomy, and quality assurance protocols.
Ensuring Data Diversity and Representativeness
A critical consideration in dataset construction is ensuring comprehensive diversity and representativeness to prevent model bias and enhance generalizability. The dataset should encompass variations across multiple dimensions:
- Patient Demographics: Age, ethnicity, and geographical factors
- Clinical Conditions: Samples from fertile and infertile individuals, various pathological conditions
- Technical Variations: Different staining methods (e.g., RAL Diagnostics staining kit [40]), microscope settings, and laboratory protocols
- Morphological Spectrum: Balanced representation of normal sperm and all categories of abnormalities [47]
A diverse dataset ensures that trained models can perform robustly across various clinical settings and population groups, rather than excelling only on data that mirrors the specific characteristics of the training set.
Strategic Framework for Data Collection & Annotation
Data Acquisition and Sample Preparation
The data acquisition process requires meticulous attention to technical consistency and biological relevance. A standardized protocol for sperm smear preparation, staining, and image capture must be established and rigorously followed [40] [2]. In the SMD/MSS (Sperm Morphology Dataset/Medical School of Sfax) dataset development, researchers included samples with a sperm concentration of at least 5 million/mL while excluding samples with high concentrations (>200 million/mL) to avoid image overlap and facilitate the capture of whole sperm [40]. On average, 37 ± 5 images were captured per sample, depending on the density and distribution of spermatozoa on the smear [40]. The MMC CASA system was employed for image acquisition using bright field mode with an oil immersion 100x objective [40]. Each image contained a single spermatozoon, comprising a head, a midpiece, and a tail, which is essential for precise morphological assessment [40].
Annotation Taxonomy and Labeling Schema
Establishing a comprehensive annotation taxonomy is fundamental for creating clinically relevant datasets for sperm morphology analysis. The modified David classification, which includes 12 classes of morphological defects, provides a structured framework for categorization [40]:
- 7 head defects: tapered, thin, microcephalous, macrocephalous, multiple, abnormal post-acrosomal region, abnormal acrosome
- 2 midpiece defects: cytoplasmic droplet, bent
- 3 tail defects: coiled, short, multiple
Each spermatozoon should be independently classified by multiple experts with extensive experience in semen analysis [40]. An Excel spreadsheet or specialized database should be created to document various morphological classes for each part of the spermatozoon, maintaining consistent labeling conventions across the entire dataset [40].
Table 2: Annotation Approaches for Sperm Morphology Analysis
| Annotation Type | Use Case in Sperm Analysis | Technical Requirements | Advantages/Limitations |
|---|---|---|---|
| Classification | Categorizing entire sperm images as normal/abnormal [47] | Whole-image labels; categorical taxonomy | Fast but provides limited morphological detail |
| Object Detection | Locating and classifying sperm parts (head, midpiece, tail) [48] | Bounding boxes around each component | Balances speed with structural information |
| Instance Segmentation | Precise pixel-level masking of sperm structures [2] | Polygon annotations defining exact boundaries | Maximum detail but computationally intensive |
| Keypoint Annotation | Marking specific landmarks (acrosome, neck junction) [48] | Coordinate points on critical features | Useful for structural alignment and measurement |
Expert Consensus and Quality Assurance
The complexity of sperm morphology classification necessitates a rigorous framework for expert consensus and quality assurance. Research indicates that inter-expert agreement distribution must be systematically analyzed to establish reliable ground truth [40]. Three agreement scenarios should be documented:
- No Agreement (NA): No consensus among experts
- Partial Agreement (PA): 2/3 experts agree on the same label for at least one category
- Total Agreement (TA): 3/3 experts agree on the same label for all categories [40]
Statistical analysis using Fisher's exact test can evaluate differences between experts in each morphology class, with significance considered at p < 0.05 [40]. This systematic approach to quantifying expert agreement helps identify ambiguous classification categories that may require refined annotation guidelines.
Technical Protocols for Dataset Enhancement
Data Preprocessing and Cleaning
Image preprocessing is essential for enhancing signal quality and standardizing inputs for AI model training. The primary steps include:
- Denoising: Removing artifacts from insufficient lighting or poorly stained semen smears [40]
- Normalization/Standardization: Resizing images with linear interpolation strategy to standardized dimensions (e.g., 80801 grayscale) [40]
- Data Cleaning: Identifying and handling missing values, outliers, or inconsistencies that might hinder model performance [40]
These preprocessing steps ensure that the AI model is not influenced by technical variations unrelated to the biological features of interest, thereby improving generalization capability and reducing confounding factors.
Data Augmentation Techniques
Data augmentation represents a crucial strategy for addressing the common challenge of limited dataset size in medical AI applications. By artificially expanding dataset diversity without collecting new images, augmentation techniques help prevent overfitting and improve model robustness [40] [47]. In the SMD/MSS dataset development, the initial collection of 1,000 images was extended to 6,035 after applying data augmentation techniques [40]. Common augmentation methods include rotation, scaling, flipping, and adding noise [47]. For sperm morphology analysis, it is essential that augmentation techniques preserve the biological validity of morphological features, as arbitrary transformations might create implausible sperm structures that could mislead the model during training.
Dataset Partitioning Strategy
Proper dataset partitioning is critical for rigorous model evaluation and preventing data leakage. The standard approach involves:
- Splitting the entire set of images randomly into subsets
- Allocating 80% of the dataset for model training
- Reserving 20% for testing model performance on unseen data [40]
- Further dividing the training subset to extract a validation set (typically 20% of the training data) for hyperparameter tuning [40]
This partitioning strategy ensures that model performance is evaluated on completely independent data, providing a more accurate assessment of real-world applicability and generalization capability.
Quality Control and Validation Framework
Annotation Consistency Metrics
Maintaining annotation consistency is paramount for dataset quality. Inter-annotator agreement (IAA) serves as a key metric for measuring consistency between different annotators [49]. High IAA indicates that annotators understand the guidelines and are aligned on how to apply them to the data [49]. However, for subjective tasks like sperm morphology assessment, perfect agreement may not always be achievable, and lower IAA can provide important signals about task difficulty or ambiguous guidelines [49]. Regular quality checks should be implemented through both automatic metrics and manual review processes to maintain high standards throughout the annotation lifecycle [48].
Continuous Quality Improvement
Quality control should be implemented as an ongoing process rather than a single final check. Two complementary approaches should be employed:
- Concurrent QC: Addressing issues as annotation occurs or in the next round, allowing for real-time guideline updates and annotator retraining [49]
- Post-annotation QC: Understanding the upper bound for model quality by thoroughly verifying annotation accuracy after completion [49]
Establishing a feedback loop where quality metrics directly inform guideline refinement is essential for continuous improvement. This iterative process helps identify systematic errors, clarify ambiguous cases, and enhance overall dataset reliability [49].
Experimental Protocols in Sperm Morphology AI Research
Dataset Creation Methodology
A representative experimental protocol for creating an annotated sperm morphology dataset follows these methodical steps:
Sample Preparation and Inclusion Criteria
- Collect semen samples from patients after obtaining informed consent
- Implement inclusion criteria: sperm concentration ≥5 million/mL
- Apply exclusion criteria: samples with high concentrations (>200 million/mL) to avoid image overlap [40]
- Prepare smears following WHO manual guidelines using standardized staining (e.g., RAL Diagnostics staining kit) [40]
Image Acquisition and Preprocessing
- Use MMC CASA system for image acquisition
- Employ bright field mode with oil immersion 100x objective
- Capture 37 ± 5 images per sample depending on density and distribution
- Ensure each image contains a single spermatozoon with head, midpiece, and tail [40]
- Apply image denoising and normalization to standardized dimensions
Expert Annotation and Consensus Building
- Engage three experts with extensive experience in semen analysis
- Conduct independent classification using modified David classification
- Document classifications in a structured database with consistent labeling
- Analyze inter-expert agreement using statistical methods (Fisher's exact test) [40]
- Resolve discrepancies through consensus meetings or adjudication
Model Training and Evaluation Protocol
Data Partitioning and Augmentation
- Randomly split dataset: 80% training, 20% testing
- Extract validation set from training subset for hyperparameter tuning
- Apply data augmentation techniques (rotation, scaling, flipping) to expand dataset diversity [40]
Algorithm Development and Training
- Implement convolutional neural network (CNN) architecture in Python
- Utilize five-stage pipeline: image preprocessing, database partitioning, data augmentation, program training, and evaluation [40]
- Employ transfer learning where appropriate to leverage pre-trained models
Performance Validation
- Apply K-Fold cross-validation scheme to maintain objectivity
- Report mean absolute error (MAE) for continuous outcomes [50]
- Calculate accuracy, precision, recall, and F-value for classification tasks [40]
- Compare model performance against expert annotations and established benchmarks
Visualization of Dataset Creation Workflow
Diagram 1: Sperm Morphology Dataset Creation Workflow
Essential Research Reagents and Materials
Table 3: Essential Research Reagents for Sperm Morphology Dataset Creation
| Reagent/Material | Specification | Function in Dataset Creation |
|---|---|---|
| Staining Kit | RAL Diagnostics staining kit [40] | Provides contrast for morphological feature visualization |
| Microscope System | MMC CASA system with digital camera [40] | Image acquisition from sperm smears |
| Microscope Objective | Oil immersion 100x objective [40] | High-resolution imaging of sperm structures |
| Annotation Software | Label Studio, CVAT, LabelImg [48] | Streamlined labeling with customizable workflows |
| Data Augmentation Tools | Python libraries (e.g., TensorFlow, PyTorch) [40] | Artificial expansion of dataset diversity |
| Statistical Analysis Software | IBM SPSS Statistics [40] | Inter-expert agreement analysis and validation |
The creation of high-quality, annotated datasets represents the foundational bottleneck in advancing AI applications for sperm morphology analysis. Addressing this challenge requires methodical approaches to data collection, expert-driven annotation, rigorous quality control, and strategic dataset enhancement. The strategies outlined in this technical guide provide a comprehensive framework for developing datasets that can support robust, clinically relevant AI models. As these datasets grow in scale and quality, they will enable increasingly sophisticated AI systems capable of transforming male fertility diagnostics through enhanced objectivity, reproducibility, and predictive accuracy. Future efforts should focus on collaborative initiatives to create large, diverse, and publicly available datasets that can accelerate innovation across the research community while maintaining the highest standards of annotation quality and biological validity.
Data Augmentation Techniques to Overcome Limited Samples and Class Imbalance
The integration of Artificial Intelligence (AI) into reproductive medicine represents a paradigm shift, offering the potential to automate and standardize diagnostic procedures that have long relied on subjective human assessment [46] [51]. A critical application lies in sperm morphology analysis, a fundamental yet challenging component of male fertility evaluation. Traditional manual analysis is slow, suffers from significant inter-observer variability, and creates bottlenecks in clinical workflows [24] [2]. While deep learning models, particularly Convolutional Neural Networks (CNNs), have demonstrated remarkable proficiency in image-based tasks, their performance is intrinsically linked to the volume and quality of training data [46] [2]. In sperm morphology analysis, researchers consistently face the dual challenges of limited sample sizes and class imbalance, where images of rare abnormal morphologies are vastly outnumbered by normal samples or other common defect types [52]. This technical guide explores how data augmentation serves as a pivotal strategy to overcome these hurdles, thereby enhancing the robustness, accuracy, and generalizability of AI models for sperm morphology analysis and accelerating their translation into clinical practice.
The Data Challenge in AI-Based Sperm Morphology Analysis
The development of robust AI models for sperm morphology analysis is fundamentally constrained by data-related challenges. The cornerstone of effective deep learning is the availability of large, well-annotated, and diverse datasets; however, this requirement is often at odds with the realities of clinical andrological research.
Limited Dataset Size and Annotation Complexity
The process of creating high-quality datasets for sperm morphology is arduous. Specimen collection and preparation must adhere to strict protocols, and expert annotation is both time-consuming and costly. Embryologists and researchers must manually label individual spermatozoa in images, often dealing with complexities such as sperm appearing intertwined or only partial structures being visible [2]. The annotation task itself is particularly challenging, as it requires simultaneous evaluation of head, vacuoles, midpiece, and tail abnormalities based on standardized criteria like those from the World Health Organization [2]. Consequently, many initial datasets are small. For instance, the SMD/MSS dataset began with only 1,000 individual spermatozoa images before augmentation [24], and other public datasets like the Modified Human Sperm Morphology Analysis Dataset (MHSMA) contain 1,540 images of sperm heads [1]. Such limited data volume is insufficient for training complex deep learning models from scratch, leading to overfitting and poor generalization to new data.
Class Imbalance in Morphological Categories
A more insidious problem is that of class imbalance. In a typical semen sample, the vast majority of spermatozoa exhibit abnormal morphology, but these abnormalities are distributed across a wide spectrum of defect types—tapered heads, coiled tails, broken necks, etc. [2]. Consequently, when categorizing sperm into specific morphological classes (e.g., normal, tapered, pyriform, amorphous), some classes become "minority classes" with very few representative samples, while others are over-represented [52]. Most conventional classifiers are biased toward the majority classes because they aim to maximize overall accuracy. This leads to poor classification performance on the minority classes, which are often clinically significant [52]. A classifier might, for instance, achieve high accuracy by simply classifying all sperm as the most common abnormal type, thereby failing to identify rare but critical morphological defects.
Core Data Augmentation Techniques and Their Performance
Data augmentation encompasses a series of techniques that generate high-quality artificial data by manipulating existing data samples [53]. By artificially enlarging and diversifying the training dataset, these techniques help models perform better on scarce or imbalanced datasets, substantially enhancing their generalization capabilities [53]. The following sections and tables summarize the key techniques and their quantified impact in biomedical image analysis, with a focus on sperm morphology.
Table 1: Fundamental Data Augmentation Techniques for Image Data
| Technique Category | Description | Key Parameters | Primary Benefit |
|---|---|---|---|
| Geometric Transformations | Alters the spatial orientation of the image. | Rotation angle, flip axis, scale ratio. | Introduces translation, rotation, and scale invariance. |
| Photometric Transformations | Alters the pixel intensity and color values. | Brightness delta, contrast range, noise variance. | Builds robustness to lighting and staining variations. |
| Mixing Methods | Blends multiple images and their labels. | Mixing ratio (α), cut region size. | Smooths decision boundaries and improves regularization. |
| Generative Methods | Generates entirely new synthetic images. | Network architecture, latent vector size. | Creates samples for rare/absent classes; addresses severe imbalance. |
The application of these techniques in real-world studies has yielded significant performance gains. In a seminal study on deep learning for sperm morphology, researchers extended their initial dataset of 1,000 images to 6,035 images by applying data augmentation techniques, which was crucial for training their Convolutional Neural Network (CNN) model. This approach resulted in a final model accuracy ranging from 55% to 92% across different morphological categories [24]. Another recent study developed an AI model for assessing unstained live sperm morphology using a ResNet50 architecture trained on a dataset of 12,683 annotated sperm images. Their model achieved a test accuracy of 93%, with precision and recall for abnormal sperm morphology at 0.95 and 0.91, respectively [1]. These figures underscore the critical role of a sufficiently large and varied training set, often achieved through augmentation, in developing high-performance models.
Table 2: Quantified Impact of Data Augmentation in Model Performance
| Study / Context | Baseline Performance (Without Augmentation) | Performance After Augmentation | Key Augmentation Techniques Used |
|---|---|---|---|
| General Image Recognition [54] | AUC ~83% | AUC ~85% (A/B tests showed 23% accuracy increase in some cases) | Flipping, Rotation, Random Cropping |
| Sperm Morphology Classification [24] | N/A (Initial dataset: 1,000 images) | Accuracy 55-92% (Trained on 6,035 augmented images) | Data augmentation techniques (unspecified) |
| Document Layout Analysis [54] | N/A | 23% drop in processing errors | Elastic Deformation |
| Imbalanced Data Classification [52] | CNN with imbalanced data performed poorly | Proposed method (without augmentation) outperformed CNN with data augmentation | Graph-based transformation (algorithm-level) |
Advanced and Generative Augmentation Strategies
While basic transformations are a good starting point, complex fields like medical imaging often require more sophisticated augmentation strategies to capture the underlying data manifold and address severe class imbalance effectively.
Mix-Based and Generative Methods
For scenarios where basic transformations plateau, mix-based methods such as MixUp and CutMix have proven effective. MixUp creates weighted average combinations of two images and their labels, which helps smooth decision boundaries and improves model calibration [54]. CutMix replaces a random patch of one image with a patch from another, preserving the spatial context and proving particularly beneficial for object detection tasks [54].
When dealing with extreme class imbalance or the need to generate entirely new, realistic samples, generative methods are employed. Generative Adversarial Networks (GANs) have been used in medical imaging to synthesize patches or entire images of rare classes, such as specific sperm morphological defects [54]. More recently, Diffusion Models have emerged as a powerful alternative for high-fidelity medical image generation. One study demonstrated that diffusion models could successfully synthesize medical images of similar styles to the original data but with dramatically varied anatomic details, providing a potential low-cost data augmentation strategy for AI applications [55].
Algorithm-Level Approaches for Imbalance
Alongside data-level solutions, algorithm-level approaches directly modify the learning process to be more robust to class imbalance. One proposed method involves a graph-based transformation that explores the relationships between a given sample and the nearest samples from both minority and majority classes [52]. This technique constructs two individual graphs to preserve the manifold structure of minority and majority classes, providing a dedicated projection matrix for each sample under test. This method has shown superior performance compared to standard CNNs, even when the CNN was supplemented with data augmentation [52].
Experimental Protocols and Research Reagents
Implementing effective data augmentation requires a structured experimental pipeline. The following workflow and toolkit outline a standard approach for a sperm morphology analysis project.
Detailed Experimental Protocol: Sperm Morphology CNN with Augmentation
The following protocol is synthesized from recent studies, notably Abdelkefi et al. (2025) and the in-house AI model development described in PMC (2025) [24] [1].
Data Acquisition:
- Specimen Collection: Collect semen samples from volunteers (e.g., n=30-100) after 2-7 days of sexual abstinence, following ethical guidelines and informed consent [1].
- Image Capture: Use a Computer-Aided Sperm Analysis (CASA) system [24] or a confocal laser scanning microscope (e.g., LSM 800 at 40x magnification) [1] to capture images of individual spermatozoa. For stained sperm analysis, slides are typically fixed and stained with Diff-Quik or similar Romanowsky stain variants [1].
- Initial Dataset: A typical starting dataset may contain 1,000-1,500 raw images, though this number can vary significantly [24] [1].
Expert Annotation & Preprocessing:
- A team of experienced embryologists manually annotates each sperm image using tools like LabelImg [1]. Annotation includes drawing bounding boxes and classifying sperm based on strict morphological criteria (e.g., WHO guidelines) into categories such as "normal," "tapered head," "pyriform head," "amorphous head," and "tail defect" [24] [2].
- Establish inter-observer reliability, with a target correlation coefficient of >0.95 between annotators [1].
- Preprocessing steps include resizing images to a uniform dimension (e.g., 512x512 pixels) and normalizing pixel values.
Data Augmentation Pipeline:
- Apply a series of augmentation techniques on-the-fly during training or pre-generate an augmented dataset. A robust pipeline for sperm images includes:
- Geometric: Random rotation (±15°), horizontal and vertical flips, slight scaling (0.9-1.1x), and translation.
- Photometric: Adjust brightness (±10%), contrast (±10%), and add minor Gaussian noise to simulate imaging variations.
- Advanced/Synthetic: For underrepresented classes, employ MixUp or CutMix. In cases of severe imbalance, train a GAN or Diffusion Model [55] on the minority class examples to generate synthetic samples.
- The goal is to significantly increase the dataset size; for example, expanding from 1,000 to over 6,000 images [24].
- Apply a series of augmentation techniques on-the-fly during training or pre-generate an augmented dataset. A robust pipeline for sperm images includes:
Model Training and Evaluation:
- Model Architecture: Select a deep learning architecture such as a custom CNN or a pre-trained model like ResNet50 using transfer learning [1].
- Training: Split the augmented dataset into training, validation, and test sets (e.g., 70/15/15). Train the model using an optimizer like Adam and a loss function such as cross-entropy.
- Evaluation: Evaluate the model on the held-out test set. Report standard metrics including Accuracy, Precision, Recall, F1-Score, and Area Under the Curve (AUC-ROC) [1] [2]. For imbalanced classes, precision and recall for the minority classes are critical.
Table 3: The Scientist's Toolkit for Sperm Morphology AI Research
| Reagent / Material / Tool | Function / Description | Example in Use |
|---|---|---|
| Confocal Laser Scanning Microscope | High-resolution imaging of unstained, live sperm at low magnification, enabling 3D Z-stack capture. | Capturing sperm images for analysis without rendering them unusable for ART [1]. |
| CASA System | Automated, objective analysis of sperm concentration, motility, and (in advanced systems) morphology. | Provides a standardized platform for initial sperm assessment and image acquisition [24] [51]. |
| Diff-Quik Stain | A variant of Romanowsky stain used to stain sperm on glass slides for detailed morphological examination. | Preparing sperm smears for conventional morphology analysis and creating stained image datasets [1]. |
| LabelImg | An open-source graphical image annotation tool. | Used by embryologists to draw bounding boxes and label sperm images for supervised learning [1]. |
| Albumentations / TorchVision | Python libraries providing a wide range of highly optimized data augmentation operations for images. | Implementing the geometric and photometric augmentation pipeline during model training [54]. |
| Generative Models (GANs, VAEs, Diffusion) | AI models that learn the data distribution of training images and can generate novel, synthetic samples. | Creating artificial images of rare sperm morphological defects to balance the training dataset [56] [55]. |
The integration of data augmentation techniques is not merely an optional step but a fundamental prerequisite for developing robust and clinically viable AI models in sperm morphology analysis. By systematically addressing the critical constraints of limited dataset size and class imbalance through a combination of geometric, photometric, mix-based, and generative methods, researchers can significantly enhance model performance and generalizability. The experimental protocols and toolkit outlined in this guide provide a roadmap for implementing these techniques effectively. As the field progresses, the synergy between advanced generative AI and algorithm-level innovations promises to further overcome data scarcity, ultimately paving the way for AI-driven tools that deliver standardized, objective, and highly accurate sperm morphology assessments in clinical practice.
The analysis of sperm morphology is a cornerstone of male fertility assessment, where the shape, size, and structural integrity of sperm are critically examined. Traditional manual evaluation, however, is plagued by significant inter-observer variability, with reported disagreement rates among experts reaching up to 40% [33] [2]. This subjectivity, combined with the time-intensive nature of the process (typically 30-45 minutes per sample), poses a substantial challenge to standardized diagnosis [33] [38]. Artificial intelligence (AI), particularly deep learning, presents a paradigm shift towards automated, objective, and highly accurate sperm morphology analysis. This technical guide explores the integration of Convolutional Block Attention Module (CBAM) and sophisticated Feature Engineering techniques—a combination demonstrated to achieve state-of-the-art performance, with accuracies exceeding 96% in sperm morphology classification [33] [38]. Framed within broader AI research in reproductive medicine, these methodologies are not merely academic exercises but are pivotal for developing reliable clinical decision-support tools that can enhance outcomes in assisted reproductive technology (ART) [1] [14].
Technical Background
The Convolutional Block Attention Module (CBAM)
CBAM is a lightweight, general-purpose attention module designed for seamless integration into any Convolutional Neural Network (CNN) architecture [57] [58]. Its core function is to enhance the representational power of a network by enabling it to focus on important features across both the channel and spatial dimensions of intermediate feature maps, sequentially [57].
- Channel Attention Module: This component identifies "what" is meaningful in an input image. It squeezes the global spatial information of a feature map into a channel descriptor using both average-pooling and max-pooling operations. The resulting vectors are processed by a shared multi-layer perceptron (MLP) to produce a channel attention map. This map is then multiplied with the input feature map, effectively highlighting the most relevant feature channels [57] [58].
- Spatial Attention Module: Building on the channel-refined features, this module identifies "where" the informative regions are. It applies average-pooling and max-pooling along the channel axis and concatenates the results to form an efficient feature descriptor. A convolutional layer then generates a spatial attention map, which is multiplied with the input features to accentuate key spatial locations [57] [58].
The synergy of these two modules allows CBAM to direct the network's focus toward critical region-specific details in sperm images, such as head shape, acrosome integrity, and tail defects, while suppressing irrelevant background noise [33].
Deep Feature Engineering (DFE)
Deep Feature Engineering represents an advanced machine learning paradigm that marries the strengths of deep learning with classical machine learning. Instead of relying solely on an end-to-end CNN, DFE involves extracting high-dimensional feature representations from intermediate layers of a pre-trained network [33]. These rich features are then subjected to:
- Dimensionality Reduction: Techniques like Principal Component Analysis (PCA) are used to reduce noise and compress the feature space without losing critical information [33] [38].
- Feature Selection: Methods including Chi-square tests, Random Forest importance, and variance thresholding identify and retain the most discriminative features for the classification task [33].
- Shallow Classifiers: The refined feature set is fed into efficient classifiers like Support Vector Machines (SVM) with RBF/Linear kernels or k-Nearest Neighbors (k-NN) for the final prediction [33].
This hybrid approach often yields higher accuracy, improved interpretability, and greater computational efficiency compared to standalone CNNs [33].
Experimental Protocols & Methodologies
A Protocol for Sperm Morphology Classification
The following workflow details the methodology for implementing a CBAM-enhanced, feature-engineered model for sperm morphology classification, as validated in recent literature [33].
Table 1: Key Research Reagent Solutions for Sperm Morphology AI
| Item Name | Function/Description | Example/Specification |
|---|---|---|
| Confocal Laser Scanning Microscope | Captures high-resolution, low-magnification Z-stack images of live, unstained sperm. | LSM 800, 40x magnification, Z-stack interval of 0.5 µm [1]. |
| Standardized Slides | Provides a consistent environment for semen sample preparation and imaging. | Two-chamber slide with a depth of 20 µm (e.g., Leja) [1]. |
| Annotation Software | Allows experts to manually label sperm images for model training and validation. | LabelImg program [1]. |
| Public Benchmark Datasets | Serves as a standardized benchmark for training and evaluating model performance. | SMIDS (3000 images, 3-class) and HuSHeM (216 images, 4-class) [33]. |
| High-Quality Custom Dataset | Provides a large, diverse, and well-annotated dataset for robust model development. | ~21,600 images captured via confocal microscopy, with 12,683 annotated sperm [1]. |
Workflow Steps:
- Data Acquisition & Annotation: Semen samples are prepared on standardized slides, and images are captured using a confocal laser scanning microscope. Embryologists and researchers then manually annotate these images, categorizing sperm into classes (e.g., normal, abnormal head, abnormal tail) based on WHO strict criteria [1] [33]. The high inter-annotator correlation (e.g., 0.95 for normal morphology) ensures label quality [1].
- Model Architecture Selection & CBAM Integration: A robust CNN like ResNet50 is selected as the backbone. The CBAM module is integrated into the network after each convolutional block, allowing the model to adaptively refine feature maps at multiple levels [33].
- Deep Feature Extraction: The CBAM-enhanced network is used as a feature extractor. Features are harvested from multiple layers, including the CBAM attention layers, Global Average Pooling (GAP), and Global Max Pooling (GMP) layers [33].
- Feature Processing and Selection: The extracted high-dimensional features are concatenated. A feature selection method, such as PCA, is applied to reduce dimensionality and mitigate the curse of dimensionality [33] [38].
- Classification: The processed feature vector is fed into a shallow classifier (e.g., SVM with RBF kernel) for the final morphology classification [33].
Diagram 1: Integrated workflow for AI-based sperm morphology analysis.
Validation and Statistical Analysis
Robust validation is critical for clinical applicability. The proposed framework should be evaluated using 5-fold cross-validation on benchmark datasets to ensure reliability [33]. Performance is measured using standard metrics:
- Accuracy, Precision, and Recall
- Statistical significance should be confirmed using tests like McNemar's test (e.g., p < 0.05) to verify that performance improvements over baselines are not due to chance [33] [38].
Quantitative Performance and Comparative Analysis
Extensive experiments demonstrate the superior performance of combining CBAM with deep feature engineering. The table below summarizes key quantitative results from a recent study that implemented this approach [33].
Table 2: Performance Comparison of Sperm Morphology Classification Models
| Model / Approach | Dataset | Accuracy | Precision | Recall | Key Findings |
|---|---|---|---|---|---|
| Baseline CNN | SMIDS | 88.00% | - | - | Baseline performance without enhancements [33]. |
| Proposed Framework (CBAM + DFE) | SMIDS | 96.08% ± 1.2 | - | - | 8.08% improvement over baseline. Best configuration: GAP + PCA + SVM RBF [33]. |
| Proposed Framework (CBAM + DFE) | HuSHeM | 96.77% ± 0.8 | - | - | 10.41% improvement over baseline on a more complex dataset [33]. |
| Conventional ML (SVM with handcrafted features) | - | ~90% (max) | - | - | Performance heavily reliant on manual feature design, limiting generalizability [2]. |
| In-house AI Model (for live sperm) | Custom | - | 0.95 (Abnormal) 0.91 (Normal) | 0.91 (Abnormal) 0.95 (Normal) | Correlated strongly with CASA (r=0.88). Processing time: ~0.0056 s/image [1]. |
The results unequivocally show that the hybrid model achieves state-of-the-art performance, outperforming not only baseline CNNs but also recent advanced architectures like Vision Transformers and ensemble methods [33].
Clinical Impact and Broader Implications in Reproductive Medicine
The integration of advanced AI models into sperm morphology analysis has profound implications for clinical practice and research in reproductive medicine.
- Standardization and Objectivity: AI models minimize the high inter-observer variability inherent in manual analysis, leading to standardized and reproducible diagnoses across different laboratories and technicians [33] [2].
- Dramatic Efficiency Gains: These systems can reduce the analysis time for a sample from 30-45 minutes to less than one minute, freeing up embryologists for higher-value tasks and potentially increasing laboratory throughput [33] [38].
- Analysis of Live, Unstained Sperm: A significant advantage is the ability to analyze unstained, living sperm using models trained on high-resolution confocal microscopy images. This allows for the selection of viable sperm for procedures like Intracytoplasmic Sperm Injection (ICSI) immediately after assessment, without the damaging effects of staining [1].
- Growing Adoption and Future Outlook: Surveys of international fertility specialists show that AI usage in IVF grew from 24.8% in 2022 to 53.2% in 2025 (including regular and occasional use), with embryo and sperm selection being primary applications. Over 80% of clinics are likely to invest in AI within the next five years, signaling strong future adoption [14].
Visualization and Interpretability
For AI models to be trusted in a clinical setting, their decision-making process must be interpretable. Grad-CAM (Gradient-weighted Class Activation Mapping) is a powerful technique that generates visual explanations for CNN-based models [33]. When applied to a CBAM-enhanced model, it produces heatmaps that highlight the precise image regions—such as a misshapen sperm head or a coiled tail—that most influenced the classification decision [33]. This provides clinicians with intuitive visual evidence to support the model's output, fostering trust and facilitating integration into the diagnostic workflow.
Diagram 2: Model interpretability via attention and Grad-CAM.
The confluence of attention mechanisms like CBAM and sophisticated deep feature engineering represents a significant leap forward for AI in sperm morphology analysis and reproductive medicine at large. This technical synergy moves beyond simple automation to create highly accurate, efficient, and interpretable diagnostic tools. By directly addressing the critical limitations of manual analysis—subjectivity, time-consumption, and the inability to safely assess live sperm—this approach paves the way for more objective fertility assessments and improved success rates in assisted reproduction. As the field evolves, the focus must remain on rigorous clinical validation, addressing ethical considerations, and ensuring these powerful technologies integrate seamlessly into clinical workflows to ultimately enhance patient care.
The application of artificial intelligence (AI) in medicine has gained significant momentum, creating new paradigms for diagnosis and treatment personalization. Within this landscape, bio-inspired optimization algorithms represent a class of computational methods that mimic natural processes and behaviors to solve complex optimization problems. These algorithms, including Ant Colony Optimization (ACO), draw inspiration from biological systems such as ant foraging behavior, bird flocking, and evolutionary selection to efficiently navigate large, complex solution spaces. In the specific domain of andrology and reproductive medicine, these algorithms are increasingly being deployed to enhance AI models, particularly for sophisticated analytical tasks like sperm morphology analysis (SMA), where they contribute to more accurate, efficient, and automated diagnostic systems [5] [59].
The integration of AI into male infertility assessment addresses critical challenges in traditional methods. Sperm morphology analysis is a cornerstone of male fertility evaluation, but it has historically been plagued by subjectivity, low reproducibility, and substantial inter-observer variability due to its reliance on manual microscopic examination [2] [6]. The process requires the classification of over 200 sperm cells into head, neck, and tail compartments based on strict World Health Organization (WHO) criteria, encompassing 26 distinct types of abnormalities—a task that is both labor-intensive and prone to human error [2]. Bio-inspired optimization algorithms are playing a pivotal role in tuning the machine learning (ML) and deep learning (DL) models that automate this process, thereby overcoming the limitations of conventional analysis and paving the way for more objective, high-throughput diagnostic tools [29] [6].
The Science of Sperm Morphology Analysis
Clinical Significance and Technical Challenges
Male factors contribute to approximately 50% of infertility cases, making accurate semen analysis a critical component of fertility diagnostics [2]. Sperm morphology is a key parameter in this evaluation, as it provides diagnostic information that not only predicts natural pregnancy outcomes but also offers insights into the functional status of the testis and epididymis [2]. The declining trend in semen quality globally, particularly in parameters like sperm concentration and total count, further underscores the need for precise and reliable assessment methods [29] [2].
The technical challenges in SMA are multifaceted. Conventional manual analysis under microscopy requires simultaneous evaluation of head, vacuole, midpiece, and tail abnormalities, which substantially increases annotation difficulty [2]. Furthermore, sperm may appear intertwined in images, or only partial structures may be visible at the image edges, complicating both image acquisition and subsequent analysis [2]. These factors contribute to the inherent variability of manual assessment, creating a compelling case for automated, AI-driven solutions that can deliver consistent, objective results.
Evolution of Automated Analysis: From CASA to AI
The initial automation of sperm analysis began with Computer-Aided Sperm Analysis (CASA) systems, which have evolved over approximately 40 years through enhancements in imaging devices, computational power, and software algorithms [6]. While foundational CASA concepts for identifying sperm and analyzing motility have remained consistent, their capabilities have expanded significantly. Modern CASA systems now integrate sophisticated AI techniques to evaluate key sperm parameters—motility, morphology, and DNA integrity—offering substantial advantages over manual methods, including enhanced objectivity, improved consistency, and the ability to detect subtle predictive patterns not discernible by human observation [6].
The transition from traditional CASA to AI-enhanced systems represents a paradigm shift in reproductive medicine. By employing a spectrum of techniques, from classic machine learning to deep learning, these advanced systems achieve more accurate, automated, and high-throughput evaluations [6]. This evolution is fueled by the emergence of extensive open datasets and big data analytics, enabling the development of more robust models that can correlate subtle variations in sperm quality with clinical outcomes, thereby facilitating personalized treatment protocols [6].
Core AI Architectures for Sperm Morphology Analysis
From Conventional Machine Learning to Deep Learning
The application of AI in sperm morphology analysis has evolved through distinct technological phases. Conventional machine learning approaches initially demonstrated considerable success in classifying sperm images. These methods typically followed a standardized pipeline where shape-based descriptors and other feature engineering techniques were used for manual extraction of sperm cell features, followed by classification using algorithms such as Support Vector Machines (SVM) or neural networks [2].
Notable examples of conventional ML applications include a Bayesian Density Estimation-based model that achieved 90% accuracy in classifying sperm heads into four morphological categories (normal, tapered, pyriform, and small/amorphous) [2]. Similarly, researchers have employed combinations of Hu moments, Zernike moments, and Fourier descriptors with K-neighbor, Simple Bayes, and decision tree classifiers [2]. While these approaches significantly advanced the field, they faced fundamental limitations due to their non-hierarchical structures and handcrafted features, which often resulted in over-segmentation or under-segmentation issues and reduced generalization capability across different datasets [2].
Table 1: Comparison of Conventional ML vs. Deep Learning for Sperm Morphology Analysis
| Feature | Conventional Machine Learning | Deep Learning |
|---|---|---|
| Feature Extraction | Manual (e.g., shape descriptors, texture) | Automatic (learned from data) |
| Representation Learning | Limited to engineered features | Hierarchical feature learning |
| Data Dependency | Works with smaller datasets | Requires large, annotated datasets |
| Performance | Prone to saturation | State-of-the-art results |
| Computational Complexity | Lower | Higher (requires GPUs) |
| Interpretability | More interpretable | "Black-box" nature |
| Typical Algorithms | SVM, K-means, Decision Trees | CNN, U-Net, YOLO |
The limitations of conventional ML prompted a shift toward deep learning algorithms, which automate the feature extraction process and learn hierarchical representations directly from image data [2]. DL approaches have demonstrated remarkable capabilities in analyzing medical imaging data related to assisted reproductive technologies, exhibiting superior ability to detect critical features in imaging data that signify underlying fertility-related problems [6].
Deep Learning Architectures and Applications
Deep learning architectures, particularly Convolutional Neural Networks (CNNs), have become the cornerstone of modern sperm morphology analysis systems. These networks excel at processing image data through multiple layers that automatically learn to detect increasingly complex features—from edges and textures in early layers to sophisticated morphological patterns in deeper layers [5] [6].
Specific DL implementations in SMA include the use of U-Net architectures for segmentation tasks, which can precisely delineate sperm components (head, neck, tail), and YOLO (You Only Look Once) variants for real-time detection and classification of sperm in images and videos [60] [2]. For instance, the YOLOv5-MS model has been adapted for real-time multi-surveillance pedestrian target detection, showcasing optimization techniques that could be transferred to sperm detection tasks [60]. These architectures have demonstrated the ability to achieve performance comparable to or exceeding human experts in specific morphological classification tasks, with some studies reporting accuracy rates exceeding 90% in distinguishing normal from abnormal sperm [2] [6].
The application of DL in SMA extends beyond basic classification to comprehensive analysis of complete sperm structure. Recent research explores the potential role of segmentation and classification of complete sperm structure based on deep learning algorithms, aiming to simultaneously evaluate head, neck, and tail abnormalities rather than focusing solely on head morphology [29] [2]. This comprehensive approach is crucial for clinical applicability, as it aligns with the WHO standards for sperm morphology assessment.
Bio-Inspired Optimization Algorithms: Theory and Mechanisms
Fundamentals of Ant Colony Optimization
Ant Colony Optimization is a metaheuristic algorithm inspired by the foraging behavior of ant colonies, particularly their ability to find the shortest paths between their nest and food sources [61] [62]. In nature, ants deposit pheromones—chemical substances that attract other ants—along the paths they travel. When faced with multiple paths to a food source, ants tend to prefer routes with stronger pheromone concentrations, creating a positive feedback loop where shorter paths accumulate pheromones faster than longer ones [61] [62].
The computational model of ACO simulates this behavior by using "artificial ants" that construct solutions to optimization problems by moving through a graph representation of the solution space. As these artificial ants traverse the graph, they deposit virtual pheromones on the edges, with the amount of pheromone proportional to the quality of the solution. Subsequent ants are then influenced by these pheromone trails when making decisions about which path to follow, gradually converging toward optimal or near-optimal solutions [61] [62].
The mathematical foundation of ACO involves probability calculations for path selection based on pheromone intensity (τ) and heuristic information (η). The probability of an ant moving from node i to node j is given by:
[ p{ij}^k = \frac{[\tau{ij}]^\alpha \cdot [\eta{ij}]^\beta}{\sum{l \in \text{allowed}} [\tau{il}]^\alpha \cdot [\eta{il}]^\beta} ]
where:
- (\tau_{ij}) represents the pheromone intensity on edge (i,j)
- (\eta_{ij}) represents the heuristic desirability of edge (i,j)
- (\alpha) and (\beta) are parameters that control the relative influence of pheromone versus heuristic information [61]
Enhanced ACO Variants for Complex Optimization
Recent advancements in ACO have led to the development of enhanced variants that address limitations of the basic algorithm, such as slow convergence speed and susceptibility to local optima [61] [63] [62]. These enhancements include:
Dynamic Weight Adjustment: Integrating dynamic weight scheduling strategies that adjust algorithm parameters in real-time based on system status, such as load changes or equipment operating parameters, to enhance search orientation and convergence [61].
Learning-Enhanced ACO (LeACO): Incorporating bandit-based learning for estimating chance constraints under uncertainty and rank-based learning for updating pheromones on edges, which has shown superior performance in integrated planning and scheduling problems [63].
Intelligently Enhanced ACO (IEACO): Implementing multiple improvement strategies including non-uniform initial pheromone distribution, ε-greedy state transition probability, adaptive adjustment of α and β parameters, and multi-objective heuristic functions that consider both target distance and turning angle [62].
These enhanced ACO variants demonstrate significant performance improvements, with studies reporting 20% reduction in average dispatch time and 15% improvement in resource utilization when dealing with large-scale power dispatching problems [61]. Similarly, in mobile robot path planning, IEACO has shown substantial advantages over traditional ACO in terms of path optimization and convergence speed [62].
Integrating Bio-Inspired Optimization with AI Models for Sperm Analysis
Optimization of Model Parameters and Hyperparameters
In the context of sperm morphology analysis, bio-inspired optimization algorithms play a crucial role in tuning the parameters and hyperparameters of AI models. Deep learning architectures for image analysis typically involve numerous configurable parameters, including learning rates, regularization factors, network depth, filter sizes, and activation functions. Manually configuring these parameters is time-consuming and often suboptimal [6] [59].
ACO and other bio-inspired algorithms can systematically explore this high-dimensional parameter space to identify configurations that maximize model performance metrics such as accuracy, precision, and recall. For instance, researchers have applied the Archimedes optimization algorithm with deep learning for breast mass classification in digital mammograms, achieving a maximum accuracy of 96.48% [60]. Similar approaches can be adapted for sperm morphology analysis, where optimization algorithms fine-tune DL model parameters to enhance classification performance.
The integration of optimization algorithms extends beyond parameter tuning to feature selection—identifying the most discriminative features for sperm classification. By reducing feature dimensionality while preserving classification accuracy, these algorithms contribute to more efficient and interpretable models [59].
Architectural Search and Workflow Optimization
Bio-inspired optimization also facilitates neural architecture search (NAS), automatically discovering optimal network architectures for specific sperm analysis tasks. Rather than relying on manually designed networks, NAS approaches use optimization algorithms to explore vast spaces of possible architectures, identifying configurations that balance complexity with performance [59].
Additionally, these algorithms optimize the end-to-end workflow in automated sperm analysis systems. This includes enhancing image preprocessing steps (e.g., segmentation, noise reduction), improving data augmentation strategies, and optimizing post-processing procedures. For example, ACO can be applied to determine optimal threshold values for sperm head segmentation or to optimize the sequence of image processing operations for maximal accuracy and efficiency [2] [6].
Table 2: Applications of Bio-Inspired Optimization in AI-Based Sperm Analysis
| Application Area | Optimization Focus | Impact on System Performance |
|---|---|---|
| Hyperparameter Tuning | Learning rates, batch size, network depth | Improves classification accuracy and training efficiency |
| Feature Selection | Identifying discriminative morphological features | Reduces computational complexity, enhances interpretability |
| Neural Architecture Search | Network connectivity, layer types | Discovers optimal architectures for specific tasks |
| Image Preprocessing | Segmentation parameters, enhancement filters | Improves input quality for downstream analysis |
| Data Augmentation | Selection of transformation strategies | Enhances model generalization and robustness |
| Workflow Scheduling | Processing order, resource allocation | Increases throughput and resource utilization |
Experimental Protocols and Methodologies
Dataset Curation and Preparation
The development of robust AI models for sperm morphology analysis requires carefully curated datasets with high-quality annotations. Several public datasets have been established to support research in this area, including:
- HSMA-DS (Human Sperm Morphology Analysis DataSet): A publicly available dataset for evaluating sperm images [2].
- MHSMA (Modified Human Sperm Morphology Analysis Dataset): Contains 1,540 images of different sperm types with features such as acrosome, head shape, and vacuoles [2].
- VISEM-Tracking: A multimodal dataset for sperm analysis [2].
- SVIA (Sperm Videos and Images Analysis) dataset: Comprises 125,000 annotated instances for object detection, 26,000 segmentation masks, and 125,880 cropped image objects for classification tasks [2].
The dataset preparation process involves several critical steps: semen sample collection, slide preparation, staining (typically using Diff-Quik or Papanicolaou stains), image acquisition using microscopy systems, and expert annotation of sperm components according to WHO guidelines [2] [6]. Annotation requires simultaneous evaluation of head, vacuoles, midpiece, and tail abnormalities, which substantially increases annotation difficulty and necessitates specialized expertise [2].
Model Training and Optimization Workflow
The experimental workflow for developing optimized AI models for sperm morphology analysis follows a systematic process:
Data Preprocessing: Apply noise reduction filters (e.g., median filtering), contrast enhancement, and normalization to improve image quality and consistency [60] [2].
Data Augmentation: Implement transformation strategies including rotation, flipping, scaling, and color adjustments to increase dataset diversity and improve model generalization [2] [6].
Model Architecture Design: Select appropriate base architectures (e.g., CNN, U-Net, YOLO) and adapt them for sperm analysis tasks [2].
Optimization Setup: Configure bio-inspired optimization algorithms (e.g., ACO, PSO) to search for optimal hyperparameters, architectural components, or feature subsets.
Cross-Validation: Implement k-fold cross-validation to ensure robust performance estimation and avoid overfitting.
Model Evaluation: Assess performance using metrics including accuracy, precision, recall, F1-score, and area under the ROC curve (AUC-ROC) [2].
Statistical Validation: Conduct significance testing to verify performance improvements resulting from optimization.
The following DOT script visualizes this integrated experimental workflow:
Performance Metrics and Validation Framework
Rigorous evaluation is essential for validating the effectiveness of bio-inspired optimization in enhancing AI models for sperm morphology analysis. Key performance metrics include:
- Classification Accuracy: Proportion of correctly classified sperm cells (both normal and abnormal) [2].
- Precision and Recall: Particularly important for detecting specific abnormality types where class imbalance may exist [2].
- Area Under ROC Curve (AUC-ROC): Measures the model's ability to distinguish between normal and abnormal sperm, with studies reporting values up to 88.59% for SVM-based classifiers [2].
- Segmentation Quality: Evaluated using Dice coefficient, Jaccard index, or boundary F1 score for sperm component segmentation [2].
- Computational Efficiency: Training time, inference time, and resource utilization, which are critical for clinical deployment [6].
Validation should include comparison against baseline models without optimization, statistical significance testing of performance differences, and clinical validation against expert andrologist assessments to ensure clinical relevance and applicability [2] [6].
Research Reagent Solutions and Computational Tools
The implementation of bio-inspired optimization for AI-enhanced sperm morphology analysis requires both wet laboratory reagents and computational resources. The following table details essential research reagents and their functions in the experimental pipeline:
Table 3: Essential Research Reagents and Computational Tools for AI-Based Sperm Analysis
| Category | Specific Items | Function/Application |
|---|---|---|
| Laboratory Reagents | Diff-Quik stain, Papanicolaou stain, Eosin-Nigrosin | Sperm staining for morphological assessment |
| Phosphate-buffered saline (PBS) | Semen sample dilution and washing | |
| Fixation solutions (e.g., methanol, formaldehyde) | Sample preservation before staining | |
| Imaging Supplies | Microscope slides and coverslips | Sample mounting for microscopy |
| Immersion oil | High-resolution microscopy | |
| Computational Tools | Python, TensorFlow, PyTorch | DL model development framework |
| OpenCV, scikit-image | Image processing and augmentation | |
| DEAP, Optuna | Optimization algorithm implementation | |
| NumPy, Pandas | Data manipulation and analysis | |
| Hardware | High-resolution microscopes with digital cameras | Image acquisition |
| GPU clusters (NVIDIA) | Accelerated model training and inference |
Future Directions and Research Challenges
Emerging Trends and Innovations
The integration of bio-inspired optimization with AI for sperm morphology analysis continues to evolve, with several emerging trends shaping future research directions. Multi-objective optimization approaches are gaining traction, simultaneously optimizing for competing objectives such as classification accuracy, computational efficiency, and model interpretability [62] [59]. Additionally, hybrid optimization algorithms that combine the strengths of different bio-inspired techniques (e.g., ACO with genetic algorithms or particle swarm optimization) show promise for addressing the complex, high-dimensional optimization landscapes presented by modern deep learning architectures [62] [59].
Another significant trend involves the application of federated learning frameworks enhanced with bio-inspired optimization, enabling model training across multiple institutions without sharing sensitive patient data. This approach addresses critical privacy concerns while leveraging diverse datasets to improve model generalization [6]. Furthermore, explainable AI (XAI) techniques, optimized using bio-inspired algorithms, are being developed to enhance the interpretability of DL models, providing clinicians with transparent insights into model decisions and increasing trust in automated sperm analysis systems [6].
Persistent Challenges and Limitations
Despite significant advancements, several challenges persist in the application of bio-inspired optimization to AI-based sperm morphology analysis. The dependency on large, high-quality annotated datasets remains a fundamental limitation, as DL models require extensive labeled data for training, and manual annotation by expert andrologists is time-consuming and expensive [2] [6]. Issues with model generalizability across diverse clinical settings, imaging protocols, and patient populations continue to pose significant hurdles for widespread clinical adoption [6].
The "black-box" nature of complex optimized models raises concerns regarding clinical validation and trust, particularly in the medically sensitive context of infertility diagnosis and treatment [6]. Additionally, computational resource requirements for both training optimized models and running bio-inspired optimization algorithms can be substantial, potentially limiting accessibility for resource-constrained healthcare settings [6] [59].
Ethical considerations surrounding data privacy, algorithmic bias, and appropriate regulatory frameworks for clinical deployment represent additional challenges that must be addressed through collaborative efforts between computer scientists, clinicians, ethicists, and regulatory bodies [6].
The integration of bio-inspired optimization algorithms with artificial intelligence represents a transformative approach to sperm morphology analysis, addressing critical limitations in conventional assessment methods while enhancing the accuracy, efficiency, and objectivity of male infertility diagnostics. Ant Colony Optimization and related algorithms provide powerful mechanisms for tuning AI models, optimizing architectures, and improving end-to-end analytical workflows. As research in this interdisciplinary field advances, focusing on addressing current challenges related to data quality, model generalizability, computational efficiency, and clinical validation will be essential for realizing the full potential of these technologies in reproductive medicine. The continued convergence of bio-inspired optimization and AI promises to reshape fertility care, paving the way for more personalized, accessible, and effective treatment strategies that can ultimately improve outcomes for individuals and couples facing infertility challenges.
The integration of artificial intelligence (AI) into sperm morphology analysis represents a paradigm shift in male fertility diagnostics, offering the potential to overcome the subjectivity and inconsistency of manual assessments [29] [18]. However, the transition from research prototypes to clinically robust tools hinges on solving the fundamental challenge of model generalizability. AI models, particularly deep learning architectures, often demonstrate exemplary performance on their training data but fail to maintain accuracy when confronted with real-world clinical data from different sources, protocols, or patient populations [64]. This performance degradation stems primarily from overfitting, where models learn spurious patterns specific to their training dataset rather than biologically relevant features of sperm morphology. The clinical implications are significant: a model that achieves >95% accuracy in a controlled research environment may provide misleading diagnostic information when deployed in a new clinic, potentially affecting treatment decisions for couples seeking infertility care [65] [1]. This technical guide examines the sources of overfitting in sperm morphology analysis and presents validated methodologies for developing models that maintain diagnostic accuracy across diverse clinical environments.
Root Causes of Overfitting in Sperm Morphology Analysis
Data Limitations and Annotation Inconsistencies
The development of robust AI models for sperm morphology analysis faces significant data-related challenges that predispose models to overfitting. A primary issue is the lack of standardized, high-quality annotated datasets with sufficient size and diversity [18]. Current publicly available datasets vary considerably in image resolution, staining protocols, and annotation criteria, forcing models to learn dataset-specific artifacts rather than generalizable morphological features. For instance, the HuSHeM dataset contains only 725 images with limited morphological classes, while the SCIAN-MorphoSpermGS dataset includes just 1,854 sperm images across five morphology classes [18]. This data scarcity compels models to memorize training examples rather than learning invariant features. Annotation inconsistency presents another critical challenge; sperm defect assessment requires simultaneous evaluation of head, vacuoles, midpiece, and tail abnormalities, leading to substantial inter-annotator variability that models may exploit as predictive features [18]. Furthermore, class imbalance problems are pervasive in sperm morphology datasets, with rare abnormality types being underrepresented, causing models to become biased toward majority classes [64].
Technical and Architectural Vulnerabilities
Beyond data limitations, certain technical approaches and architectural choices inherently increase vulnerability to overfitting. Conventional machine learning approaches for sperm analysis often rely on handcrafted features (e.g., shape descriptors, texture features) that may not capture the full complexity of morphological variations [18] [66]. Deep learning models, while capable of automated feature extraction, typically contain millions of parameters that require extensive regularization when training data is limited [64]. Models focused exclusively on sperm head morphology while neglecting other structural components (mid-piece, tail) develop an incomplete understanding of sperm morphology, limiting their ability to generalize to comprehensive clinical assessments [64]. Additionally, the dependency on single-model architectures rather than ensemble approaches increases sensitivity to noise and dataset-specific biases [64]. Training protocols that lack domain-specific augmentation strategies fail to expose models to the full spectrum of image variations encountered across different clinical settings, further exacerbating generalization issues [67].
Methodologies for Enhancing Model Generalization
Advanced Learning Paradigms
Contrastive Meta-Learning with Auxiliary Tasks (HSHM-CMA)
Meta-learning frameworks have demonstrated remarkable effectiveness in improving cross-domain generalization for sperm morphology classification. The HSHM-CMA (Contrastive Meta-Learning with Auxiliary Tasks) algorithm addresses gradient conflicts in multi-task learning by separating meta-training tasks into primary and auxiliary tasks [67]. This approach integrates localized contrastive learning in the outer loop of meta-learning to exploit invariant sperm morphology features across domains, significantly improving task convergence and adaptation to new categories.
Table 1: Performance of HSHM-CMA Under Different Generalization Scenarios
| Testing Objective | Dataset Relationship | Accuracy | Generalization Challenge |
|---|---|---|---|
| Same dataset, different HSHM categories | Fixed source domain, novel classes | 65.83% | Recognizing unseen morphological classes within similar image characteristics |
| Different datasets, same HSHM categories | Novel domain, known classes | 81.42% | Maintaining accuracy on known morphology types despite domain shift |
| Different datasets, different HSHM categories | Novel domain, novel classes | 60.13% | Simultaneous adaptation to new data sources and new morphological classes |
Experimental Protocol: The HSHM-CMA framework was evaluated using three distinct testing objectives to rigorously assess generalizability. Implementation requires (1) constructing a diverse task distribution from multiple sperm morphology datasets (HuSHeM, SCIAN-SpermMorphoGS, SMIDS), (2) applying episodic training where each episode contains a support set (for model adaptation) and query set (for evaluation), (3) employing contrastive learning to maximize similarity between embeddings of the same morphology class across different domains, and (4) optimizing the model using a meta-objective that explicitly minimizes loss on unseen tasks after adaptation [67].
Multi-Level Ensemble Learning with Feature Fusion
Ensemble-based classification approaches that combine convolutional neural network (CNN)-derived features using both feature-level and decision-level fusion techniques have demonstrated superior generalization capabilities compared to single-model architectures [64]. This methodology leverages complementary strengths from different feature representations, effectively creating a more robust morphological assessment system.
Experimental Protocol: The implementation involves (1) extracting features from multiple EfficientNetV2 variants as base architectures, (2) applying feature-level fusion by concatenating penultimate layer representations, (3) classifying fused features using diverse classifiers (Support Vector Machines, Random Forest, and Multi-Layer Perceptron with Attention), and (4) applying decision-level fusion via soft voting to enhance robustness [64]. This approach was validated on the Hi-LabSpermMorpho dataset containing 18 distinct sperm morphology classes and 18,456 image samples, where it achieved 67.70% accuracy, significantly outperforming individual classifiers [64].
Table 2: Performance Comparison of Ensemble vs. Single-Model Approaches
| Model Architecture | Accuracy | Key Strengths | Generalization Limitations |
|---|---|---|---|
| Single EfficientNetV2 Baseline | 58.2% | Architectural optimization for image classification | Vulnerable to domain shift in staining protocols |
| SVM on Traditional Features | 62.1% | Interpretable decision boundaries | Limited feature representation capacity |
| Feature-Level Fusion (Proposed) | 65.3% | Combines multi-scale feature representations | Increased computational complexity |
| Decision-Level Fusion (Proposed) | 64.8% | Robust to individual classifier failures | Requires training multiple architectures |
| Full Multi-Level Ensemble | 67.7% | Maximizes complementary strengths | Implementation complexity in clinical workflows |
Data-Centric Generalization Strategies
Cross-Domain Validation Frameworks
Rigorous validation methodologies are essential for accurately assessing true generalizability before clinical deployment. The cross-dataset validation protocol provides a realistic measure of performance in diverse clinical environments by testing trained models on completely external datasets with different acquisition protocols [1].
Experimental Protocol: Implementation requires (1) training models on one or multiple source datasets (e.g., VISEM-Tracking, SVIA dataset), (2) applying the trained model without fine-tuning to completely external datasets (e.g., HuSHeM, SCIAN-MorphoSpermGS), (3) measuring performance degradation across domains to identify vulnerability points, and (4) analyzing failure cases to understand specific domain shifts causing performance drops [18] [1]. This approach was utilized in developing an AI model for assessing unstained live sperm morphology, which demonstrated strong correlation (r=0.88) with computer-aided semen analysis when validated across multiple clinical sites [1].
Multi-Center Data Acquisition and Standardization
Addressing the fundamental data limitations in sperm morphology analysis requires systematic approaches to dataset creation. The establishment of standardized, high-quality annotated datasets through multi-center collaborations represents the most effective strategy for building models that generalize across clinical environments [18].
Experimental Protocol: Key steps include (1) establishing standardized protocols for sperm morphology slide preparation, staining, and image acquisition across participating centers, (2) implementing multi-tier annotation systems with expert consensus for challenging cases, (3) incorporating comprehensive morphological classes covering head, neck, and tail abnormalities, and (4) applying rigorous quality control measures for annotation consistency [18]. Recent initiatives like the SVIA (Sperm Videos and Images Analysis) dataset, comprising 125,000 annotated instances for object detection and 26,000 segmentation masks, demonstrate the scalability of this approach [18].
Implementation Framework for Clinical Deployment
Integrated Workflow for Robust Model Development
The transition from research to clinical implementation requires a systematic framework that incorporates generalization as a core requirement rather than an afterthought. The following workflow integrates the methodologies discussed previously into a comprehensive pipeline for developing clinically robust sperm morphology analysis systems.
Research Reagent Solutions for Sperm Morphology Analysis
The experimental methodologies described require specific technical resources and reagents to implement successfully. The following table details essential research reagents and their functions in developing generalizable AI models for sperm morphology analysis.
Table 3: Essential Research Reagents and Resources for Robust Sperm Morphology Analysis
| Reagent/Resource | Specifications | Function in Experimental Protocol |
|---|---|---|
| Confocal Laser Scanning Microscope | LSM 800, 40× magnification, Z-stack interval 0.5 μm | High-resolution imaging of unstained live sperm for model training [1] |
| Standardized Staining Kits | Diff-Quik stain (Romanowsky variant) | Consistent morphological visualization across multiple centers [1] |
| Annotated Datasets | SVIA Dataset: 125,000 instances, 26,000 masks | Training and validation of generalizable models [18] |
| Computational Framework | TensorFlow/PyTorch with multi-GPU support | Efficient training of ensemble and meta-learning models [64] |
| Validation Datasets | Hi-LabSpermMorpho (18 classes, 18,456 samples) | Cross-dataset generalization testing [64] |
| Sperm Slide Preparation | LEJA chambers (20 μm depth) | Standardized sample preparation for consistent imaging [1] |
The clinical translation of AI-based sperm morphology analysis depends critically on addressing the challenges of overfitting and limited generalizability. Through the implementation of advanced learning paradigms like contrastive meta-learning with auxiliary tasks and multi-level ensemble approaches, researchers can develop models that maintain diagnostic accuracy across diverse clinical environments. The methodologies presented in this technical guide—including cross-domain validation frameworks, systematic multi-center data acquisition, and integrated regularization strategies—provide a roadmap for creating robust, clinically applicable systems. As these approaches become more widely adopted, AI-powered sperm morphology analysis has the potential to standardize male fertility assessment globally, ultimately improving diagnostic accuracy and treatment outcomes for couples facing infertility.
Benchmarks and Efficacy: Validating AI Models Against Gold Standards and Clinical Outcomes
The integration of Artificial Intelligence (AI) into clinical diagnostics represents a paradigm shift in how medical data is interpreted. In the specific field of reproductive medicine, AI models for sperm morphology analysis are being developed to automate and standardize a process traditionally prone to subjectivity and inter-observer variability [1] [2]. The performance of these models has direct implications for clinical decision-making, patient diagnosis, and treatment success in assisted reproductive technology (ART) [6]. Therefore, moving beyond a simple measure of "accuracy" to a nuanced understanding of a suite of performance metrics is not merely an academic exercise but a clinical necessity. These metrics, including accuracy, precision, recall, and mean Average Precision (mAP), form the core language for evaluating, validating, and trusting AI tools before they can be safely integrated into patient care pathways.
This guide provides a detailed framework for researchers and clinicians to interpret these metrics within the context of AI-driven sperm morphology analysis. It outlines the fundamental definitions, explains their clinical significance, and presents structured data and methodologies from contemporary studies. Furthermore, it offers best practices for selecting and interpreting these metrics to ensure that AI models are not only technically proficient but also clinically reliable and effective.
Core Performance Metrics: Definitions and Clinical Interpretations
At its core, the evaluation of a classification AI model is based on counting how many times it was correct or incorrect in its predictions, broken down into four fundamental categories: True Positives (TP), True Negatives (TN), False Positives (FP), and False Negatives (FN). These counts are organized into a confusion matrix, which serves as the foundation for calculating all subsequent metrics [68].
The following diagram illustrates the logical relationships between the core confusion matrix elements and the primary performance metrics derived from them.
Metric Definitions and Clinical Implications
Table 1: Core Performance Metrics for Clinical AI Classification Models
| Metric | Calculation | Clinical Interpretation | Question Answered |
|---|---|---|---|
| Accuracy | (TP + TN) / Total Population | The overall proportion of correct sperm classifications (both normal and abnormal). | How often is the model correct overall? |
| Precision (PPV) | TP / (TP + FP) | When the model flags a sperm as abnormal, how often is it correct? A high precision minimizes false alarms. | How reliable is a positive (abnormal) result? |
| Recall (Sensitivity) | TP / (TP + FN) | The ability to find all truly abnormal sperm. High recall minimizes missed abnormalities. | What proportion of actual abnormalities does the model find? |
| F1-Score | 2 × (Precision × Recall) / (Precision + Recall) | The harmonic mean of precision and recall. Useful when a balanced view of both FP and FN is needed. | What is the balanced performance between precision and recall? |
| Specificity (TNR) | TN / (TN + FP) | The ability to correctly identify normal sperm. | What proportion of actual normal sperm does the model correctly identify? |
| mAP | Mean of Average Precision across classes | Used in object detection (e.g., locating sperm parts). Averages precision across all recall levels for multiple classes. | How accurate is the model at both finding and classifying objects? |
In a clinical setting, the choice of which metric to prioritize is dictated by the clinical consequence of error. For instance, in a diagnostic scenario for male infertility, a false negative (missing an abnormal sperm that indicates a potential fertility issue) could lead to a missed diagnosis and lack of appropriate treatment. Conversely, a false positive (misclassifying a normal sperm as abnormal) might lead to unnecessary further testing or the unjustified discarding of a viable sperm in ART [68]. Therefore, high recall (sensitivity) is critical when the cost of missing a positive case is high, while high precision is vital when the cost of a false alarm is high [68].
Metrics like mAP are particularly relevant for more complex AI tasks in sperm analysis, such as object detection, where the model must both locate and classify individual sperm or their subcellular components (head, neck, tail) within an image. A study on bovine sperm morphology using YOLOv7 reported a global mAP@50 of 0.73, indicating a reasonably good performance in correctly identifying and classifying sperm structures [11].
Performance Metrics in Practice: Sperm Morphology Analysis
The theoretical framework of performance metrics comes to life when applied to real-world AI research in sperm morphology. Recent studies demonstrate a trade-off between different metrics and highlight how model architecture and dataset quality directly influence outcomes.
Table 2: Reported Performance Metrics from Recent AI Studies in Sperm Morphology
| Study / Model | Task | Reported Accuracy | Reported Precision | Reported Recall | Other Key Metrics |
|---|---|---|---|---|---|
| In-house AI Model (ResNet50) [1] | Classification of normal/abnormal unstained live sperm | Test Accuracy: 0.93 | Abnormal: 0.95Normal: 0.91 | Abnormal: 0.91Normal: 0.95 | Correlation with CASA: r=0.88 |
| Bovine Sperm Analysis (YOLOv7) [11] | Object detection & morphological classification | - | 0.75 | 0.71 | mAP@50: 0.73 |
| Bull Sperm Analysis (YOLO Networks) [69] | Classification of viability and morphology | 0.82 | 0.85 | - | - |
| SMD/MSS Dataset (CNN) [40] | Multi-class classification of sperm defects | Range: 0.55 - 0.92 | - | - | (Performance varied by class) |
| Hybrid Diagnostic Framework [66] | Male fertility diagnosis from clinical profiles | 0.99 | - | 1.00 | - |
The variation in reported metrics underscores the importance of context. For example, the high accuracy (0.93) and strong precision/recall values of the ResNet50 model [1] reflect a well-trained system for a specific binary classification task. In contrast, the YOLOv7 model for bovine sperm [11], which performs the more complex task of object detection and multi-class classification, reports a mAP of 0.73, a solid result for such a task. The wide accuracy range (0.55 - 0.92) in the SMD/MSS study [40] highlights a common challenge: performance can significantly differ across morphological classes, especially with imbalanced datasets or for rare defect types.
Experimental Protocols in AI for Sperm Morphology
A critical factor in achieving reliable performance metrics is a robust experimental methodology. The following workflow visualizes a standardized pipeline for developing and evaluating an AI model for sperm morphology analysis, synthesized from current research practices [1] [11] [40].
Key Stages in the Workflow:
Sample Preparation & Imaging: Semen samples are processed onto slides, often stained (e.g., Diff-Quik, RAL stain), and imaged using microscopes, sometimes with specialized systems like confocal laser scanning microscopy [1] or bright-field microscopy with a CASA system [40]. Standardization here is crucial for image quality.
Data Annotation: This is a critical step for generating "ground truth" labels. Expert embryologists or trained analysts manually annotate thousands of sperm images, classifying them into categories like "normal," "abnormal head," "bent neck," etc. [1] [40]. Studies often report inter-expert agreement coefficients (e.g., 0.95) to establish label reliability [1].
Data Preprocessing: The raw images are prepared for model consumption. This involves:
- Cleaning: Removing noise and artifacts from images [40].
- Normalization: Resizing images and scaling pixel values to a standard range (e.g., 0-1) [66] [40].
- Augmentation: Techniques like rotation, flipping, and scaling are used to artificially expand the dataset and improve model generalizability, especially for rare defect classes [40]. One study increased its dataset from 1,000 to 6,035 images via augmentation [40].
Model Training & Validation: The dataset is split (e.g., 80% for training, 20% for testing). A deep learning model, typically a Convolutional Neural Network (CNN) or an object detection framework like YOLO, is trained on the training set. Its performance is periodically checked on a validation set to tune parameters and prevent overfitting [1] [11] [69].
Performance Evaluation & Clinical Validation: The final model is evaluated on the held-out test set, and metrics like accuracy, precision, and recall are calculated. For clinical relevance, the AI's performance is often directly compared against existing standards like Computer-Aided Semen Analysis (CASA) or Conventional Semen Analysis (CSA) through correlation analysis [1].
The Scientist's Toolkit: Essential Research Reagents and Materials
The development and validation of AI models for sperm morphology rely on a foundation of specialized laboratory equipment, software, and datasets. The following table details key resources referenced in recent studies.
Table 3: Essential Research Reagents and Solutions for AI-Based Sperm Morphology Analysis
| Item / Resource | Specification / Example | Primary Function in the Workflow |
|---|---|---|
| Microscopy Systems | Confocal Laser Scanning Microscope (e.g., LSM 800) [1]; Bright-field microscope (e.g., Optika B-383Phi) [11]; CASA-integrated microscope [40] | High-resolution image acquisition of sperm cells for dataset creation. |
| Staining Kits | Diff-Quik stain (Romanowsky variant) [1]; RAL Diagnostics staining kit [40] | Enhances contrast of sperm structures (head, midpiece, tail) for morphological assessment. |
| Sample Preparation Aids | Standardized slides (e.g., Leja) [1]; Trumorph system for fixation [11]; Optixcell extender [11] | Standardizes sperm immobilization and presentation for consistent imaging. |
| Annotation Software | LabelImg program [1]; Roboflow [11] | Allows experts to draw bounding boxes and assign class labels to sperm in images, creating ground truth data. |
| Public Datasets | HSMA-DS [1] [2]; MHSMA [2]; VISEM-Tracking [2]; SVIA [1] [2] | Provides benchmark data for training and comparing AI models, fostering reproducibility. |
| Programming Frameworks | Python (v3.8) [40]; Deep Learning libraries (e.g., for YOLOv7 [11], ResNet50 [1], CNNs [40]) | Provides the software environment to build, train, and evaluate AI models. |
Best Practices for Metric Selection and Interpretation in Clinical Context
Selecting and interpreting the right metrics requires a strategy aligned with clinical goals. The European Society of Medical Imaging Informatics recommends using task-specific performance metrics and considering the deployment context when assessing AI performance [68]. The following guidelines translate this principle into actionable steps for sperm morphology AI research:
Align Metrics with the Clinical Task: Determine the primary goal of the AI tool.
- Screening for Defects: If the goal is to identify every potential abnormality for further review, high recall is paramount to minimize false negatives.
- Confirming Normality for ART: If the goal is to select unequivocally normal sperm for injection (e.g., in ICSI), high precision is critical to ensure that sperm classified as "normal" are indeed normal, minimizing false positives [68].
Go Beyond a Single Metric: Never rely on accuracy alone, especially with imbalanced datasets. A model can achieve high accuracy by simply always predicting the majority class. Always report a suite of metrics, including precision, recall (sensitivity), and specificity, to provide a complete picture of model behavior [2] [68].
Validate on Independent, Local Datasets: Performance on a clean, curated research dataset may not translate to a different clinical lab. Conduct local validation using an independent dataset that reflects your institution's patient demographics, imaging protocols, and staining methods. This is essential for ensuring the claimed performance holds in a real-world setting [68].
Report Prevalence and Use Prevalence-Dependent Metrics: Disease prevalence in the test population directly impacts the clinical meaning of a result. Calculate and report outcome-based metrics like Positive Predictive Value (PPV, synonymous with precision) and Negative Predictive Value (NPV), as these depend on prevalence and tell a clinician the probability that a positive (or negative) AI result is correct in their patient population [68].
Conduct Comparative Analysis: To establish clinical utility, compare the AI model's performance and outputs against the current gold-standard methods, such as CASA or manual assessment by senior embryologists. Reporting correlation coefficients, as done in a study which found a correlation of r=0.88 between an AI model and CASA [1], provides strong evidence for validity.
The accurate interpretation of performance metrics is the cornerstone of translating AI research in sperm morphology into trustworthy clinical tools. As the field progresses, a sophisticated understanding of what accuracy, precision, recall, and mAP represent in a diagnostic context is mandatory for researchers and clinicians alike. By adhering to rigorous experimental protocols, selecting metrics that reflect the clinical stakes, and validating models in real-world settings, the promise of AI to bring unprecedented objectivity, efficiency, and success to the diagnosis and treatment of male infertility can be fully realized. This disciplined approach ensures that these powerful new tools are not only technically impressive but also clinically impactful and safe for patient care.
Sperm morphology analysis is a critical component of male fertility assessment, providing vital diagnostic and prognostic information for clinical outcomes in assisted reproductive technology (ART). For decades, the field has relied on two primary methodologies: manual assessment by trained embryologists and Computer-Aided Sperm Analysis (CASA) systems. Manual assessment, while considered the traditional standard, is inherently subjective and suffers from significant inter-observer variability [2]. Traditional CASA systems introduced a degree of automation but often relied on simplified algorithms and required sperm staining, which renders sperm unusable for subsequent procedures [1].
The emergence of Artificial Intelligence (AI), particularly deep learning, represents a paradigm shift. AI-powered systems offer the potential for fully automated, highly accurate, and objective sperm analysis. This whitepaper provides a comparative analysis of these three methodologies—AI, manual embryologist assessment, and traditional CASA—framed within the broader thesis that AI research is fundamentally advancing sperm morphology analysis from a subjective art to a quantitative, data-driven science. The integration of AI not only enhances current capabilities but also opens new avenues for non-invasive, real-time assessment that was previously impossible [1] [70].
Technical Performance Comparison
A direct comparison of key performance metrics reveals the distinct advantages and limitations of each sperm morphology assessment method. The following table synthesizes quantitative and qualitative data from recent studies.
Table 1: Technical Performance Comparison of Sperm Morphology Assessment Methods
| Feature | AI-Driven Systems | Manual Embryologist Assessment | Traditional CASA Systems |
|---|---|---|---|
| Correlation with CASA | Strong (r=0.88) [1] | Moderate (r=0.76) [1] | (Self) |
| Correlation with Manual Assessment | Moderate to Strong (r=0.76) [1] | (Self) | Weaker (r=0.57) [1] |
| Analysis Accuracy | High (e.g., Test Accuracy: 93%, Precision: 91-95%) [1] | Variable (subject to observer experience and fatigue) [2] | Lower than AI and Manual [1] |
| Objectivity | High (Minimizes subjectivity) [1] [70] | Low (High inter-observer variability) [2] | Medium (Rule-based, but limited by algorithms) |
| Key Advantage | Objective, automated, high accuracy, can use live/unstained sperm [1] | Considered the traditional gold standard, requires no capital equipment | Provides some quantitative data beyond human perception |
| Key Limitation | Requires large, high-quality datasets for training [2] | Subjective, labor-intensive, inconsistent [2] | Often requires staining; lower accuracy and correlation [1] |
| Sperm Status | Can assess unstained, live sperm [1] | Requires stained, fixed sperm [1] | Typically requires stained, fixed sperm [1] |
Experimental Protocols in AI Sperm Morphology Research
The development of robust AI models for sperm morphology requires meticulously designed experimental protocols. The following section details the methodology from a seminal study that developed an in-house AI model for assessing unstained live sperm, providing a template for research in this field.
Study Design and Sample Preparation
In a 2025 experimental study, 30 healthy male volunteers (aged 18-40) were enrolled. Participants maintained 2-7 days of sexual abstinence before providing an ejaculate via masturbation. Each semen sample was divided into three aliquots for parallel assessment by the three methods: the in-house AI model, a commercial CASA system (IVOS II), and conventional semen analysis (CSA) performed by embryologists according to Björndahl guidelines and the WHO laboratory manual [1].
AI Model Development Workflow
The core of the AI methodology involved a multi-stage process for data acquisition, annotation, and model training. The workflow is summarized in the diagram below.
Image Acquisition and Dataset Curation: The critical first step involved creating a novel, high-resolution dataset. Sperm images were captured using confocal laser scanning microscopy (LSM 800) at 40x magnification in confocal mode (Z-stack). This produced high-resolution images (512x512 pixels) without the need for staining, preserving sperm viability [1].
Annotation and Labeling: Embryologists and researchers manually annotated well-focused sperm images using the LabelImg program. Each sperm was categorized into one of nine datasets based on strict WHO criteria for normal and abnormal morphology (e.g., smooth oval head, no vacuoles, normal tail). A high inter-annotator agreement was reported, with a correlation coefficient of 0.95 for normal sperm and 1.0 for abnormal sperm detection [1].
Model Architecture and Training: The study employed a deep learning approach using the ResNet50 architecture, a well-established convolutional neural network (CNN) for image classification. The model was trained using a transfer learning strategy on a subset of 9,000 images (4,500 normal and 4,500 abnormal) to minimize the difference between predicted and actual labels. The model's performance was evaluated on a separate, unseen test dataset [1].
The Scientist's Toolkit: Key Research Reagents and Materials
The experimental protocol for advanced AI-based sperm morphology analysis relies on a specific set of reagents, equipment, and software. The following table details these essential components and their functions, serving as a guide for researchers seeking to replicate or build upon this work.
Table 2: Essential Research Materials for AI-Based Sperm Morphology Analysis
| Category | Item / Technology | Specification / Function |
|---|---|---|
| Core Imaging Equipment | Confocal Laser Scanning Microscope | e.g., LSM 800; enables high-resolution, label-free imaging of live sperm via Z-stack scanning [1]. |
| Clinical Analysis Equipment | Computer-Aided Sperm Analysis (CASA) System | e.g., IVOS II (Hamilton Thorne); provides automated, quantitative sperm analysis for comparative studies [1]. |
| Software & Algorithms | ResNet50 Deep Learning Model | A Convolutional Neural Network (CNN) architecture used for image classification via transfer learning [1]. |
| Annotation Software | LabelImg Program | Open-source tool for manually annotating and labeling sperm images to create ground truth data for model training [1]. |
| Clinical Consumables | Standard Two-Chamber Slide | e.g., Leja slide (20 µm depth); provides a standardized environment for imaging live sperm [1]. |
| Staining Reagents (For Comparator Methods) | Diff-Quik Stain | A Romanowsky stain variant used to prepare sperm for traditional CASA and conventional semen analysis [1]. |
Analysis and Future Directions
The quantitative data and experimental details presented confirm that AI-driven systems are establishing a new benchmark for sperm morphology analysis. Their superior correlation with existing methods, combined with high accuracy and the unique ability to use unstained, viable sperm, positions them as a transformative technology [1]. This capability is crucial for ART, as it allows for the selection of high-quality sperm with normal morphology that can be used immediately in intracytoplasmic sperm injection (ICSI), potentially leading to improved fertility outcomes [1].
However, significant challenges remain for widespread adoption. A primary hurdle is the lack of standardized, high-quality annotated datasets needed to train robust deep learning models [2]. Barriers such as high implementation costs, lack of training for clinical staff, and ethical concerns regarding over-reliance on technology also temper the pace of adoption, as evidenced by global surveys of fertility specialists [14]. Furthermore, it is crucial to distinguish between true AI, which uses adaptive algorithms for predictive analytics, and simple automation, which follows pre-set rules [71]. This distinction is vital for managing expectations and making informed technological investments.
Future research will likely focus on creating large, multi-center, standardized datasets to improve model generalizability. Furthermore, the integration of AI with other advanced, label-free imaging modalities, such as fluorescence lifetime imaging microscopy (FLIM) and holographic microscopy, promises to add a new dimension of metabolic and biophysical data to morphological assessment [70]. As these technologies mature, AI is poised to move from a research tool to an integral component of a fully objective, efficient, and predictive clinical workflow in reproductive medicine.
Sperm DNA fragmentation (SDF) has emerged as a critical parameter in male fertility assessment that conventional semen analysis fails to evaluate adequately [21]. While routine semen analysis provides basic parameters like concentration and motility, it offers limited insight into the molecular integrity of sperm DNA, which is now recognized as crucial for successful fertilization and embryonic development [72]. Male factors contribute to approximately 50% of infertility cases, with unexplained infertility detected in about 30% of these couples [72]. In a substantial portion of males identified as having unexplained infertility, high levels of fragmented sperm DNA are often the underlying cause [72]. This diagnostic gap has accelerated the development of artificial intelligence (AI) tools that can predict DNA fragmentation status from standard phase-contrast microscopy images, creating an urgent need for robust validation against functional biochemical assays [21].
The clinical significance of DNA fragmentation cannot be overstated. High DNA fragmentation index (DFI) is associated with increased miscarriage rates and lower live birth rates, making it an essential parameter for comprehensive fertility assessment [72]. Consequently, validating AI predictions against established functional assays represents a critical step toward clinical adoption, potentially enabling real-time sperm selection based on DNA integrity for therapeutic applications [21]. This technical guide examines the current methodologies for correlating AI-based morphological assessments with functional DNA fragmentation tests, with particular emphasis on validation frameworks, experimental protocols, and performance benchmarks.
DNA Fragmentation Assays: Gold Standards for Validation
Terminal Deoxynucleotidyl Transferase dUTP Nick End Labeling (TUNEL) Assay
The TUNEL assay stands as one of the most robust and widely recognized methods for detecting sperm DNA fragmentation [21]. This biochemical assay operates on the principle of using fluorescent nucleotides to identify DNA 'nicks' or free ends through the enzyme terminal deoxynucleotidyl transferase (TdT) [72]. The fundamental mechanism involves TdT catalyzing the addition of fluorescently-labeled dUTP to the 3'-hydroxyl termini of DNA breaks, allowing for direct visualization and quantification of DNA damage in individual spermatozoa.
When employed as a validation reference for AI tools, the TUNEL assay provides binary classification of sperm as either DNA fragmented or intact. In a landmark validation study, an AI tool designed to detect SDF through digital analysis of phase contrast microscopy images utilized TUNEL as the gold standard reference [21]. The AI methodology leveraged the established link between sperm morphology and DNA integrity, employing a morphology-assisted ensemble model that combined image processing techniques with state-of-the-art transformer-based machine learning models (GC-ViT) for predicting DNA fragmentation in sperm from phase contrast images [21].
Sperm Chromatin Dispersion (SCD) Test
The SCD test, also referred to as the halo test, provides an alternative methodology for DNA fragmentation assessment based on the differential dispersion of nuclear proteins and DNA loops [72]. The underlying principle of this assay centers on the fact that sperm with fragmented DNA fail to create the distinctive halo of dispersed DNA loops that are characteristic of non-fragmented sperm after acid denaturation and removal of nuclear proteins [72]. This assay classifies sperm into multiple categories based on halo dispersion patterns: big halo (BH), medium halo (MH), small halo (SH), and degraded (DEG), with BH and MH indicating intact DNA and SH and DEG indicating poor DNA integrity [72].
The SCD test offers practical advantages for AI validation studies due to its straightforward methodology that doesn't require sophisticated equipment [72]. However, a significant consideration for validation frameworks is the potential interobserver subjectivity in classifying halo patterns, which can be mitigated through standardized AI annotation protocols [72]. In research settings, the SCD test has been utilized to generate large datasets for AI training, with one study compiling 24,415 images from 30 patients, which were then classified into both binary (halo/no halo) and multiclass (BH/MH/SH/DEG) configurations for model development [72].
Comparative Analysis of DNA Fragmentation Assays
Table 1: Comparison of Key DNA Fragmentation Assays Used for AI Validation
| Assay Type | Underlying Principle | Detection Method | Classification Output | Equipment Requirements | Advantages | Limitations |
|---|---|---|---|---|---|---|
| TUNEL [21] [72] | Enzymatic labeling of DNA strand breaks | Fluorescence microscopy | Binary (fragmented/intact) | Fluorescence microscope/flow cytometer | High specificity and accuracy; Considered gold standard | Higher cost; Requires specialized equipment |
| SCD (Halo Test) [72] | Differential DNA dispersion patterns | Bright-field microscopy | Multiclass (BH, MH, SH, DEG) | Standard optical microscope | Low cost; Simple protocol; No specialized equipment | Interobserver variability in halo classification |
| SCSA [72] | DNA susceptibility to denaturation | Flow cytometry | Sperm Chromatin Structure Assay | Flow cytometer | High throughput; Quantitative | Requires flow cytometry; Complex data analysis |
| COMET [72] | Electrophoretic DNA migration | Fluorescence microscopy | Continuous DNA damage measurement | Electrophoresis + fluorescence microscope | Sensitive; Quantifies various DNA damage types | Labor-intensive; Not suitable for rapid diagnosis |
AI Architectures for DNA Fragmentation Prediction
Model Architectures and Performance
Multiple AI architectures have been developed to correlate sperm morphology with DNA fragmentation status, with varying levels of complexity and performance characteristics. The most promising approaches utilize ensemble methods and deep learning architectures that can extract nuanced morphological features associated with DNA integrity.
Table 2: AI Models for Predicting DNA Fragmentation from Morphology
| Model Architecture | Input Data | Validation Assay | Key Performance Metrics | Advantages | Limitations |
|---|---|---|---|---|---|
| Morphology-Assisted Ensemble AI [21] | Phase contrast images | TUNEL | Sensitivity: 60%; Specificity: 75% | Combines image processing with transformer models; Non-destructive | Moderate sensitivity; Requires further optimization |
| Pure Transformer Vision Model [21] | Phase contrast images | TUNEL | Benchmark against ensemble | State-of-the-art architecture; Direct feature learning | Performance details not fully specified |
| Convolutional Neural Network (CNN) [40] | Bright-field stained images | Expert annotation (David classification) | Accuracy: 55%-92% (class-dependent) | Handles complex feature hierarchies; Proven image classification capability | Wide accuracy range suggests class imbalance issues |
| Custom Vision (Azure) [72] | SCD test images | SCD (manual annotation) | Binary F1-score: 0.81; Multiclass F1-score: 0.72 | Leverages transfer learning; Effective with limited data | Multiclass performance significantly lower than binary |
The morphology-assisted ensemble model represents a particularly innovative approach, combining traditional image processing techniques with state-of-the-art transformer-based machine learning models (GC-ViT) for predicting DNA fragmentation in sperm from phase contrast images [21]. This hybrid methodology achieves a promising balance between sensitivity (60%) and specificity (75%) when validated against TUNEL assay results [21]. The ensemble approach benchmarks performance against both pure transformer 'vision' models and 'morphology-only' models, establishing a robust framework for comparative analysis [21].
Experimental Workflow for AI Validation
The validation of AI models for DNA fragmentation prediction follows a systematic experimental workflow that integrates both computational and biochemical components. The process begins with sample preparation and proceeds through image acquisition, biochemical assay processing, AI model training, and statistical correlation analysis.
AI Validation Workflow: From Sample Collection to Clinical Deployment
Research Reagent Solutions and Experimental Materials
Essential Materials for Validation Studies
Table 3: Key Research Reagents and Experimental Materials
| Category | Specific Product/Type | Application/Function | Implementation Example |
|---|---|---|---|
| Staining Kits | RAL Diagnostics staining kit [40] | Sperm morphology visualization | Sample preparation for the SMD/MSS dataset [40] |
| DNA Fragmentation Assays | Sperm Chroma Kit (Cryotec) [72] | SCD test performance | Standardized halo pattern generation for AI training [72] |
| Microscopy Systems | MMC CASA System [40] | Image acquisition | Digital capture of sperm images with 100x oil immersion objective [40] |
| Image Annotation Tools | Custom Vision (Azure) [72] | Automated image classification | Transfer learning and data augmentation for model training [72] |
| Data Augmentation Tools | Python 3.8 with augmentation libraries [40] | Dataset expansion | Rotation, saturation, and Gaussian blur/noise application [40] |
Performance Metrics and Validation Frameworks
Quantitative Performance Analysis
The performance of AI models in predicting DNA fragmentation from morphological features varies significantly based on architecture, training data, and validation methods. Recent studies demonstrate a range of efficacy metrics that highlight both the potential and limitations of current approaches.
In binary classification tasks (e.g., fragmented vs. non-fragmented), AI models generally demonstrate stronger performance. A study utilizing Azure's Custom Vision for SCD test interpretation achieved an F1-score of 0.81 for binary classification (fragmented/unfragmented) compared to 0.72 for multiclass classification (big/medium/small/degraded) [72]. Similarly, accuracy metrics showed better performance for binary approaches (80.15%) versus multiclass approaches (75.25%) [72].
For CNN architectures applied to morphological classification, performance shows considerable variation across different abnormality classes, with accuracy ranging from 55% to 92% depending on the specific morphological defect [40]. This wide range underscores the challenge of developing unified models that perform consistently across diverse morphological anomalies.
Inter-Expert Agreement and Ground Truth Establishment
A critical aspect of validation framework development involves addressing the inherent subjectivity in morphological classification. Studies implementing rigorous inter-expert agreement protocols reveal the complexity of establishing reliable ground truth data. Research utilizing three independent experts reported varying agreement levels: no agreement (NA) among experts, partial agreement (PA) where 2/3 experts concurred on labels, and total agreement (TA) where 3/3 experts agreed on all categories [40]. Statistical analysis using Fisher's exact test revealed significant differences between expert classifications in various morphology classes (p < 0.05), highlighting the critical importance of standardized annotation protocols for training reliable AI models [40].
Future Directions and Clinical Implementation
The integration of AI-based morphological assessment with DNA fragmentation validation represents a paradigm shift in male fertility evaluation. Current research demonstrates promising correlations between morphological features and DNA integrity, enabling non-destructive sperm selection for assisted reproductive technologies. The ensemble approach combining image processing with transformer models achieves clinically relevant performance levels (60% sensitivity, 75% specificity) when validated against TUNEL assays [21].
Future developments in this field will likely focus on multi-modal AI architectures that integrate morphological, motile, and biochemical parameters to enhance predictive accuracy. Additionally, standardization of validation protocols across research institutions will be essential for clinical translation. As these technologies mature, AI-powered sperm analysis systems capable of predicting DNA fragmentation from standard microscopy images have the potential to revolutionize clinical andrology laboratories, making advanced fertility assessment more accessible and cost-effective.
Artificial Intelligence (AI) is fundamentally transforming the field of reproductive biology, enabling unprecedented precision in the assessment of gamete quality. This transition from subjective, manual evaluations to automated, data-driven diagnostics is particularly impactful in sperm morphology analysis—a critical determinant of male fertility. Within both human andrology and veterinary medicine, AI-powered systems are now capable of extracting subtle, predictive patterns from sperm images that elude human visual inspection [6]. This technical guide explores the operational frameworks of these AI technologies through concrete case studies, with a specific focus on bull sperm analysis—a domain where genetic improvement and economic outcomes provide a compelling context for innovation. The integration of machine learning (ML) and deep learning (DL) algorithms is not merely automating existing procedures; it is reshaping diagnostic standards, enhancing reproducibility, and forging a new pathway for objective male fertility assessment [5] [2].
Technical Foundations of AI in Sperm Morphology Analysis
The application of AI in sperm analysis spans a hierarchy of computational techniques, each with distinct capabilities and requirements.
From Conventional Machine Learning to Deep Learning
Conventional machine learning approaches have historically been applied to sperm image analysis. These methods, including Support Vector Machines (SVM), K-means clustering, and decision trees, rely heavily on manually engineered features such as shape descriptors (e.g., Hu moments, Zernike moments), grayscale intensity, and texture patterns for segmentation and classification [2]. For instance, Bayesian Density Estimation and Fourier descriptors have been used to classify sperm heads into morphological categories with up to 90% accuracy [2]. However, their performance is limited by their dependence on these handcrafted features, which often struggle with the complex and variable nature of sperm morphology, leading to challenges in generalizing across different datasets and imaging conditions [2].
In contrast, deep learning (DL), a subset of AI based on artificial neural networks with multiple layers (hence "deep"), automates the feature extraction process. DL models, particularly convolutional neural networks (CNNs), can learn hierarchical representations directly from raw pixel data, capturing intricate features from sperm images without human intervention [5] [6]. This capability makes DL exceptionally suited for complex tasks like segmenting complete sperm structures (head, neck, and tail) and classifying a wide spectrum of abnormalities [2].
Key Algorithmic Architectures
Several neural network architectures are central to modern sperm analysis:
- YOLO (You Only Look Once) Networks: A type of CNN used for real-time object detection and classification. A recent study developed a YOLO-based algorithm to identify sperm cells in microscope-acquired images, establish their viability, and classify their morphology with an accuracy of 82% and a precision of 85% [69].
- Deep Neural Networks for Classification: Multi-layered networks are trained on large datasets of sperm images to differentiate between "normal" and "abnormal" morphologies, and to categorize specific defect types [2].
Table 1: Comparison of AI Approaches to Sperm Morphology Analysis.
| Feature | Conventional Machine Learning | Deep Learning |
|---|---|---|
| Feature Extraction | Manual, based on expert-defined parameters (e.g., shape, texture). | Automatic, learned directly from data. |
| Data Dependency | Performs well with smaller datasets. | Requires large, annotated datasets for training. |
| Complexity Handling | Struggles with complex structures and high variability. | Excels at managing complex patterns and variations. |
| Representative Algorithms | Support Vector Machine (SVM), K-means, Decision Trees. | YOLO, Convolutional Neural Networks (CNNs). |
| Reported Accuracy | Up to 90% for specific tasks like head classification [2]. | Up to 85-90% precision for holistic morphology classification [69] [2]. |
Case Study: AI-Driven Bull Sperm Morphology Evaluation
The evaluation of bull sperm is a critical component in the artificial insemination (AI) industry, directly impacting genetic progress and economic returns. The following case study exemplifies the practical implementation and validation of an AI system in this context.
Experimental Protocol and Workflow
A seminal study aimed to develop an AI algorithm for the automated classification of bull spermatozoa morphology, moving away from the subjective visual assessments guided by the Society for Theriogenology (SFT) standards [69].
1. Sample Preparation and Imaging:
- Sample Source: Semen samples were obtained from bulls.
- Imaging: Sperm cells were visualized using bright-field microscopy to acquire high-resolution digital images.
2. Dataset Curation and Annotation:
- A substantial dataset of 8,243 microscope-acquired images was compiled.
- Experts then labeled and annotated each image, using bounding boxes to segment individual sperm cells and classify them according to a simplified scheme (e.g., normal, major defect, minor defect, non-sperm cell) [69]. This annotated dataset is the fundamental resource for training the AI model.
3. AI Model Training and Validation:
- Algorithm: A YOLO (You Only Look Once) network, a type of convolutional neural network (CNN) designed for fast and efficient object detection, was employed [69].
- Process: The model was trained on the annotated dataset, learning to extract features and associate them with the expert-provided labels.
- Performance Metrics: The model's performance was evaluated based on its accuracy (82%) and precision (85%) in correctly classifying spermatozoa morphology [69].
The following diagram illustrates the end-to-end experimental workflow.
Performance Results and Clinical Relevance
The AI model demonstrated high efficacy in automating a traditionally labor-intensive task. The achieved precision of 85% indicates a low rate of false positives, which is crucial for reliable diagnosis [69]. While the overall accuracy of 82% was high, the study noted that performance varied across different defect classes, highlighting an area for continued model refinement. This level of accuracy supports the model's potential for use in bull breeding soundness evaluations (BBSE), offering a more standardized and objective technique. This is especially valuable for implementing genomic selection in young bulls, where accurate assessment of sperm abnormalities that affect freezing suitability and fertilizing capacity is paramount [69].
Case Study: Standardizing Sperm Concentration Analysis in a Multi-Laboratory Setting
While morphology is crucial, the accurate assessment of sperm concentration is equally fundamental. A multi-laboratory study focused on standardizing bull sperm concentration analysis demonstrates how technology and rigorous protocol can ensure data reliability across different sites.
Experimental Protocol for Standardization
The study was conducted across seven commercial bovine semen processing laboratories to assess the effectiveness of a standardization program [73].
1. Instrumentation and Reference Standards:
- Primary Instrument: The NucleoCounter SP-100, an automated system that quantifies fluorescently labelled sperm, was used as the standard instrument [73].
- Reference Standards: Frozen semen samples with known concentrations were used as reference standards to calibrate the instruments and monitor performance over time.
2. Standardized Procedures and Personnel Training:
- Standard Operating Procedures (SOPs): All participating laboratories followed the same SOPs for semen analysis.
- E-Learning Training: A customized e-learning training program was implemented to harmonize technician skills across laboratories. The impact of this training was quantitatively assessed.
3. Multi-Laboratory Validation:
- Design: Ten batches of frozen semen were produced, with three replicates of each batch coded differently to blind the technicians.
- Testing: Sperm concentration was evaluated by nine technicians across six laboratories during three test periods over one year to measure both intra-technician and inter-laboratory variability [73].
Results and Impact on Data Integrity
The standardization program yielded highly precise and accurate results. Key findings included:
- Low Variability: The intra-technician coefficient of variation (CV) was 3.4 ± 3.1%, and the intra-batch CV was 4.6 ± 2.2%, indicating high repeatability and consistency [73].
- Effect of Training: Following e-learning training, the proportion of sample duplicates with results differing by more than 10% significantly decreased from 8.1% to 6.9% [73].
- Minimal Bias: Bland-Altman analysis confirmed that the mean difference between individual results and the overall mean was close to zero, demonstrating a lack of significant systematic bias across laboratories [73].
This case underscores that precise and accurate concentration results in a real-world, multi-laboratory setting are achievable through the combination of robust technology (NucleoCounter), standardized procedures, and effective personnel training [73].
Table 2: Key Performance Metrics from Multi-Laboratory Standardization Study [73].
| Metric | Pre-Training Result | Post-Training Result |
|---|---|---|
| Coefficient of Variation (CV) for Duplicate Results | 3.2 ± 3.8% | 3.0 ± 3.2% |
| Samples with Duplicate Results >10% Difference | 8.1% | 6.9% |
| Overall Intra-Technician CV | - | 3.4 ± 3.1% |
| Overall Intra-Batch CV | - | 4.6 ± 2.2% |
The following diagram visualizes the structured approach of this multi-laboratory validation study.
The Scientist's Toolkit: Essential Research Reagents and Materials
The advancement and application of AI in sperm analysis are underpinned by a suite of specialized reagents, instruments, and computational tools.
Table 3: Essential Research Reagents and Materials for AI-Based Sperm Analysis.
| Item | Function/Application | Example Use-Case |
|---|---|---|
| NucleoCounter SP-100 | Automated, precise quantification of sperm concentration using fluorescence. | Standardized sperm concentration analysis in multi-laboratory settings [73]. |
| Percoll Density Gradient Centrifugation (PDGC) | Technique for separating spermatozoa based on density; used in sperm sexing and quality enrichment. | Optimization of sexing protocols for Holstein-Friesian bull sperm; 20%-65% gradient showed superior performance [74]. |
| Droplet-Loaded Microfluidic Chips | Single-use, disposable chips for consistent sample loading and imaging in portable analyzers. | User-friendly sample handling in the iSperm portable analyzer for on-farm boar semen evaluation [75]. |
| YOLO (You Only Look Once) Networks | A class of convolutional neural networks for real-time object detection and image classification. | Automated classification of bull sperm vitality and morphology from microscope images [69]. |
| Annotated Sperm Image Datasets | Curated, labeled datasets of sperm images (e.g., SVIA, MHSMA) used for training and validating AI models. | Training deep learning models for sperm head, neck, and tail segmentation and defect classification [2]. |
The integration of artificial intelligence into reproductive medicine represents a paradigm shift from subjective assessment to quantitative, data-driven science. As demonstrated by the case studies in bull sperm analysis, AI systems are not merely replicating human expertise but are enhancing it, providing levels of standardization, throughput, and analytical depth that were previously unattainable. The successful application of YOLO networks for morphology classification and the rigorous multi-laboratory standardization of concentration measurements prove that these technologies are mature for real-world deployment.
The future trajectory of this field points towards even greater integration. Portable systems like the iSperm analyzer combine microfluidics and mobile AI for on-farm diagnostics [75], while ongoing research focuses on overcoming the challenges of dataset standardization and the "black-box" nature of complex algorithms [2] [6]. Ultimately, the continuous refinement of these AI tools promises to reshape the landscape of both human and veterinary reproduction, enabling more precise fertility diagnoses, optimizing assisted reproductive outcomes, and accelerating genetic progress in livestock industries.
The integration of Artificial Intelligence (AI) into sperm morphology analysis represents a paradigm shift in male fertility assessment, offering the potential to overcome long-standing limitations of conventional semen analysis. Traditional sperm morphology assessment is inherently subjective, prone to significant inter-observer variability, and hampered by methodological inconsistencies [2] [17]. While AI-powered systems promise objectivity, reproducibility, and enhanced accuracy, their transition from research laboratories to clinical settings requires rigorous standardization and robust regulatory approval pathways. This technical guide examines the core requirements for clinical deployment of AI-based sperm morphology analysis systems, focusing on validation frameworks, data standardization, regulatory considerations, and implementation protocols essential for researchers, scientists, and drug development professionals working in this field.
The clinical imperative for standardization is underscored by performance variations observed in current assessment methods. Studies demonstrate that untrained morphologists exhibit high variability (CV = 0.28) and accuracy as low as 53% when using complex 25-category classification systems, though standardized training can improve accuracy to 90% even for intricate classification schemes [17]. AI models have demonstrated strong correlations (r = 0.88) with computer-aided semen analysis while offering the distinct advantage of analyzing unstained, live sperm—a capability that preserves sperm viability for subsequent assisted reproductive technology (ART) procedures [1]. This technical advancement highlights the transformative potential of AI in clinical andrology, provided that appropriate standardization and regulatory frameworks are established.
Performance Validation and Benchmarking
Comprehensive performance validation against established standards forms the foundation of clinical deployment for AI-based sperm morphology systems. This requires rigorous comparison with both conventional semen analysis (CSA) and computer-aided semen analysis (CASA) methods across multiple performance dimensions.
Table 1: Performance Metrics of AI Sperm Morphology Analysis Compared to Conventional Methods
| Parameter | AI Model Performance | Conventional CSA | CASA Systems | Clinical Validation Requirements |
|---|---|---|---|---|
| Correlation with Reference Methods | r = 0.88 with CASA [1] | r = 0.76 with AI [1] | r = 0.57 with CSA [1] | Minimum r > 0.85 with expert consensus |
| Analysis Capabilities | Unstained, live sperm; subcellular features [1] | Stained, fixed sperm only [1] | Stained sperm primarily [1] | Must maintain viability for ART use |
| Classification Accuracy | 93% overall accuracy; 95% precision for abnormal sperm [1] | Variable (53-81%) based on training [17] | Manufacturer-dependent | >90% accuracy across sperm subtypes |
| Processing Speed | 0.0056 seconds per image [1] | 4.9-9.5 seconds per image [17] | Variable | Must support clinical workflow demands |
| Inter-Method Variability | Reduced subjectivity [1] | High without standardization (CV=0.28) [17] | Moderate | CV < 0.10 for normal morphology |
The performance benchmarking process must extend beyond technical metrics to encompass clinical utility validation. This includes demonstrating improved pregnancy outcomes, enhanced embryo quality selection for intracytoplasmic sperm injection (ICSI), and correlation with DNA fragmentation indices [1] [51]. Recent surveys of fertility specialists indicate that 21.64% report regular use of AI in clinical practice, with 31.58% reporting occasional use—reflecting growing adoption despite persistent barriers including cost (38.01%) and training limitations (33.92%) [14].
Data Standardization and Annotation Protocols
The performance of deep learning models for sperm morphology analysis is fundamentally constrained by the quality, diversity, and standardization of training datasets. Current limitations in publicly available datasets represent significant barriers to clinical-grade model development.
Table 2: Current Sperm Morphology Datasets and Their Limitations for Clinical AI Development
| Dataset | Image Characteristics | Sample Size | Annotation Level | Key Limitations for Clinical Use |
|---|---|---|---|---|
| HSMA-DS [2] | 40-60× magnification | 1,475 images [1] | Morphology classification | Limited sample size, insufficient categories |
| MHSMA [1] [2] | Sperm head images | 1,540 images [1] | Head morphology focus | Exclusive head focus, no full sperm analysis |
| SVIA [1] [2] | Videos and images | 101 videos, 4,041 images [1] | Object detection, segmentation | Limited clinical correlation data |
| VISEM-Tracking [2] | Video data | 123 samples [2] | Motility and basic morphology | Limited morphological detail |
| Proprietary Clinical Datasets [1] | Confocal microscopy, Z-stack | 21,600 images [1] | Multi-frame validation | Lack of standardization, accessibility |
Expert Annotation and Ground Truth Establishment
Establishing reliable ground truth labels requires a rigorous expert consensus process analogous to methodologies used in machine learning. Studies demonstrate that expert morphologists agree on normal/abnormal classification for only 73% of sperm images when working independently [17]. This inherent subjectivity necessitates a formal consensus framework:
- Multi-expert Review Panel: Assembly of at least three certified andrology specialists with minimum 5 years of experience
- Blinded Independent Assessment: Initial classification by each expert without consultation
- Consensus Development: Discussion and reconciliation of discrepant classifications
- Quality Assurance: Validation against external reference standards when available
- Documentation: Comprehensive recording of decision rationales for model interpretation
The annotation protocol must encompass the complete sperm structure, including head (length-to-width ratio 1.5-2, vacuolation, acrosome appearance), neck (slender and regular), and tail (uniform calibre, cytoplasmic droplets <1/3 head size) according to WHO sixth edition criteria [1]. For clinical deployment, models should be validated across multiple classification systems (2-category, 5-category, 8-category, and 25-category) with demonstrated accuracy exceeding 90% for even the most complex schemas [17].
Figure 1: Expert Consensus Protocol for Ground Truth Establishment in Sperm Morphology Annotation
Regulatory Pathways and Quality Management
Evolving Regulatory Frameworks
The regulatory landscape for AI-based medical devices, including sperm analysis systems, is rapidly evolving. The European Union's AI Act categorizes reproductive medicine applications as high-risk, requiring conformity assessment, quality management system implementation, and clinical evaluation [76]. In the United States, the FDA's Digital Health Center of Excellence has established frameworks for software as a medical device (SaMD) with particular emphasis on algorithm transparency and performance consistency across diverse populations [51].
Recent expert recommendations from the French BLEFCO Group question the clinical prognostic value of traditional sperm morphology parameters before ART procedures, highlighting the need for demonstrated clinical utility rather than merely technical equivalence [10]. This shifting perspective underscores that regulatory submissions must include clinical outcome data rather than simple correlation with existing methods.
Quality Management System Requirements
Clinical deployment necessitates implementation of comprehensive quality management systems encompassing:
- Analytical Validation: Demonstration of accuracy, precision, specificity, and sensitivity across the intended use population
- Clinical Validation: Evidence of improved diagnostic outcomes or clinical decision-making
- Algorithmic Transparency: Documentation of model architecture, training data characteristics, and performance limitations
- Post-Market Surveillance: Protocols for continuous performance monitoring and model drift detection
For AI-based systems with continuous learning capabilities, regulatory frameworks require controlled update cycles with re-validation requirements and change control documentation [27]. The "black box" problem inherent in some complex deep learning models presents additional regulatory challenges, with increasing emphasis on explainable AI (XAI) approaches that provide interpretable decision support [76].
Experimental Protocols for Clinical Validation
Protocol 1: Multi-Center Performance Validation
Objective: Establish analytical and clinical performance of AI sperm morphology analysis across multiple clinical sites with diverse patient populations.
Materials and Methods:
- Samples: Collect 300 semen samples across fertility clinics, ensuring representation of diverse morphology patterns (normal, teratozoospermia, oligoasthenoteratozoospermia)
- Staining: Prepare slides using Diff-Quik stain following WHO sixth edition specifications [1]
- Imaging: Capture images using standardized microscopy protocols (100× oil immersion for reference methods; 40× confocal z-stack for AI analysis) [1]
- Reference Method: Perform blinded assessment by three independent expert morphologists following consensus protocols
- AI Analysis: Process images through the candidate AI system with pre-specified classification criteria
- Statistical Analysis: Calculate concordance statistics (Cohen's kappa), correlation coefficients, and diagnostic performance metrics with 95% confidence intervals
Validation Endpoints:
- Primary: Concordance with expert consensus for normal morphology classification (kappa >0.8)
- Secondary: Correlation with clinical outcomes (fertilization rates, embryo quality) in ART cycles
Protocol 2: Clinical Utility Assessment
Objective: Determine whether AI-based sperm morphology analysis improves clinical decision-making or ART outcomes compared to conventional methods.
Study Design: Prospective randomized controlled trial comparing standard care versus AI-informed sperm selection for ICSI.
Participants: 200 couples undergoing ICSI with male factor infertility contribution.
Intervention: Laboratory embryologists randomized to use either conventional morphology assessment or AI-based assessment for sperm selection during ICSI procedures.
Outcome Measures:
- Primary endpoint: Fertilization rate (normal fertilization assessed 16-18 hours post-insemination)
- Secondary endpoints: Embryo quality metrics (day 3 morphology, blastulation rate), clinical pregnancy rate, usable embryo rate per injected oocyte
Statistical Considerations: Power calculation based on 10% improvement in fertilization rate (80% power, α=0.05) requires 100 cycles per arm.
Implementation Framework and Integration Pathway
Successful clinical deployment requires careful attention to practical implementation challenges beyond technical validation. The following framework supports effective integration into clinical andrology workflows:
Figure 2: Clinical Deployment Pathway for AI Sperm Morphology Analysis Systems
Implementation Checklist
- Infrastructure Requirements: Compatibility with existing laboratory information systems, data storage capacity for image archives, computational resources for model inference
- Personnel Training: Certification programs for technical staff, continuing education on system limitations, result interpretation guidelines
- Quality Control Procedures: Daily verification testing, periodic comparison with reference methods, participation in external quality assurance programs
- Result Reporting Framework: Standardized report templates, interpretative comments for clinicians, uncertainty quantification in results
Essential Research Reagent Solutions
Table 3: Key Research Reagents and Materials for AI Sperm Morphology Analysis Development
| Reagent/Material | Specification | Research Function | Clinical Validation Role |
|---|---|---|---|
| Standardized Stains | Diff-Quik, Papanicolaou | Reference method establishment | Method comparison and validation |
| Slide Systems | LEJA standard two-chamber (20μm depth) [1] | Consistent sample preparation | Reproducible imaging conditions |
| Quality Control Samples | Fixed sperm suspensions with characterized morphology | Analytic performance verification | Daily quality assurance testing |
| Image Annotation Tools | LabelImg program or equivalent [1] | Ground truth establishment | Expert consensus development |
| Reference Image Sets | Curated images with expert consensus classification | Model training and testing | Ongoing competency assessment |
| Confocal Microscopy Systems | LSM 800 with Z-stack capability [1] | High-resolution image acquisition | Unstained live sperm analysis |
The clinical deployment of AI-based sperm morphology analysis systems requires a methodical, evidence-based approach that prioritizes standardization, validation, and integration within clinical workflows. By addressing the requirements outlined in this technical guide—including robust performance validation against consensus standards, comprehensive data standardization, navigation of evolving regulatory frameworks, and implementation of rigorous clinical validation protocols—researchers and developers can advance these promising technologies from research tools to clinically valuable diagnostic systems. The future of AI in sperm morphology analysis lies not merely in technical achievement but in demonstrated clinical utility that improves patient care and reproductive outcomes.
Conclusion
AI-powered sperm morphology analysis represents a paradigm shift in reproductive diagnostics, transitioning from a subjective, labor-intensive manual task to an objective, high-throughput, and data-driven process. The synthesis of this review confirms that deep learning models, particularly CNNs and transformers, consistently meet or exceed expert-level accuracy in classifying morphological defects, while also demonstrating nascent capability in predicting functional parameters like DNA integrity. Key hurdles for widespread clinical and research adoption remain, primarily the creation of large, standardized, and diverse datasets and ensuring model generalizability across different populations and imaging protocols. Future directions should focus on the development of integrated, explainable AI systems that not only classify morphology but also provide actionable insights for drug discovery, toxicology studies, and personalized treatment planning in assisted reproductive technologies, ultimately bridging the gap between seminal analysis and clinical outcomes.
References
- 1. Artificial intelligence model for the assessment of unstained ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12060770/]
- 2. Deep learning-based approach for sperm morphology analysis [https://bmcurol.biomedcentral.com/articles/10.1186/s12894-025-01946-w]
- 3. Sperm morphology: What does it mean? [https://www.mayoclinic.org/diseases-conditions/male-infertility/expert-answers/sperm-morphology/faq-20057760]
- 4. The relevance of sperm morphology in male infertility - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9580829/]
- 5. Artificial Intelligence in Andrology: From Semen Analysis to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10782130/]
- 6. Revolutionizing Sperm Analysis with AI: A Review of ... [https://www.mdpi.com/2079-3197/13/6/132]
- 7. Global, regional and national burden of male infertility in 204 ... [https://bmcpublichealth.biomedcentral.com/articles/10.1186/s12889-023-16793-3]
- 8. Global, regional and national burden of male infertility in ... [https://pubmed.ncbi.nlm.nih.gov/37940907/]
- 9. Global, regional, and national burden and trends of ... [https://www.frontiersin.org/journals/endocrinology/articles/10.3389/fendo.2025.1506229/full]
- 10. An Expert Review From the French BLEFCO Group [https://pubmed.ncbi.nlm.nih.gov/41185394/]
- 11. Design and Implementation of a Deep Learning System to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12567593/]
- 12. Validation of AI-based semen analysis by urologists in training [https://pmc.ncbi.nlm.nih.gov/articles/PMC12634335/]
- 13. AI-Guided Sperm Analysis Results In First Successful ... [https://www.drugs.com/news/ai-guided-sperm-analysis-results-first-successful-pregnancy-127418.html]
- 14. Global Trends in the Use of Artificial Intelligence (AI) in ... [https://jivfww.scholasticahq.com/article/140673-global-trends-in-the-use-of-artificial-intelligence-ai-in-reproductive-medicine-insights-from-surveys-of-international-fertility-specialists]
- 15. Assessing human sperm morphology: top models, underdogs ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3739679/]
- 16. Human live spermatozoa morphology assessment using ... [https://www.nature.com/articles/s41598-022-08798-6]
- 17. Use of a sperm morphology assessment standardisation ... [https://www.nature.com/articles/s41598-025-07515-3]
- 18. Deep learning-based approach for sperm morphology analysis [https://pmc.ncbi.nlm.nih.gov/articles/PMC12538767/]
- 19. Performance evaluation of sperm concentration, motility ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10756940/]
- 20. Synthetic Post-Contrast Imaging through Artificial Intelligence [https://pmc.ncbi.nlm.nih.gov/articles/PMC9695136/]
- 21. Validation of an Artificial Intelligence Tool for the Detection ... [https://arxiv.org/abs/2510.11142]
- 22. Frontiers | Validation of AI-based semen analysis by urologists in training: clinical impact after varicocelectomy [https://www.frontiersin.org/journals/urology/articles/10.3389/fruro.2025.1702569/full]
- 23. Unveiling the capabilities of vision transformers in sperm ... [https://peerj.com/articles/cs-3173/]
- 24. Deep-learning based model for sperm morphology ... [https://pubmed.ncbi.nlm.nih.gov/41203404/]
- 25. Unveiling the capabilities of vision transformers in sperm ... [https://pubmed.ncbi.nlm.nih.gov/40989372/]
- 26. The prospect of artificial intelligence to personalize ... [https://www.nature.com/articles/s41746-024-01006-x]
- 27. Artificial intelligence in assisted reproductive technology [https://www.sciencedirect.com/science/article/pii/S1472648325000628]
- 28. Current Updates on Involvement of Artificial Intelligence ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10890589/]
- 29. Deep learning-based approach for sperm morphology ... [https://pubmed.ncbi.nlm.nih.gov/41121112/]
- 30. Sperm morphology: Evaluating its clinical relevance in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11194684/]
- 31. Sperm morphology (shape) patient education fact sheet [https://www.reproductivefacts.org/news-and-publications/fact-sheets-and-infographics/sperm-morphology-shape-does-it-affect-fertility/]
- 32. Accuracy comparison study of new smartphone-based ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8566791/]
- 33. Deep feature engineering for accurate sperm morphology ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0330914]
- 34. Deep Learning Models for Multi-Part Morphological ... [https://www.mdpi.com/1424-8220/25/10/3093]
- 35. Super-resolution microscopy and deep learning methods ... [https://www.frontiersin.org/journals/neuroinformatics/articles/10.3389/fninf.2025.1630133/full]
- 36. Recent Advancements in Microscopy Image Enhancement ... [https://arxiv.org/html/2509.15363v2]
- 37. A multi-sperm dynamic tracking algorithm based on ... [https://www.sciencedirect.com/science/article/abs/pii/S1746809425006834]
- 38. Deep feature engineering for accurate sperm morphology ... [https://pubmed.ncbi.nlm.nih.gov/40929291/]
- 39. Sperm Morphology (Size and Shape) [https://lomalindafertility.com/infertility/men/sperm-morphology/]
- 40. Deep-learning based model for sperm morphology ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12599357/]
- 41. THE FUTURE OF SEMEN ANALYSIS: HOW ARTIFICIAL ... [https://mensreproductivehealth.com/the-future-of-semen-analysis-how-artificial-intelligence-is-transforming-andrology/]
- 42. Specific sperm morphological abnormalities and their ... [https://pubmed.ncbi.nlm.nih.gov/41204830/]
- 43. -based Apoptosis Cell Classification Using AI - Phase ... contrast Images [https://pubmed.ncbi.nlm.nih.gov/38423642/]
- 44. A Guide to Phase Contrast | Principles, Applications… [https://www.scientifica.uk.com/learning-zone/a-guide-to-phase-contrast]
- 45. Phase Contrast Microscopy - Molecular Expressions [https://micro.magnet.fsu.edu/primer/techniques/phasecontrast/phaseindex.html]
- 46. Artificial intelligence in reproductive medicine - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6733338/]
- 47. How to build high-quality annotated datasets for AI image ... [https://www.tencentcloud.com/techpedia/125210]
- 48. Data Collection and Annotation Strategies for Computer ... [https://docs.ultralytics.com/guides/data-collection-and-annotation/]
- 49. Annotation Best Practices for Building High-Quality Datasets [https://www.grammarly.com/blog/engineering/annotation-best-practices/]
- 50. Deep learning-based sperm motility and morphology ... [https://www.sciencedirect.com/science/article/pii/S2352914824000157]
- 51. The integration of artificial intelligence in assisted ... [https://www.frontiersin.org/journals/reproductive-health/articles/10.3389/frph.2025.1520919/full]
- 52. Imbalanced data classification using graph based ... [https://www.nature.com/articles/s41598-025-18804-2]
- 53. A Comprehensive Survey on Data Augmentation [https://arxiv.org/html/2405.09591v3]
- 54. Data Augmentation: Techniques That Work in Real-World ... [https://labelyourdata.com/articles/data-augmentation]
- 55. Diffusion Model-based Medical Image Generation as a ... [https://pubmed.ncbi.nlm.nih.gov/40908695/]
- 56. Exploring the potential of generative artificial intelligence in ... [https://www.sciencedirect.com/science/article/pii/S258975002500072X]
- 57. [1807.06521] CBAM: Convolutional Block Attention Module [https://arxiv.org/abs/1807.06521]
- 58. Convolutional Block Attention Module (CBAM) Overview [https://medium.com/thedeephub/convolutional-block-attention-module-cbam-overview-118e14048980]
- 59. A Comprehensive Review of Bio-Inspired Optimization ... [https://www.mdpi.com/2313-7673/8/3/278]
- 60. Bioinspired Artificial Intelligence Applications 2023 - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10886490/]
- 61. Optimization of A comprehensive dispatching system ... [https://www.nature.com/articles/s41598-025-22913-3]
- 62. An Intelligently Enhanced Ant Colony Optimization ... [https://www.mdpi.com/1424-8220/25/5/1326]
- 63. A learning-enhanced ant colony optimization algorithm for ... [https://www.sciencedirect.com/science/article/abs/pii/S2210650225000094]
- 64. Advanced Multi-Level Ensemble Learning Approaches for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12192502/]
- 65. HKUMed develops world-first AI-powered fertilisation ... [https://www.hku.hk/press/press-releases/detail/28565.html]
- 66. Enhancing the precision of male fertility diagnostics ... [https://www.nature.com/articles/s41598-025-25052-x]
- 67. A generalized classification of human sperm head ... [https://www.sciencedirect.com/science/article/pii/S0925231225011269]
- 68. ESR Essentials: common performance in metrics —practice... AI [https://link.springer.com/article/10.1007/s00330-025-11890-w]
- 69. Evaluation of an artificial intelligence system for bull sperm ... [https://www.sciencedirect.com/science/article/pii/S0093691X25002304]
- 70. Innovations in assisted reproductive technology through ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12645012/]
- 71. The Importance of Distinguishing Between Automated ... [https://www.mispeces.com/en/news/The-Importance-of-Distinguishing-Between-Automated-System-and-AI-Driven-systems/]
- 72. Deep Learning-Based Robust Automated System for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10256941/]
- 73. Standardization of bovine sperm concentration analysis in ... [https://www.sciencedirect.com/science/article/abs/pii/S0093691X25001980]
- 74. Analysis of kinetic parameters of sexed Holstein-Friesian ... [https://veterinaryworld.org/Vol.18/February-2025/5.php]
- 75. A Mobile Sperm Analyzer with User-Friendly Microfluidic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12190950/]
- 76. Editorial: Artificial intelligence in assisted reproductive ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12521430/]