Implementing Convolutional Neural Networks for Human Sperm Morphology Classification: A Comprehensive Guide for Biomedical Research
This article provides a comprehensive exploration of the implementation of Convolutional Neural Networks (CNNs) for the automated classification of human sperm morphology, a critical parameter in male fertility assessment.
Implementing Convolutional Neural Networks for Human Sperm Morphology Classification: A Comprehensive Guide for Biomedical Research
Abstract
This article provides a comprehensive exploration of the implementation of Convolutional Neural Networks (CNNs) for the automated classification of human sperm morphology, a critical parameter in male fertility assessment. Tailored for researchers, scientists, and drug development professionals, it covers the foundational motivation for automating this traditionally subjective analysis, delves into specific methodological approaches and CNN architectures, addresses common troubleshooting and optimization challenges, and presents rigorous validation and performance comparison frameworks. By synthesizing current research and clinical applications, this guide aims to equip professionals with the knowledge to develop robust, AI-driven tools that enhance the standardization, accuracy, and efficiency of semen analysis in clinical and research settings.
The Why and What: Foundations of AI in Sperm Morphology Analysis
Male factor infertility is a significant public health issue, substantially contributing to approximately 50% of all infertility cases among couples [1]. The initial and cornerstone investigation for male infertility is the semen analysis, among which sperm morphology assessment—the evaluation of sperm size, shape, and structure—is considered one of the most clinically informative yet challenging parameters [2]. Traditionally, this assessment is performed manually by technicians using microscopy, a method notoriously plagued by high subjectivity and inter-laboratory variability due to its reliance on individual expertise [2]. This manual process is slow, difficult to standardize, and can lead to inconsistent clinical diagnoses.
The integration of Convolutional Neural Networks (CNNs), a class of deep learning algorithms, presents a paradigm shift for andrology laboratories. CNNs are uniquely suited for image analysis tasks as they can learn hierarchical features directly from pixel data, automating the classification process and minimizing human bias [3] [4]. This document outlines the application of a CNN-based framework for the standardization of human sperm morphology classification, detailing the experimental protocols, data handling procedures, and technical specifications required for robust implementation.
The following tables summarize the key quantitative aspects of developing a CNN model for sperm morphology classification, from dataset composition to model performance.
Table 1: SMD/MSS Dataset Composition and Augmentation
| Component | Description | Quantity |
|---|---|---|
| Initial Image Collection | Individual sperm images acquired via MMC CASA system (100x oil immersion) | 1,000 images [2] |
| Data Augmentation | Application of techniques to create variant images (e.g., rotation, scaling) to balance classes and increase dataset size | Final dataset: 6,035 images [2] |
| Expert Classification | Three independent experts classifying based on modified David criteria (12 defect classes) | 3 experts per image [2] |
| Inter-Expert Agreement | Percentage of images where all three experts assigned identical labels for all categories | "Total Agreement" (TA) on a subset of images [2] |
Table 2: CNN Model Configuration and Performance Metrics
| Parameter | Specification | Value / Range |
|---|---|---|
| Programming Environment | Language and key libraries | Python 3.8 [2] |
| Image Pre-processing | Resizing, normalization, denoising | 80x80 pixels, grayscale [2] |
| Data Partitioning | Train/Test split | 80% Training, 20% Testing [2] |
| Reported Model Accuracy | Performance on the test set | 55% to 92% [2] |
Experimental Protocols & Workflows
Protocol 1: Sample Preparation and Data Acquisition
This protocol ensures the consistent creation of high-quality sperm image smears for subsequent digitization.
Materials:
- Fresh semen sample (sperm concentration ≥ 5 million/mL and < 200 million/mL) [2]
- RAL Diagnostics staining kit or equivalent [2]
- Microscope slides and coverslips
- Optical microscope with 100x oil immersion objective and digital camera (e.g., MMC CASA system) [2]
Procedure:
- Smear Preparation: Prepare a thin smear of the semen sample on a clean glass slide, following the guidelines outlined in the WHO laboratory manual [2].
- Staining: Apply the RAL Diagnostics stain according to the manufacturer's instructions to enhance cellular contrast and detail.
- Image Acquisition: Using the MMC CASA system or equivalent, capture images of individual spermatozoa with the 100x oil immersion objective in bright-field mode.
- Data Export: Ensure each captured image contains a single spermatozoon. Save images in a standard format (e.g., PNG, JPEG) and assign a unique filename.
Protocol 2: Image Labeling and Ground Truth Establishment
This protocol defines the process for creating a reliable "ground truth" dataset, which is critical for supervised learning.
Materials:
- Acquired sperm images (from Protocol 1)
- Standardized classification form (e.g., Excel spreadsheet) based on modified David classification [2]
Procedure:
- Expert Panel: Provide the set of images to three independent experts, each with extensive experience in semen analysis.
- Blinded Classification: Each expert independently classifies every spermatozoon into one or more of the 12 morphological classes defined by the modified David criteria (e.g., tapered head, microcephalous, bent midpiece, coiled tail) [2].
- Data Consolidation: Compile all classifications from the experts into a single ground truth file. This file should link each image filename to the classifications from all three experts and any associated morphometric data.
- Consensus Analysis: Analyze the level of inter-expert agreement. Images with total agreement (TA) among all three experts provide the highest confidence labels for model training.
Protocol 3: CNN Model Development and Training
This protocol covers the computational steps for building and training the deep learning model.
Materials:
- Hardware: Computer with a dedicated Graphics Processing Unit (GPU)
- Software: Python 3.8 with deep learning libraries (e.g., TensorFlow, PyTorch, Keras) [2] [4]
Procedure:
- Image Pre-processing:
- Resize: Scale all images to a uniform size of 80x80 pixels using a linear interpolation strategy.
- Grayscale Conversion: Convert color images to single-channel grayscale to simplify the initial model input.
- Normalization: Normalize pixel intensity values to a range of 0 to 1 to aid model convergence. [2]
- Data Partitioning: Randomly split the entire dataset of 6,035 images into a training set (80% of data) for model learning and a hold-out test set (20% of data) for final performance evaluation. [2]
- Model Training:
- Architecture Definition: Design a CNN architecture comprising convolutional layers for feature extraction, pooling layers for dimensionality reduction, and fully connected layers for final classification. [3] [4]
- Compilation: Define a loss function (e.g., categorical cross-entropy) and an optimizer (e.g., Adam).
- Iteration: Train the model by iteratively presenting batches of images from the training set, adjusting the model's internal weights to minimize classification error.
Visualization of Workflows
The following diagrams, generated with Graphviz DOT language, illustrate the logical relationships and workflows described in the protocols.
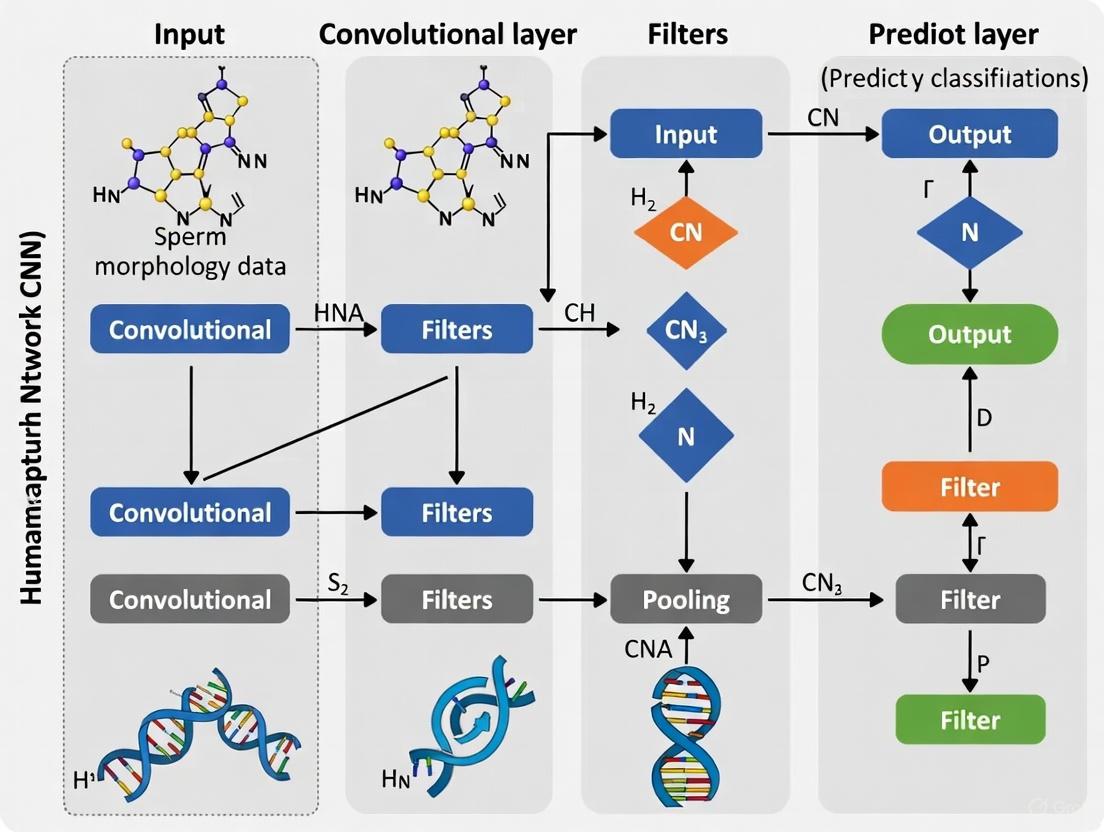
Sperm Morphology Analysis Workflow
Convolutional Neural Network Architecture
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials and Reagents for CNN-based Sperm Morphology Analysis
| Item | Function / Application |
|---|---|
| MMC CASA System | An integrated hardware and software system for the automated, sequential acquisition of images from sperm smears using a microscope-equipped camera. [2] |
| RAL Diagnostics Staining Kit | A ready-to-use staining solution used to prepare semen smears for morphological analysis, enhancing the contrast and visibility of sperm structures under a microscope. [2] |
| Modified David Classification Sheet | A standardized form detailing 12 specific classes of sperm defects (affecting the head, midpiece, and tail) used by experts to generate consistent ground truth labels. [2] |
| GPU-Accelerated Computing Workstation | A computer equipped with a dedicated Graphics Processing Unit (GPU) essential for performing the vast number of calculations required to train deep learning models in a feasible timeframe. [4] |
| Python with Deep Learning Libraries (TensorFlow/PyTorch) | The core programming environment and software libraries that provide the tools and functions necessary to define, train, and evaluate convolutional neural network models. [2] [4] |
The diagnostic evaluation of male infertility relies heavily on semen analysis, with sperm morphology assessment representing one of its most prognostically significant yet challenging components. For decades, this analysis has been performed manually by trained technicians observing stained sperm smears under a microscope, a method subject to significant subjectivity and variability. The introduction of Computer-Assisted Semen Analysis (CASA) systems promised to revolutionize the field by introducing automation, objectivity, and standardization. However, current CASA methodologies exhibit considerable limitations that impact their reliability and clinical utility, particularly in morphological assessment. This application note critically examines the limitations inherent in both manual and conventional CASA approaches, contextualized within the framework of emerging convolutional neural network (CNN) technologies that offer potential solutions to these longstanding challenges.
Manual Sperm Morphology Assessment: Inherent Subjectivity and Variability
Fundamental Methodological Constraints
Traditional manual sperm morphology assessment follows World Health Organization (WHO) guidelines, requiring technicians to classify at least 200 spermatozoa into normal and abnormal categories based on strict morphological criteria. The process involves staining semen smears (typically with RAL, Papanicolaou, or Diff-Quik stains) and systematic examination under high-power magnification (100x oil immersion) [2]. Despite standardized protocols, this method suffers from inherent limitations:
- Subjectivity in Classification: Borderline morphological features often receive different classifications between technicians, even within the same laboratory.
- Cognitive Fatigue: The visual intensity of scanning and classifying hundreds of sperm cells leads to declining accuracy over time.
- Inter-laboratory Variability: Differences in staining techniques, microscope calibration, and technician training create substantial inconsistencies between facilities.
Quantitative Evidence of Manual Method Limitations
Table 1: Documented Variability in Manual Sperm Morphology Assessment
| Parameter | Evidence of Variability | Impact on Diagnostic Reliability |
|---|---|---|
| Inter-observer Agreement | Kappa values as low as 0.05-0.15 reported between technicians [5] | Poor diagnostic reproducibility even among trained experts |
| Time Consumption | 30-45 minutes per sample for proper assessment [5] | Practical limitations in high-volume clinical settings |
| Disagreement Rate | Up to 40% coefficient of variation between evaluators [5] | Significant potential for misclassification and diagnostic error |
Computer-Assisted Semen Analysis (CASA): Persistent Technological Limitations
Conventional CASA systems utilize optical microscopy coupled with digital cameras and specialized software to capture and analyze sperm images. The general workflow involves:
- Sample Preparation: Semen samples are loaded into specialized chambers (e.g., Makler, MicroCell) of standardized depth
- Image Acquisition: Multiple digital images or videos are captured under phase-contrast or bright-field microscopy
- Software Analysis: Proprietary algorithms identify sperm cells, distinguish them from debris, and calculate parameters
- Data Output: Quantitative results for concentration, motility, and in some systems, morphological metrics
Despite four decades of technological evolution, current CASA systems face significant challenges in accurate morphological classification due to fundamental limitations in image analysis capabilities and algorithmic approaches.
Specific Limitations of Conventional CASA Systems
Table 2: Documented Limitations of Conventional CASA Systems in Morphology Assessment
| Limitation Category | Specific Technical Challenges | Impact on Analysis |
|---|---|---|
| Image Resolution & Quality | Limited ability to distinguish subtle morphological features; difficulty with overlapping sperm or debris-filled samples [6] [2] | Inaccurate detection and classification of abnormal forms |
| Algorithmic Constraints | Inability to properly classify midpiece and tail abnormalities; poor performance with complex defects [2] | Systematic under-reporting of specific abnormality types |
| Standardization Issues | High sensitivity to instrument settings (illumination, contrast, chamber depth) [7] | Poor inter-system reproducibility and comparability |
| Concentration Dependency | Increased variability in low (<15 million/mL) and high (>60 million/mL) concentration specimens [6] | Restricted reliable operating range |
| Morphological Heterogeneity | Difficulty handling the natural shape variation within samples and subjects [6] | Oversimplification of complex morphological patterns |
Experimental Protocols for Methodology Comparison
Protocol 1: Manual Morphology Assessment (WHO Standard)
Principle: Visual classification of stained spermatozoa based on standardized morphological criteria.
Materials:
- RAL, Papanicolaou, or Diff-Quik staining kits
- Microscope with 100x oil immersion objective
- Tally counters or specialized data entry software
- Standardized data collection forms
Procedure:
- Prepare thin semen smears on clean glass slides and air dry
- Fix and stain according to manufacturer protocols for selected stain
- Systematically scan slides using meander pattern at 100x magnification
- Classify each of 200+ consecutive spermatozoa into:
- Normal
- Head defects (tapered, thin, microcephalic, macrocephalic, multiple, abnormal acrosome)
- Midpiece defects (bent, cytoplasmic droplet)
- Tail defects (coiled, short, multiple, broken)
- Calculate percentage of normal forms and specific defect categories
- Record all data with quality control documentation
Quality Control: Participation in external quality assurance programs; regular inter-technician comparison exercises [8].
Protocol 2: Conventional CASA Morphology Analysis
Principle: Automated image capture and analysis of sperm morphological parameters.
Materials:
- CASA system (e.g., Hamilton Thorne IVOS/CEROS, SCA, SQA-V)
- Disposable counting chambers (e.g., Leja, MicroCell)
- Quality control beads (e.g., latex Accu-Beads) [6]
- Temperature-controlled stage (if assessing motility)
Procedure:
- Calibrate system using quality control beads according to manufacturer specifications
- Pre-warm counting chamber and stage to 37°C if analyzing motility
- Load appropriately diluted semen sample (following manufacturer recommendations)
- Set acquisition parameters:
- Number of fields to analyze (minimum 10)
- Sperm detection thresholds (size, intensity, contrast)
- Morphology classification criteria (based on WHO standards)
- Execute automated acquisition and analysis
- Review results for artifacts or misclassifications; manually override if necessary
- Generate and export comprehensive report
Quality Control: Regular calibration and validation; standardized operating procedures for all technicians; documentation of all instrument settings [7].
Visualizing Methodological Limitations and Solutions
Comparative Workflow: Traditional vs. AI-Enhanced Analysis
Technical Pathway: CASA System Operation and Failure Points
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Research Reagent Solutions for Sperm Morphology Analysis
| Reagent/Material | Function/Application | Specific Examples & Notes |
|---|---|---|
| Staining Kits | Cellular staining for manual morphology assessment | RAL Diagnostics kit [2], Papanicolaou, Diff-Quik |
| Standardized Chambers | Consistent sample depth for analysis | Leja 20μm chambers [7], MicroCell, Makler |
| Quality Control Beads | System calibration and validation | Latex Accu-Beads [6] |
| CASA Systems | Automated semen analysis | Hamilton Thorne IVOS/CEROS [6], SCA Microptics [6] |
| Dataset Images | Algorithm training and validation | HuSHeM (216 images) [5], SCIAN (1,854 images) [9], SMIDS (3,000 images) [5] |
| Deep Learning Frameworks | CNN model development | YOLOv7 [10], VGG16 [11], ResNet50 [5], DenseNet169 [12] |
The limitations of current sperm morphology assessment methodologies—both manual and CASA-based—represent significant challenges in male infertility diagnostics. Manual methods suffer from irreproducible subjectivity and substantial inter-observer variability, while conventional CASA systems demonstrate inadequate performance in morphological classification, particularly for complex defects and challenging samples. These limitations necessitate technological innovation, with deep learning approaches—particularly CNN-based architectures—emerging as promising solutions. The integration of AI technologies offers the potential to overcome longstanding limitations through automated, standardized, and highly accurate sperm morphology classification, ultimately advancing both clinical diagnostics and research capabilities in reproductive medicine.
The morphological evaluation of human sperm is a cornerstone of male fertility assessment, providing critical diagnostic and prognostic information. Traditional manual analysis, however, is notoriously subjective, time-consuming, and plagued by significant inter-observer variability, with reported disagreement rates among experts reaching up to 40% [5]. This lack of standardization directly impacts the reliability of infertility diagnostics and treatment planning.
Convolutional Neural Networks (CNNs) offer a powerful solution to these challenges by enabling the automation, standardization, and acceleration of sperm morphology analysis. This document outlines the foundational classification task for a CNN-based system, defining the core categories of "Normal" versus "Abnormal" and detailing the key morphological defects that the model must learn to identify. By establishing a clear and consistent classification framework, researchers can develop robust models that enhance objectivity and reproducibility in reproductive medicine [2] [11].
Defining the Classification Classes
The primary task for a CNN in sperm morphology analysis is a classification problem. The system must analyze an input image of an individual spermatozoon and assign it to one of several predefined categories. These categories are hierarchically organized, starting with the broad distinction between normal and abnormal forms, followed by a more granular classification of specific defect types and their locations.
The "Normal" Spermatozoon
A morphologically normal spermatozoon is the reference point for all classification. According to World Health Organization (WHO) guidelines, it is characterized by the following features [5]:
- Head: Smooth, oval configuration with a well-defined acrosome covering 40-70% of the head area. The length should be between 4.0–5.5 µm and the width between 2.5–3.5 µm.
- Midpiece: Slender, approximately the same length as the head, and axially attached.
- Tail: A single, unbroken tail that is thinner than the midpiece and approximately 45 µm long, without sharp bends or coils.
Any deviation from this strict definition qualifies the sperm as abnormal. In clinical practice, a sample with ≥ 4% normal forms is generally considered within the normal range, though this threshold can vary [13].
The "Abnormal" Spermatozoon: A Framework of Defects
Abnormal sperm are categorized based on the specific part of the sperm cell that is defective. The most comprehensive systems, such as the modified David classification, define numerous specific anomaly types [2]. For a CNN-based system, these can be consolidated into a structured hierarchy of defects.
Table 1: Key Morphological Defects for CNN Classification
| Defect Category | Specific Defect Types | Key Morphological Characteristics |
|---|---|---|
| Head Defects | Tapered, Thin, Microcephalous (small), Macrocephalous (large), Multiple heads, Abnormal acrosome, Abnormal post-acrosomal region [2] | Irregular head shape (pyriform, round, amorphous), vacuolization, size discrepancies, disordered acrosome [9] |
| Midpiece Defects | Bent midpiece, Cytoplasmic droplet [2] | Thickened, asymmetrical, or bent midpiece; presence of a cytoplasmic remnant >1/3 the head size [9] |
| Tail Defects | Coiled tail, Short tail, Multiple tails [2] | Absent, broken, coiled, or multiple tails; sharp angular bends [9] |
It is common for a single spermatozoon to exhibit multiple defects across different compartments (e.g., a microcephalic head with a coiled tail). This is classified as a sperm with associated anomalies [2]. Some studies also include a distinct "Non-Sperm" class to identify cellular debris or other artifacts that are not sperm cells, which is crucial for reducing false positives in an automated system [14].
Quantitative Performance of CNN Models
Deep learning approaches have demonstrated significant success in automating the classification task. Performance varies based on the model architecture, dataset size and quality, and the specific classification scheme used.
Table 2: Reported Performance of Selected CNN Models for Sperm Morphology Classification
| Model Architecture / Approach | Dataset(s) Used | Reported Performance | Key Highlights |
|---|---|---|---|
| CBAM-enhanced ResNet50 with Deep Feature Engineering [5] | SMIDS (3-class), HuSHeM (4-class) | Accuracy: 96.08% (SMIDS), 96.77% (HuSHeM) | Integrates attention mechanisms; uses feature selection & SVM classifier. |
| Multi-model CNN Fusion (Soft-Voting) [14] | SMIDS, HuSHeM, SCIAN-Morpho | Accuracy: 90.73% (SMIDS), 85.18% (HuSHeM), 71.91% (SCIAN) | Fuses six different CNN models for robust prediction. |
| VGG16 with Transfer Learning [11] | HuSHeM, SCIAN | True Positive Rate: 94.1% (HuSHeM), 62% (SCIAN) | Applies transfer learning from ImageNet, avoiding manual feature extraction. |
| Custom CNN [2] | SMD/MSS (12-class) | Accuracy: 55% to 92% (varies by class) | Based on the modified David classification with 12 detailed defect classes. |
Experimental Protocol for CNN-Based Classification
The following protocol provides a detailed methodology for developing and validating a CNN model for human sperm morphology classification, synthesizing best practices from recent literature.
Sample Preparation and Image Acquisition
- Sample Preparation: Collect semen samples after obtaining informed consent. Prepare smears on glass slides according to WHO guidelines [2]. Stain using a standardized protocol (e.g., RAL Diagnostics staining kit) to ensure consistent contrast and nuclear/morphological detail [2].
- Image Acquisition: Use an optical microscope equipped with a high-resolution digital camera and a 100x oil immersion objective [2]. The CASA (Computer-Assisted Semen Analysis) system's morphometric tool can be utilized to capture images and initially determine basic dimensions of the head and tail [2]. Capture images of individual spermatozoa, ensuring they are well-separated to avoid overlap.
Data Preprocessing and Annotation
- Image Preprocessing:
- Cleaning: Handle missing values or inconsistent data.
- Normalization: Resize images to a uniform dimension (e.g., 80x80 pixels) and convert to grayscale to standardize input and reduce computational load [2].
- Denoising: Apply techniques like wavelet denoising to remove noise from insufficient lighting or poor staining, which can improve model accuracy [2] [15].
- Expert Annotation (Ground Truth Creation): Each sperm image must be independently classified by multiple experienced embryologists or technicians (e.g., three experts) [2]. The annotation should follow a standardized classification system (e.g., WHO criteria or modified David classification). A ground truth file is compiled, listing the image name and the classifications from all experts [2].
- Data Augmentation: To balance morphological classes and increase dataset size, apply augmentation techniques such as rotation, flipping, scaling, and changes in brightness and contrast. For example, one study expanded a dataset from 1,000 to 6,035 images through augmentation [2].
Model Training and Evaluation
- Dataset Partitioning: Randomly split the entire dataset into a training set (80%) and a testing set (20%) [2]. A portion of the training set (e.g., 20%) can be used as a validation set for hyperparameter tuning.
- Model Selection and Training:
- Architecture Choice: Select a CNN architecture (e.g., ResNet50, VGG16, custom CNN) [14] [11] [5].
- Transfer Learning: Consider using a pre-trained model (on datasets like ImageNet) and fine-tune it on the sperm morphology dataset, which can be effective, especially with limited data [11].
- Training: Train the model using the training set. Employ a suitable optimizer (e.g., Adam) and a loss function (e.g., categorical cross-entropy).
- Model Evaluation:
- Metrics: Evaluate the model on the held-out test set using metrics such as Accuracy, True Positive Rate (Sensitivity), Precision, and F1-Score [11] [5].
- Validation Technique: Use k-fold cross-validation (e.g., k=5 or k=10) to ensure the model's performance is consistent and not dependent on a particular data split [14] [16].
- Comparison to Baseline: Compare the model's performance (e.g., using Mean Absolute Error) against a simple baseline model (ZeroR) to demonstrate its predictive power [16].
The following workflow diagram summarizes the complete experimental pipeline:
CNN-Based Sperm Morphology Classification Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials and Reagents for Sperm Morphology Analysis Research
| Item | Function / Application | Examples / Specifications |
|---|---|---|
| Staining Kits | Provides contrast for microscopic examination of sperm structures (head, acrosome, midpiece). | RAL Diagnostics kit [2] |
| Public Datasets | Benchmarks for training, validating, and comparing CNN models. | SMIDS, HuSHeM, SCIAN-MorphoSpermGS, SMD/MSS Dataset [2] [14] [5] |
| Deep Learning Frameworks | Software libraries for building, training, and deploying CNN models. | Python with TensorFlow/Keras or PyTorch [2] [16] |
| Microscopy Systems | Image acquisition for creating new datasets or validating model predictions. | Microscope with 100x oil objective, digital camera, CASA system [2] |
| Pre-trained Models | Accelerates development via transfer learning, improving performance with limited data. | VGG16, ResNet-50, InceptionV3 [11] [16] [5] |
Core Datasets in the Field: SMD/MSS, MHSMA, VISEM, and HuSHeM
The implementation of Convolutional Neural Networks (CNNs) for human sperm morphology classification represents a paradigm shift in male fertility research, offering a path to standardize a traditionally subjective and variable analysis. The robustness of any deep learning model is intrinsically linked to the quality, size, and diversity of the dataset used for its training. This application note provides a detailed examination of four core public datasets—SMD/MSS, MHSMA, VISEM-Tracking, and HuSHeM—that are pivotal for developing and benchmarking CNN-based sperm morphology analysis systems. We present structured quantitative comparisons, detailed experimental protocols for dataset utilization, and a scientist's toolkit to guide researchers in selecting and applying these resources effectively within a computational andrology framework.
Core Dataset Specifications and Quantitative Comparison
A critical first step in experimental design is the selection of an appropriate dataset. The core datasets vary significantly in their focus, encompassing static morphology from stained samples and dynamic motility from live video recordings. The quantitative specifications and primary applications of the SMD/MSS, MHSMA, VISEM-Tracking, and HuSHeM datasets are summarized in Table 1.
Table 1: Quantitative Comparison of Core Sperm Morphology and Motility Datasets
| Dataset Name | Primary Focus | Original Sample Size | Augmented/Extended Size | Annotation & Classification Standard | Key Strengths |
|---|---|---|---|---|---|
| SMD/MSS [2] [17] | Static Morphology | 1,000 images | 6,035 images (after augmentation) | Modified David Classification (12 defect classes) by 3 experts | Comprehensive defect annotation across head, midpiece, and tail; expert consensus |
| MHSMA [18] | Static Morphology | 1,540 images | Not specified | WHO-based guidelines for head, acrosome, and vacuole defects | Freely available; benchmark for head/acrosome/vacuole classification |
| VISEM-Tracking [19] | Motility & Tracking | 20 videos (29,196 frames) | 166 additional unlabeled video clips | Bounding boxes with tracking IDs; labels: normal, pinhead, cluster | Rich motility data; manually annotated tracking coordinates |
| HuSHeM [20] | Static Morphology (Head) | Not specified in detail | Not specified | Five head morphology categories (e.g., normal, tapered, pyriform) | Focused on sperm head morphology classification |
Diagram 1: Logical relationship between dataset type and CNN model development focus
Diagram 1 Title: Dataset Type Drives CNN Application Focus
Detailed Experimental Protocols
Protocol 1: Implementing a CNN for Morphology Classification Using SMD/MSS
The SMD/MSS dataset, with its detailed annotations based on the modified David classification, is ideal for training a CNN to perform multi-class defect identification [2].
Sample Preparation & Image Acquisition (as per SMD/MSS protocol):
- Prepare semen smears from samples with a concentration of at least 5 million/mL, excluding samples >200 million/mL to prevent image overlap.
- Stain smears using the RAL Diagnostics staining kit, following WHO guidelines.
- Acquire images of individual spermatozoa using an MMC CASA system, employing a bright field mode with an oil immersion x100 objective [2].
Data Pre-processing for CNN Input:
- Data Cleaning: Inspect images for artifacts and inconsistencies. The SMD/MSS protocol involves handling missing values and outliers to ensure data quality.
- Normalization: Resize all images to a consistent dimension (e.g., 80x80 pixels) and convert to grayscale. Normalize pixel values to a common scale, e.g., [0, 1], to stabilize and accelerate CNN training [2].
- Data Partitioning: Randomly split the augmented dataset of 6,035 images into training (80%), validation (10%), and testing (10%) subsets. Ensure stratification to maintain class distribution across splits [2].
CNN Training & Evaluation:
- Architecture Selection: Implement a CNN architecture such as ResNet50, which has demonstrated effectiveness on this dataset [21]. The network should culminate in a softmax output layer corresponding to the 12 morphological classes plus a 'normal' class.
- Training Configuration: Use categorical cross-entropy loss and the Adam optimizer. Mitigate overfitting by employing techniques like dropout and data augmentation (e.g., rotations, flips) beyond the initial dataset augmentation.
- Performance Metrics: Evaluate the model on the held-out test set using accuracy, precision, recall, and F1-score. The SMD/MSS study reported accuracies ranging from 55% to 92% across different morphological classes, reflecting the varying complexity of the classification task [2] [17].
Protocol 2: Sperm Detection and Motility Analysis Using VISEM-Tracking
The VISEM-Tracking dataset enables the development of models for sperm detection and movement analysis in video sequences, a key step towards automated CASA systems [19].
Data Acquisition (as per VISEM-Tracking protocol):
- Place unstained semen samples on a heated microscope stage (37°C) and examine under 400x magnification using a microscope with phase-contrast optics.
- Record videos using a microscope-mounted camera (e.g., IDS UI-2210C). VISEM-Tracking provides 20 annotated videos of 30 seconds each, saved as AVI files [19].
Data Pre-processing and Annotation:
- Frame Extraction: Decompose videos into individual frames for processing.
- Bounding Box Formatting: The dataset provides annotations in YOLO format, which includes normalized bounding box coordinates and class labels (0: normal sperm, 1: sperm clusters, 2: small/pinhead sperm) [19].
YOLO Model for Detection and Tracking:
- Model Training: Train a YOLO-based object detection model (e.g., YOLOv5 or YOLOv7) on the annotated frames. This model will learn to localize and classify spermatozoa in each frame [19] [10].
- Tracking Implementation: Utilize the provided tracking identifiers to link detections of the same sperm across consecutive frames. This allows for the computation of kinematic parameters (e.g., velocity, linearity).
- Baseline Performance: The dataset publishers established a baseline using YOLOv5, demonstrating the dataset's suitability for training complex deep learning models for sperm detection [19].
Diagram 2: Workflow for CNN-based Sperm Morphology Classification
Diagram 2 Title: End-to-End Morphology Classification Workflow
The Scientist's Toolkit: Research Reagent Solutions
Successful implementation of the protocols above requires both wet-lab materials and computational tools. The following table details key reagents, software, and datasets essential for research in this field.
Table 2: Essential Research Reagents and Resources for Automated Sperm Analysis
| Category | Item / Resource | Specification / Example | Primary Function in Research |
|---|---|---|---|
| Wet-Lab Reagents | RAL Diagnostics Staining Kit | As used in SMD/MSS protocol [2] | Provides contrast for detailed morphological analysis of sperm structures in static images. |
| Optixcell Extender | As used in bovine studies [10] | Preserves sperm viability and morphology during sample preparation and imaging. | |
| Non-Capacitating / Capacitating Media | As used in 3D-SpermVid dataset [22] | Enables study of sperm motility under different physiological conditions. | |
| Software & Tools | Python with Deep Learning Libraries | Python 3.8, PyTorch/TensorFlow [2] | Core programming environment for building, training, and evaluating CNN models. |
| YOLO Framework | YOLOv5, YOLOv7 [19] [10] | Real-time object detection and tracking of sperm in video sequences. | |
| LabelBox | Commercial annotation tool [19] | Facilitates manual annotation of bounding boxes for creating ground-truth datasets. | |
| Public Datasets | SMD/MSS, MHSMA, VISEM-Tracking | See Table 1 | Benchmark datasets for training and validating models on morphology and motility. |
| Synthetic Data | AndroGen Software | Open-source synthetic generator [23] | Generates customizable, annotated sperm images to augment real datasets and address data scarcity. |
Discussion and Future Perspectives
The reviewed datasets provide foundational resources for automating sperm analysis, yet they present complementary strengths and limitations. SMD/MSS offers exceptional morphological detail via expert annotation but is limited to static images [2]. Conversely, VISEM-Tracking provides rich motility data but less granular morphological classification [19]. MHSMA is a valuable, publicly available benchmark, though it may have limitations in resolution and sample size [18]. A significant challenge across the field is the lack of standardized, high-quality annotated datasets, which is crucial for developing robust, generalizable models [20].
Future research will likely focus on multi-dimensional datasets that combine high-resolution morphology with 3D motility tracking, as seen in emerging resources like the 3D-SpermVid dataset [22]. Furthermore, to combat data scarcity, the use of synthetic data generation tools like AndroGen provides a promising avenue to create large, balanced, and annotated datasets for training more accurate models without privacy concerns [23]. The clinical integration of these AI tools is advancing, with recent expert reviews providing a positive opinion on their use after rigorous qualification and validation within individual laboratories [13]. This progression from bespoke manual analysis to standardized, AI-driven pipelines heralds a new era of objectivity and efficiency in male fertility assessment.
The Role of Deep Learning and CNNs in Revolutionizing Reproductive Biology
Infertility represents a significant global health challenge, affecting approximately 15% of couples worldwide, with male factors contributing to nearly half of all cases [2] [20]. The morphological analysis of sperm remains a cornerstone in male fertility assessment, providing critical diagnostic and prognostic value for natural conception and assisted reproductive technologies (ART) [24] [25]. Traditional manual sperm morphology assessment, however, suffers from substantial limitations including subjectivity, extensive time requirements (30-45 minutes per sample), and significant inter-observer variability with reported disagreement rates reaching up to 40% among experts [2] [5].
The emergence of deep learning, particularly convolutional neural networks (CNNs), is transforming reproductive biology by introducing automated, standardized, and highly accurate analytical capabilities [24]. These artificial intelligence technologies demonstrate remarkable potential to exceed human expert performance in sperm classification tasks, offering improved reliability, throughput, and diagnostic consistency across laboratories [11] [25]. This paradigm shift addresses fundamental challenges in reproductive medicine while opening new avenues for precise male fertility assessment.
Deep Learning Architectures for Sperm Analysis
Convolutional Neural Network Fundamentals
CNNs represent a specialized class of deep neural networks particularly suited for processing structured grid data such as images [11]. Their architecture typically consists of multiple convolutional layers that automatically learn hierarchical feature representations directly from raw pixel data, followed by pooling layers for spatial invariance, and fully-connected layers for final classification [26] [5]. This endogenous feature learning capability eliminates the need for manual feature engineering, allowing CNNs to discern subtle morphological patterns often imperceptible to human observers [11].
Specialized Architectures and Performance
Recent research has investigated numerous CNN architectures optimized for sperm analysis, demonstrating exceptional classification performance across various morphological parameters:
Table 1: Performance of Deep Learning Models in Sperm Morphology Classification
| Architecture | Dataset | Classes | Performance | Reference |
|---|---|---|---|---|
| VGG16 (Transfer Learning) | HuSHeM | 5 WHO categories | 94.1% TPR | [11] |
| Custom CNN | SMD/MSS | 12 David classes | 55-92% Accuracy | [2] |
| CBAM-ResNet50 + DFE | SMIDS | 3-class | 96.08% Accuracy | [5] |
| CBAM-ResNet50 + DFE | HuSHeM | 4-class | 96.77% Accuracy | [5] |
| Sequential DNN | MHSMA | Head/Vacuole/Acrosome | 89-92% Accuracy | [26] |
| Specialized CNN | SCIAN | 5 WHO categories | 88% Recall | [25] |
The integration of attention mechanisms with traditional CNNs represents a significant advancement. The Convolutional Block Attention Module (CBAM) enhanced ResNet50 architecture sequentially applies channel-wise and spatial attention to feature maps, enabling the network to focus on diagnostically relevant sperm structures while suppressing irrelevant background information [5]. When combined with deep feature engineering pipelines incorporating multiple feature selection methods, this approach has achieved state-of-the-art performance with accuracy improvements of 8.08-10.41% over baseline CNN models [5].
Experimental Protocols for CNN-Based Sperm Morphology Classification
Dataset Curation and Preparation
Protocol 1: SMD/MSS Dataset Development [2]
Sample Collection and Preparation: Collect semen samples from patients with sperm concentration ≥5 million/mL. Prepare smears following WHO manual guidelines and stain with RAL Diagnostics staining kit.
Image Acquisition: Capture individual sperm images using MMC CASA system with bright field mode under oil immersion at 100x objective magnification. Ensure each image contains a single spermatozoon with clearly visible head, midpiece, and tail structures.
Expert Annotation and Ground Truth Establishment: Engage three independent experts with extensive semen analysis experience to classify each spermatozoon according to modified David classification (12 morphological classes). Resolve disagreements through consensus review.
Data Augmentation and Balancing: Apply transformation techniques including rotation, scaling, and flipping to address class imbalance. Expand original dataset from 1,000 to 6,035 images to enhance model generalizability.
Protocol 2: Deep Feature Engineering Pipeline [5]
Backbone Feature Extraction:
- Implement CBAM-enhanced ResNet50 architecture pre-trained on ImageNet
- Extract multi-level feature representations from CBAM, Global Average Pooling (GAP), Global Max Pooling (GMP), and pre-final layers
Feature Selection and Optimization:
- Apply multiple feature selection methods including Principal Component Analysis (PCA), Chi-square test, Random Forest importance, and variance thresholding
- Evaluate intersection combinations of selection methods to identify optimal feature subsets
Classification:
- Implement Support Vector Machines with RBF and Linear kernels
- Utilize k-Nearest Neighbors algorithms as complementary classifiers
- Apply 5-fold cross-validation for robust performance estimation
Model Training and Validation
Protocol 3: Transfer Learning Implementation [11]
Network Adaptation:
- Modify VGG16 architecture by replacing final fully-connected layers with task-specific classifiers
- Maintain pre-trained weights from ImageNet for initial feature detection capabilities
Progressive Fine-Tuning:
- Initially freeze early layers, training only replacement layers for 100 epochs
- Gradually unfreeze and fine-tune intermediate layers with reduced learning rates (0.0004)
- Employ Adam optimizer with default parameters from Keras Python package
Validation Strategy:
- Implement strict train-test splits (80-20%)
- Utilize ten-fold cross-validation to account for limited dataset sizes
- Apply early stopping with 15-epoch patience to prevent overfitting
Diagram 1: Sperm Morphology Analysis Workflow
Table 2: Essential Research Reagents and Computational Resources
| Resource Category | Specific Examples | Function/Application | Implementation Notes |
|---|---|---|---|
| Public Datasets | HuSHeM [11], SCIAN [25], SMD/MSS [2], MHSMA [26], VISEM [27] | Model training and benchmarking | Annotated by domain experts; Varied classification schemes (WHO/David) |
| Staining Reagents | RAL Diagnostics staining kit [2] | Sperm structural visualization | Follow WHO manual protocols for consistent results |
| Imaging Systems | MMC CASA System [2] | Digital image acquisition | 100x oil immersion objective; Bright field mode |
| CNN Architectures | VGG16 [11], ResNet50 [5], Custom CNN [2], Sequential DNN [26] | Feature extraction and classification | Transfer learning from ImageNet; Attention mechanisms (CBAM) |
| Data Augmentation | Rotation, scaling, flipping [2] | Dataset expansion and balancing | Address class imbalance; Improve model generalization |
| Programming Tools | Python 3.8 [2], Keras [16] | Algorithm implementation | Open-source libraries; GPU acceleration support |
Analytical Framework and Validation Methodologies
Performance Metrics and Statistical Validation
Rigorous validation constitutes an essential component of CNN implementation for sperm morphology classification. Established metrics include true positive rate (TPR), accuracy, recall, and mean absolute error (MAE) [11] [27]. Statistical significance testing, such as McNemar's test, validates performance improvements against baseline models and establishes clinical reliability [5].
Cross-validation strategies, particularly k-fold (k=5 or k=10) approaches, mitigate overfitting concerns with limited dataset sizes [16] [5]. The implementation of multiple agreement scenarios (no agreement, partial agreement, total agreement among experts) further strengthens validation frameworks by acknowledging the inherent subjectivity in morphological assessment [2].
Diagram 2: CNN Architecture with Feature Engineering
Clinical Implementation Considerations
The transition from experimental validation to clinical implementation necessitates addressing several practical considerations. Computational efficiency remains paramount, with processing times reduced from 30-45 minutes for manual assessment to under 1 minute per sample with optimized CNN implementations [5]. Real-time classification capabilities (approximately 25 milliseconds per sperm) enable comprehensive morphological analysis of sufficient sperm populations (200+ cells) as recommended by WHO guidelines [26].
Model interpretability, facilitated through Grad-CAM attention visualization, provides clinical transparency by highlighting the specific morphological features influencing classification decisions [5]. This explainability component enhances clinician trust and supports diagnostic verification, accelerating adoption within clinical workflows.
Deep learning approaches, particularly CNNs, are fundamentally transforming reproductive biology by addressing critical limitations in traditional sperm morphology assessment. The implementation of specialized architectures, comprehensive feature engineering pipelines, and rigorous validation frameworks has established new standards for accuracy, efficiency, and reproducibility in male fertility evaluation.
Future research directions include the development of multi-modal models integrating morphological, motile, and clinical parameters; expansion of classification schemes to encompass rare morphological variants; and standardization of validation protocols across institutions. As these technologies continue to mature, their integration into clinical practice promises to enhance diagnostic precision, personalize treatment strategies, and ultimately improve outcomes for couples facing infertility challenges.
Building the Model: CNN Architectures and Practical Implementation
The implementation of Convolutional Neural Networks (CNNs) for human sperm morphology classification research hinges on the quality and integrity of the digital image data fed into these models. The process of transforming a biological sample into a curated, analysis-ready digital dataset is critical, as the performance of any deep learning system is fundamentally bounded by its input data. This document details standardized protocols for acquiring and preparing microscopic images of human sperm, providing a foundational framework for building robust and reliable CNN-based classification systems within reproductive research and diagnostics.
Microscopy Acquisition Modalities
The choice of microscopy technique directly influences the type and quality of morphological information that can be extracted. The following modalities are particularly relevant for sperm analysis.
Conventional Optical Microscopy
Principle: This is the traditional method for semen analysis, often involving stained sperm smears examined under brightfield illumination. Staining (e.g., with hematoxylin and eosin) enhances the contrast of sperm structures, facilitating visual distinction between the head, midpiece, and tail [28].
Protocol for Stained Smear Preparation:
- Fixation: Place a drop of liquefied semen on a clean glass slide and allow it to air dry. Fix the smear by immersing it in 95% ethanol for 10 minutes. Air dry once more.
- Staining: Follow a standardized staining protocol, such as the Papanicolaou method or Quick-Diff stain, as recommended by the WHO laboratory manual.
- Mounting: Apply a coverslip using an appropriate mounting medium.
- Image Acquisition: Use a high-magnification oil immersion objective (e.g., 100x) on a brightfield microscope. Capture images from multiple, randomly selected fields to ensure a representative sample of the sperm population. Consistent Köhler illumination is crucial for uniform image quality.
Digital Holographic Microscopy (DHM)
Principle: DHM is a label-free, interferometric technique that quantifies the phase shift of light passing through a specimen. This allows for the reconstruction of three-dimensional topographic profiles of live, unstained spermatozoa [29].
Protocol for Live Sperm DHM Imaging:
- Sample Preparation: Use intact, live spermatozoa directly after semen liquefaction. Place a small droplet (~5-10 µL) of the sample on a microscope slide. For wet mounting, a coverslip can be applied.
- Acquisition: The DHM system records holograms via a CCD camera by interfering the laser beam that has passed through the sample with a reference beam.
- Reconstruction: Numerically back-propagate the recorded hologram to reconstruct the optical wavefront of the sperm cells. This process generates quantitative phase images.
- 3D Parameter Extraction: Extract novel 3D morphological parameters, such as head height (hh), acrosome/nucleus height (anh), and head/midpiece height (hmh), which have been shown to be less variable in sperm from fertile men [29].
Image-Based Flow Cytometry (IBFC)
Principle: IBFC combines the high-throughput capabilities of traditional flow cytometry with high-speed, single-cell imaging. It allows for the rapid collection of thousands of individual sperm images, which is ideal for building large-scale datasets for deep learning [28].
Protocol for Sperm Imaging via IBFC:
- Sample Fixation: Fix sperm samples in 2% formaldehyde for 40 minutes at room temperature. Wash and resuspend in phosphate-buffered saline (PBS) for analysis [28].
- Instrument Configuration: Use an instrument such as the ImageStreamX Mark II, which can be fitted with 20x, 40x, and 60x objective lenses.
- Acquisition: Hydrodynamically focus the sperm suspension to pass cells single-file past the objective. Trigger the camera to capture a brightfield image of each individual spermatozoon. Magnifications of 60x are recommended for high-fidelity morphological classification [28].
Table 1: Comparison of Microscopy Modalities for Sperm Image Acquisition
| Modality | Sample State | Key Advantages | Key Limitations | Suitability for CNN |
|---|---|---|---|---|
| Conventional (Stained) | Fixed & Stained | High contrast, standardized protocols, familiar to clinics | Staining artefacts, 2D only, destructive process | High, but may not reflect live-state morphology |
| Digital Holographic (DHM) | Live & Unstained | Label-free, provides 3D parameters, non-invasive | Specialized equipment, complex data reconstruction | High for novel 3D feature extraction |
| Image-Based Flow Cytometry | Fixed or Live | Very high throughput, single-cell images, scalable | Lower resolution per image compared to microscopy, cost | Excellent for building large training datasets |
From Raw Images to Analysis-Ready Data
Once acquired, raw images must undergo a series of preprocessing and annotation steps to be usable for supervised CNN training.
Image Preprocessing
The goal of preprocessing is to standardize images and enhance relevant features.
- Contrast Enhancement: Apply techniques like histogram equalization to improve the distinction between sperm structures and the background, which is especially critical for unstained images with low signal-to-noise ratios [30].
- Denoising: Use algorithms or self-supervised deep learning models (e.g., Noise2Void) to reduce noise without altering the underlying morphological structures [31].
- Normalization: Scale pixel intensity values to a standard range (e.g., 0 to 1) to ensure consistent input for the CNN and stabilize the training process.
Annotation and Ground Truth Labeling
For supervised learning, every image in the training set requires accurate annotation. This is a critical and time-consuming step.
- Multi-Part Segmentation: The most detailed annotation involves pixel-level segmentation of each sperm component. As per recent research, advanced models like Mask R-CNN, YOLOv8, and U-Net are used to segment the head, acrosome, nucleus, neck, and tail [30]. Annotators use software to manually outline these regions, creating "ground truth" masks.
- Whole-Cell Classification: For broader categorization, entire sperm cells are labeled according to WHO morphology classes (e.g., "normal", "head defect", "coiled tail") or specific research criteria [32] [33].
- Annotation Tools: Utilize specialized software that supports instance segmentation, such as those integrated in ZeroCostDL4Mic, or other bioimage analysis platforms [31].
Data Augmentation
Data augmentation artificially expands the size and diversity of the training dataset by applying random, realistic transformations to the original images. This technique is vital for improving model robustness and reducing overfitting.
- Common Techniques: Include random rotations, flipping (horizontal/vertical), zooming, shearing, and adjusting brightness/contrast.
- Implementation: This can be performed on-the-fly during CNN training using built-in functions in deep learning frameworks (e.g., Keras
ImageDataGenerator) or using Python packages like Augmentor or imgaug, which are integrated into platforms like ZeroCostDL4Mic [31].
Experimental Workflow for Data Acquisition and Model Training
The following diagram summarizes the integrated workflow from sample preparation to CNN model evaluation.
Table 2: Key Research Reagent Solutions and Computational Tools
| Category | Item / Tool | Function / Application |
|---|---|---|
| Wet Lab Reagents | Formaldehyde (2%) | Fixation of sperm for IBFC or stained smears to preserve morphology [28]. |
| Papanicolaou Stain | Standardized staining solution for enhancing contrast of sperm structures in brightfield microscopy. | |
| Phosphate-Buffered Saline (PBS) | Washing and suspension medium for sperm samples. | |
| Percoll Gradient | Density gradient medium for selecting morphologically normal spermatozoa for specific studies [29]. | |
| Computational Tools | ZeroCostDL4Mic | A cloud-based platform (Google Colab) providing free-access Jupyter Notebooks for training DL models (U-Net, YOLO, CARE) without coding expertise [31]. |
| Mask R-CNN / YOLOv8 / U-Net | Deep learning models for instance segmentation (Mask R-CNN, YOLO) and semantic segmentation (U-Net) of sperm components [30]. | |
| ResNet-50 | A deep CNN architecture used for classification tasks, such as assigning sperm motility categories from video data [16]. | |
| Augmentor / imgaug | Python packages for implementing data augmentation to increase the effective size and diversity of training datasets [31]. |
Quantitative Performance Metrics for Data Quality and Model Evaluation
Establishing quantitative metrics is essential for evaluating both the quality of the annotations and the performance of the trained CNN model.
Table 3: Key Quantitative Metrics for Segmentation and Classification
| Metric | Definition | Interpretation and Target Value | ||||||
|---|---|---|---|---|---|---|---|---|
| Intersection over Union (IoU) | Area of Overlap / Area of Union between predicted and ground truth mask. | Measures segmentation accuracy. A score of 0.8 or higher is generally considered reliable [34]. For sperm nuclei, advanced models can achieve ~0.97 [30]. | ||||||
| Dice Coefficient (F1 Score) | 2 × ( | Prediction ∩ Truth | ) / ( | Prediction | + | Truth | ) | Similar to IoU, it quantifies the overlap between segmentation masks. Values closer to 1.0 indicate better performance. |
| Precision | True Positives / (True Positives + False Positives) | Measures the reliability of positive predictions. High precision means fewer false alarms. | ||||||
| Recall | True Positives / (True Positives + False Negatives) | Measures the ability to find all relevant positive cases. High recall means fewer missed detections. | ||||||
| Mean Absolute Error (MAE) | Average absolute difference between predicted and actual values. | Used in regression or motility classification. A lower MAE is better. For 3-category motility classification, MAE can be as low as 0.05 [16]. |
Pre-processing and Augmentation Techniques to Enhance Model Robustness
The manual assessment of sperm morphology is a cornerstone of male fertility diagnosis but remains highly subjective, prone to significant inter-observer variability, and challenging to standardize across laboratories [2] [9]. Convolutional Neural Networks (CNNs) offer a promising path toward the automation, standardization, and acceleration of this analysis [2] [24]. However, the performance and robustness of these deep learning models are critically dependent on the quality and quantity of the training data. This document provides detailed Application Notes and Protocols for the essential pre-processing and data augmentation techniques required to build reliable CNN models for human sperm morphology classification, directly supporting the broader objectives of thesis research in this field.
Core Techniques and Their Impact
Effective data preparation involves a pipeline of techniques designed to clean the data, expand its diversity, and ultimately teach the model to focus on biologically relevant features while ignoring irrelevant noise. The following table summarizes the quantitative impact of these techniques as reported in recent literature.
Table 1: Impact of Pre-processing and Augmentation Techniques on Model Performance
| Technique Category | Specific Method | Reported Outcome/Performance | Source/Context |
|---|---|---|---|
| Dataset Augmentation | Multiple techniques (e.g., geometric, noise) | Expanded dataset from 1,000 to 6,035 images; Model accuracy 55-92% | SMD/MSS Dataset Development [2] |
| Deep Feature Engineering | CBAM + ResNet50 + PCA + SVM RBF | Accuracy of 96.08% on SMIDS dataset; ~8% improvement over baseline CNN | Sperm Classification with Feature Engineering [5] |
| Deep Feature Engineering | CBAM + ResNet50 + PCA + SVM RBF | Accuracy of 96.77% on HuSHeM dataset; ~10.4% improvement over baseline CNN | Sperm Classification with Feature Engineering [5] |
| Object Detection | YOLOv7 on bovine sperm | Global mAP@50: 0.73; Precision: 0.75; Recall: 0.71 | Veterinary Reproduction Study [35] [10] |
Detailed Experimental Protocols
Protocol 1: Image Pre-processing for Sperm Morphology Analysis
This protocol outlines the essential steps for preparing raw sperm images for CNN model training, aiming to reduce noise and standardize input data [2].
3.1.1 Materials and Equipment
- Source Images: Raw sperm images acquired via CASA system or bright-field microscope [2].
- Analysis Software: Python 3.8 with libraries (e.g., OpenCV, SciKit-Image, TensorFlow/PyTorch) [2] [5].
3.1.2 Step-by-Step Procedure
- Data Cleaning: Identify and handle missing values, outliers, or any inconsistencies in the dataset. Cleaning the data ensures the model is not influenced by noise or inaccuracies that hinder performance [2].
- Grayscale Conversion: Convert all input images to a single-channel (grayscale) format to reduce computational complexity and focus on morphological structures rather than potential staining color variations [2].
- Resizing: Resize all images to a consistent dimensions using a linear interpolation strategy. A common size used in research is 80x80 pixels [2]. This standardization is required for batch processing in CNN.
- Normalization: Normalize pixel intensity values to a common scale, typically [0, 1], by dividing all values by the maximum possible value (e.g., 255). This ensures no particular feature dominates the learning process due to differences in magnitude and improves numerical stability during training [2] [36].
Protocol 2: Data Augmentation for Enhanced Generalization
This protocol describes methods to artificially expand the training dataset, which is crucial for improving model robustness and mitigating overfitting, especially given the common challenge of limited and class-imbalanced medical datasets [2] [9].
3.2.1 Materials and Equipment
- Pre-processed Images: The standardized images output from Protocol 1.
- Software: Python with deep learning frameworks (e.g., TensorFlow/Keras, PyTorch) that include built-in image transformation functions.
3.2.2 Step-by-Step Procedure Apply a series of geometric and pixel-wise transformations to generate new training samples from the existing dataset. The following transformations are recommended:
- Geometric Transformations:
- Rotation: Randomly rotate images by a defined range of angles (e.g., ±15°).
- Flipping: Randomly flip images horizontally and/or vertically.
- Shearing and Zooming: Apply slight shearing and zoom transformations to simulate different perspectives.
- Pixel-wise Transformations:
3.2.3 Implementation Note These transformations can be applied in real-time during training (on-the-fly augmentation) or as a pre-processing step to create a larger, static dataset. The parameters for each transformation should be chosen to create plausible sperm images without distorting critical morphological features.
The workflow below illustrates the sequential steps of a robust data preparation pipeline for sperm morphology classification.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials and Reagents for Sperm Morphology Analysis Experiments
| Item Name | Function/Application | Example/Specification |
|---|---|---|
| CASA System | Automated image acquisition and initial morphometric analysis of sperm cells. | MMC CASA system [2] |
| Optical Microscope | High-resolution imaging of sperm smears. | Microscope with oil immersion 100x objective in bright-field mode [2] |
| Staining Kit | Enhances contrast for visual and computational analysis of sperm structures. | RAL Diagnostics staining kit [2] |
| Annotation Software | For labeling images and creating ground truth data for model training. | Software like Roboflow [35] |
| Deep Learning Framework | Provides the programming environment to build, train, and test CNN models. | Python 3.8 with TensorFlow/PyTorch [2] [5] |
Advanced Integrated Framework
For researchers aiming to achieve state-of-the-art performance, combining advanced architectural components with pre-processing and augmentation yields significant benefits. The following workflow integrates an attention mechanism and feature engineering into a high-accuracy classification system.
Procedure for Advanced Framework Implementation:
- Model Architecture Setup: Employ a pre-trained CNN (e.g., ResNet50) as a feature extraction backbone. Integrate a Convolutional Block Attention Module (CBAM) that sequentially applies channel and spatial attention to intermediate feature maps. This mechanism enables the network to focus on the most relevant sperm features (e.g., head shape, acrosome, tail) while suppressing background or noise [5].
- Deep Feature Extraction & Selection: Instead of using the CNN for direct end-to-end classification, extract high-dimensional feature representations from the network's intermediate layers. Subsequently, apply classical feature selection and dimensionality reduction techniques, such as Principal Component Analysis (PCA), to this feature set. This process reduces noise and creates a more compact and discriminative feature vector for classification [5].
- Hybrid Classification: Feed the optimized feature vector into a shallow classifier, such as a Support Vector Machine (SVM) with an RBF kernel. This hybrid approach (CNN + feature engineering + SVM) has been demonstrated to yield superior performance compared to standalone CNNs, achieving accuracy levels above 96% on benchmark datasets [5].
The implementation of Convolutional Neural Networks (CNNs) for human sperm morphology classification represents a paradigm shift in male fertility assessment. Traditional manual analysis is highly subjective, time-intensive (taking 30–45 minutes per sample), and suffers from significant inter-observer variability, with studies reporting up to 40% disagreement between expert evaluators [5] [37]. Automated CNN-based systems address these limitations by providing objective, standardized assessments that can reduce analysis time to under one minute per sample while improving diagnostic consistency across laboratories [5] [37]. These systems are particularly valuable in clinical settings where subtle morphological differences in sperm head shape, acrosome integrity, and tail structure must be consistently identified according to World Health Organization (WHO) criteria [20] [11].
The evolution of CNN architectures for this specialized domain has progressed from using pre-trained networks as feature extractors to developing sophisticated custom hybrids that integrate attention mechanisms and ensemble strategies. ResNet50 has emerged as a particularly effective backbone architecture due to its residual learning framework, which mitigates vanishing gradient problems in deep networks and enables effective training even with limited medical imaging data [5]. More recently, researchers have enhanced ResNet50 with Convolutional Block Attention Modules (CBAM) to help networks focus on morphologically discriminative sperm regions while suppressing irrelevant background information [5] [37]. Simultaneously, ensemble approaches combining multiple EfficientNetV2 variants have demonstrated robust performance across diverse abnormality classes by leveraging complementary feature representations [32].
Landscape of CNN Architectures: Performance Comparison
Table 1: Quantitative Performance Comparison of CNN Architectures for Sperm Morphology Classification
| Architecture | Key Features | Dataset | Classes | Performance |
|---|---|---|---|---|
| CBAM-Enhanced ResNet50 + Deep Feature Engineering | Attention mechanism + PCA + SVM classifier | SMIDS | 3 | 96.08% ± 1.2% accuracy [5] |
| HuSHeM | 4 | 96.77% ± 0.8% accuracy [5] [37] | ||
| Multi-Level Ensemble (EfficientNetV2) | Feature-level & decision-level fusion | Hi-LabSpermMorpho | 18 | 67.70% accuracy [32] |
| VGG16 with Transfer Learning | Fine-tuning pre-trained weights | HuSHeM | 5 | 94.1% true positive rate [11] |
| SCIAN | 5 | 62% true positive rate [11] | ||
| Custom CNN | Five convolutional layers | SMD/MSS | 12 | 55-92% accuracy range [2] |
Table 2: Technical Specifications of Featured CNN Architectures
| Architecture | Feature Extraction Method | Classifier | Attention Mechanism | Data Augmentation |
|---|---|---|---|---|
| CBAM-Enhanced ResNet50 | Multiple layers (CBAM, GAP, GMP, pre-final) | SVM with RBF/Linear kernels | CBAM (Channel & Spatial) | Not specified [5] |
| Multi-Level Ensemble | Multiple EfficientNetV2 variants | SVM, Random Forest, MLP-Attention | MLP-Attention | Yes (dataset specific) [32] |
| VGG16 Transfer Learning | Pre-trained on ImageNet | Fine-tuned fully connected layers | None | Not specified [11] |
| Custom CNN | Five convolutional layers | Fully connected layers | None | Yes (6035 images from 1000 originals) [2] |
Experimental Protocols for CNN Implementation
Protocol 1: CBAM-Enhanced ResNet50 with Deep Feature Engineering
Purpose: To implement an attention-based deep learning framework combining ResNet50 with comprehensive feature engineering for superior sperm morphology classification [5] [37].
Materials and Reagents:
- SMIDS dataset (3000 images, 3-class) or HuSHeM dataset (216 images, 4-class)
- Python 3.x with TensorFlow/PyTorch frameworks
- Hardware: GPU-enabled computing environment
- Stained semen smears following WHO guidelines [20]
Procedure:
- Backbone Architecture Preparation:
- Implement ResNet50 architecture as feature extraction backbone
- Integrate Convolutional Block Attention Module (CBAM) sequentially after residual blocks
- Configure CBAM to perform channel attention first, followed by spatial attention
Multi-Layer Feature Extraction:
- Extract deep features from four strategic layers: CBAM attention layers, Global Average Pooling (GAP), Global Max Pooling (GMP), and pre-final layer
- Concatenate features from multiple layers to capture both spatial and semantic information
Feature Selection and Dimensionality Reduction:
- Apply Principal Component Analysis (PCA) to reduce feature dimensionality while preserving discriminative information
- Evaluate alternative feature selection methods including Chi-square test, Random Forest importance, and variance thresholding
- Select optimal feature subset based on classification performance
Classifier Training and Evaluation:
- Implement Support Vector Machine (SVM) classifier with RBF and linear kernels
- Train classifier on reduced feature set using 5-fold cross-validation
- Evaluate performance using accuracy, precision, recall, and F1-score metrics
- Generate Grad-CAM visualizations to interpret model focus areas
Validation: Perform statistical significance testing using McNemar's test (p < 0.05) to compare against baseline CNN performance [5] [37].
Protocol 2: Multi-Level Ensemble Learning with EfficientNetV2
Purpose: To develop an ensemble framework combining multiple CNN architectures through feature-level and decision-level fusion for comprehensive sperm morphology classification across 18 distinct morphological classes [32].
Materials and Reagents:
- Hi-LabSpermMorpho dataset (18,456 images across 18 classes)
- Multiple EfficientNetV2 variants (S, M, L)
- Support Vector Machines (SVM), Random Forest, and MLP-Attention classifiers
- Data augmentation pipeline for class imbalance mitigation
Procedure:
- Multi-Model Feature Extraction:
- Implement multiple EfficientNetV2 variants (S, M, L) as parallel feature extractors
- Extract deep features from the penultimate layer of each network
- Apply dimensionality reduction using dense-layer transformations
Feature-Level Fusion:
- Concatenate features from all EfficientNetV2 variants into a unified feature representation
- Normalize fused features to ensure balanced contribution from each network
- Apply feature selection techniques to reduce dimensionality while preserving discriminative power
Classifier Implementation and Decision-Level Fusion:
- Train multiple classifier types (SVM, Random Forest, MLP-Attention) on fused features
- Implement soft voting mechanism for decision-level fusion
- Optimize fusion weights based on individual classifier performance
- Apply weighted averaging of class probabilities for final prediction
Class Imbalance Mitigation:
- Analyze performance across minority and majority classes
- Implement data augmentation strategies specific to underrepresented classes
- Adjust classification thresholds or loss functions to address imbalance
Validation: Evaluate framework using stratified k-fold cross-validation, with particular attention to performance consistency across all 18 morphological classes [32].
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Essential Research Reagents and Computational Tools for CNN-Based Sperm Morphology Analysis
| Item | Specification | Function/Application |
|---|---|---|
| Benchmark Datasets | HuSHeM (216 images), SCIAN, SMIDS (3000 images), Hi-LabSpermMorpho (18,456 images) | Model training, validation, and benchmarking [32] [5] [11] |
| Data Augmentation Tools | Python libraries (TensorFlow, PyTorch, OpenCV) | Address class imbalance, expand training data, improve generalization [2] |
| Attention Mechanisms | Convolutional Block Attention Module (CBAM) | Enhance focus on discriminative morphological features [5] [37] |
| Feature Selection Methods | PCA, Chi-square test, Random Forest importance, variance thresholding | Dimensionality reduction, noise suppression, performance optimization [5] |
| Classification Algorithms | SVM (RBF/Linear), Random Forest, MLP-Attention | Final morphology classification using deep features [32] [5] |
| Evaluation Metrics | Accuracy, Precision, Recall, F1-Score, McNemar's test | Performance assessment and statistical validation [32] [5] |
Architectural Decision Framework and Clinical Implementation
The selection of appropriate CNN architecture for sperm morphology classification depends on multiple factors including dataset characteristics, computational resources, and clinical requirements. For laboratories with limited data (200-1000 images), transfer learning with VGG16 or ResNet50 provides robust performance without extensive training data requirements [11]. When classifying a broad spectrum of morphological abnormalities (10+ classes), ensemble approaches with EfficientNetV2 variants offer superior performance through complementary feature representation, though at increased computational cost [32]. For maximum classification accuracy on well-defined morphological categories, CBAM-enhanced ResNet50 with deep feature engineering currently represents the state-of-the-art, achieving up to 96.77% accuracy on benchmark datasets [5] [37].
Clinical implementation requires careful consideration of interpretability needs alongside raw performance. Attention mechanisms like CBAM not only improve accuracy but generate Grad-CAM visualizations that help clinicians understand model decisions and build trust in automated systems [5]. Furthermore, the significant time reduction from 30-45 minutes to under 1 minute per sample represents a substantial efficiency gain for clinical workflows, potentially increasing laboratory throughput and standardizing diagnostic criteria across institutions [5] [37]. As these technologies mature, integration with existing laboratory information systems and validation against clinical outcomes (pregnancy success rates) will be essential for widespread adoption in reproductive medicine.
The implementation of Convolutional Neural Networks (CNNs) for human sperm morphology classification presents a significant paradigm shift in male fertility diagnostics. This critical analysis within the broader thesis research context compares two fundamental training approaches: transfer learning, which leverages pre-existing knowledge from pre-trained models, and training from scratch, which builds models exclusively on target domain data. The morphological classification of human sperm is a well-established indicator of biological function and male fertility, yet manual assessment remains laborious, time-consuming, and subject to inter-observer variability [38] [25]. Deep learning approaches offer the potential to automate, standardize, and accelerate this analysis, with the choice of training strategy profoundly impacting model performance, computational efficiency, and practical applicability in clinical and research settings [2].
Comparative Analysis of Training Strategies
Theoretical Foundations and Performance Characteristics
Transfer Learning utilizes knowledge gained from solving a source problem (S) to improve learning efficiency and effectiveness on a target problem (T), where the domains or tasks may differ [39]. In practical implementation, this typically involves pre-training a model on a large, general dataset (e.g., ImageNet) followed by fine-tuning on the specific target task with a smaller dataset [38] [40]. This approach is particularly valuable in medical imaging domains where annotated data is scarce and expert labeling is costly.
Training from Scratch involves initializing model parameters randomly and training exclusively on the target dataset. This approach requires no pre-trained models but typically demands larger quantities of labeled target data to achieve competitive performance [25]. While this method avoids potential domain mismatch between source and target tasks, it faces significant challenges in medical imaging applications where data limitations are common.
Table 1: Quantitative Performance Comparison of Training Strategies in Medical Imaging
| Application Domain | Model Architecture | Training Strategy | Performance Metrics | Reference |
|---|---|---|---|---|
| Sperm Head Classification | Modified AlexNet | Transfer Learning | 96.0% Accuracy, 96.4% Precision | [38] |
| Sperm Head Classification | Custom CNN | Training from Scratch | 88.0% Recall (SCIAN) | [25] |
| Sperm Morphology Assessment | CNN with Augmentation | Training from Scratch | 55-92% Accuracy Range | [2] |
| Cross-Modality Medical Imaging | MobileNetV3 | Cross-Modality Transfer Learning | 0.99 Accuracy | [40] |
| Brain Tumor Segmentation | DeepLabv3+ with EfficientNet | Transfer Learning | 99.53% Accuracy | [41] |
Strategic Advantages and Limitations
Transfer Learning demonstrates remarkable effectiveness in scenarios with limited training data. The modified AlexNet approach for sperm head classification achieved 96.0% accuracy despite the small HuSHeM dataset containing only 216 images [38]. This strategy significantly reduces training time and computational resources compared to training from scratch [39]. Cross-modality and cross-organ transfer learning further expand its applicability, as demonstrated by MobileNetV3 pre-trained on mammograms achieving 0.99 accuracy on prostate data [40]. However, potential limitations include overfitting during fine-tuning and domain mismatch when source and target distributions differ substantially [42].
Training from Scratch offers the advantage of complete specialization to the target domain without potential bias from pre-training on dissimilar datasets. Custom architectures can be meticulously designed to address domain-specific challenges, such as the specialized CNN for sperm head classification that achieved 88% recall on the SCIAN dataset [25]. The primary limitation remains the substantial data requirement, with performance highly dependent on dataset size and quality. Training from scratch typically demands more extensive data augmentation and longer training times to achieve convergence [2].
Experimental Protocols for Sperm Morphology Classification
Protocol 1: Transfer Learning Implementation
A. Dataset Preparation and Preprocessing
- Acquire human sperm images using standardized microscopy protocols (e.g., bright field mode with oil immersion 100x objective) [38] [2]
- Apply automated preprocessing using OpenCV-based pipelines: denoise images, convert to grayscale, apply Sobel operator for gradient detection, use adaptive thresholding for binarization, perform morphological operations (erosion/dilation) to eliminate interference [38]
- Extract sperm heads through elliptical fitting of contours and crop to standardized dimensions (e.g., 64×64 pixels) [38]
- Align sperm head orientation to uniform direction to reduce rotational variance [38]
- Implement dataset partitioning (80% training, 20% testing) with random shuffling to minimize bias [2]
B. Model Selection and Adaptation
- Select appropriate pre-trained architecture (AlexNet, VGG16, or MobileNetV3) based on computational constraints and task complexity [38] [40]
- Replace final classification layer with domain-specific head matching sperm morphology categories (normal, tapered, pyriform, amorphous) [38]
- Integrate Batch Normalization layers to improve training stability and convergence [38]
- Optionally freeze early convolutional layers to preserve general feature detectors while allowing deeper layers to adapt to domain-specific features [39]
C. Fine-Tuning and Optimization
- Initialize with pre-trained weights from large-scale datasets (e.g., ImageNet)
- Employ moderate learning rates (0.01-0.001) with SGD or Adam optimizers to balance adaptation and preservation of transferred knowledge [38]
- Implement early stopping based on validation accuracy to prevent overfitting
- Apply gentle data augmentation (horizontal flipping, minor rotations) to increase effective dataset size while preserving biological validity [2]
Protocol 2: Training from Scratch Implementation
A. Comprehensive Data Preparation
- Curate extensive sperm image datasets with expert annotations following WHO standards or modified David classification [2] [25]
- Implement aggressive data augmentation to address limited dataset size: rotational transformations, scaling variations, intensity adjustments, and elastic deformations [2]
- Balance class distributions through oversampling of rare morphological classes or synthetic data generation
- Employ multiple expert annotations with consensus mechanisms to establish reliable ground truth and address inter-expert variability [2]
B. Custom Architecture Design
- Develop domain-optimized architectures with careful consideration of sperm image characteristics (small size, limited structural complexity)
- Design efficient networks with multiple filter sizes but fewer parameters to prevent overfitting on limited data [25]
- Incorporate regularization techniques: Dropout layers (0.1-0.5 rate), Batch Normalization, and L2 weight regularization [42]
- Implement progressive training strategies, starting with simpler architectures and gradually increasing complexity
C. Specialized Training Methodology
- Employ sophisticated initialization strategies (He/Xavier) to promote stable gradient flow
- Utilize dynamic learning rate schedules with reduction on plateau to refine convergence
- Implement strong data augmentation throughout training to effectively increase dataset diversity
- Apply model checkpointing and ensemble methods to enhance final performance and robustness
Workflow Visualization
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Research Materials and Computational Tools for Sperm Morphology CNN Research
| Research Component | Specification/Example | Function/Purpose | Implementation Notes |
|---|---|---|---|
| Imaging System | MMC CASA System with 100x oil immersion objective | High-quality sperm image acquisition | Standardize acquisition parameters across samples [2] |
| Staining Protocol | RAL Diagnostics staining kit | Enhance contrast for morphological features | Follow WHO standardized protocols [2] |
| Annotation Framework | Multi-expert consensus (3 specialists) | Establish reliable ground truth | Resolve discrepancies through majority voting [2] |
| Public Datasets | HuSHeM (216 images), SCIAN (1854 images), SMD/MSS (1000 images) | Benchmarking and comparative analysis | HuSHeM covers 4 classes; SCIAN includes 5 categories [38] [25] |
| Data Augmentation | Rotation, flipping, intensity adjustment | Address limited dataset size | Apply biologically plausible transformations only [2] |
| Pre-trained Models | AlexNet, VGG16, MobileNetV3 | Transfer learning foundation | AlexNet offers efficiency; VGG16 provides depth [38] [40] |
| Optimization Algorithms | SGD with momentum, Adam | Model parameter optimization | SGD with learning rate 0.01 effective for fine-tuning [38] |
| Regularization Techniques | Dropout (0.1-0.5), Batch Normalization | Prevent overfitting | Batch Normalization improves training stability [38] |
The selection between transfer learning and training from scratch for CNN implementation in human sperm morphology classification depends primarily on dataset characteristics, computational resources, and performance requirements. Transfer learning demonstrates superior performance in data-scarce environments, achieving up to 96% accuracy with limited samples [38], while training from scratch offers competitive results (88-95% recall) with sufficient data and appropriate architectural design [25].
For most research and clinical applications in sperm morphology classification, transfer learning provides the most practical and effective approach, particularly given the typical challenges of limited annotated data and computational constraints. The integration of cross-modality pre-training [40] and advanced fine-tuning strategies can further enhance performance. Training from scratch remains valuable for specialized applications requiring complete domain specificity or when substantial datasets with comprehensive expert annotations are available. Future research directions should explore hybrid approaches, domain adaptation techniques, and automated architecture search to optimize model performance while maintaining computational efficiency for clinical deployment.
Male infertility is a significant global health concern, contributing to approximately 50% of infertility cases among couples [20]. Semen analysis serves as a cornerstone in male fertility assessment, with sperm morphology representing one of the most clinically relevant parameters for predicting fertility potential [2]. Traditional manual morphology assessment suffers from substantial subjectivity, inter-laboratory variability, and reliance on expert technicians, making standardization challenging [2] [20]. While computer-assisted semen analysis (CASA) systems were developed to address these limitations, they often struggle with accurately distinguishing spermatozoa from cellular debris and classifying specific midpiece and tail abnormalities [2].
The emergence of deep learning, particularly convolutional neural networks (CNNs), offers promising solutions for automating sperm analysis while improving accuracy and standardization. This case study explores the implementation of CNNs for analyzing unstained live sperm, focusing on morphological classification within a research framework aimed at enhancing diagnostic precision in male infertility evaluation.
Current Landscape of Sperm Morphology Datasets
The development of robust CNN models depends critically on the availability of high-quality, well-annotated datasets. Significant variability exists among publicly available sperm image datasets in terms of staining methods, resolution, annotation quality, and morphological classifications.
Table 1: Overview of Human Sperm Morphology Datasets
| Dataset Name | Year | Characteristics | Annotation Type | Image Count | Key Features |
|---|---|---|---|---|---|
| SMD/MSS [2] | 2025 | Unstained | Classification (Modified David) | 1,000 → 6,035 (after augmentation) | 12 morphological defect classes; single sperm images |
| VISEM [9] | 2019 | Unstained, grayscale | Regression | Multi-modal with videos | From 85 participants; includes biological data |
| VISEM-Tracking [9] | 2023 | Unstained, grayscale | Detection, tracking, regression | 656,334 annotated objects | Extensive tracking details |
| SVIA [20] [9] | 2022 | Unstained, grayscale | Detection, segmentation, classification | 125,880 cropped objects | Multiple annotation types; 26,000 segmentation masks |
| MHSMA [9] | 2019 | Unstained, noisy, low resolution | Classification | 1,540 grayscale images | Focus on sperm head features |
| HSMA-DS [9] | 2015 | Unstained, noisy, low resolution | Classification | 1,457 images from 235 patients | Early benchmark dataset |
The SMD/MSS (Sperm Morphology Dataset/Medical School of Sfax) dataset represents a recent contribution specifically designed for deep learning applications, utilizing the modified David classification system that includes 12 classes of morphological defects across head, midpiece, and tail compartments [2]. This classification system encompasses seven head defects (tapered, thin, microcephalous, macrocephalous, multiple, abnormal post-acrosomal region, abnormal acrosome), two midpiece defects (cytoplasmic droplet, bent), and three tail defects (coiled, short, multiple) [2].
A fundamental challenge in dataset creation is the substantial inter-expert variability in morphological assessment. In the SMD/MSS development, three experts performed independent classifications, with statistical analysis revealing varying agreement levels: no agreement (NA), partial agreement (PA: 2/3 experts), and total agreement (TA: 3/3 experts) [2]. This heterogeneity in ground truth annotation directly impacts model training and performance evaluation.
CNN Architecture Design and Implementation
Preprocessing Pipeline for Unstained Sperm Images
Unstained sperm images present unique challenges including low contrast, noise from optical microscopy, and interfering cellular debris [2]. An effective preprocessing pipeline is essential for optimal CNN performance:
- Data Cleaning: Identification and handling of missing values, outliers, or inconsistencies [2]
- Normalization/Standardization: Rescaling pixel values to a common range to standardize input distributions
- Image Resizing: Uniform resizing to dimensions compatible with CNN architecture (e.g., 80×80×1 for grayscale) using linear interpolation [2]
- Grayscale Conversion: Transformation of color images to single-channel grayscale to reduce computational complexity [2]
Data Augmentation Strategies
To address limited dataset size and class imbalance, strategic data augmentation techniques are employed:
- Geometric transformations (rotation, scaling, translation)
- Elastic deformations
- Intensity variations
- Noise injection
In the SMD/MSS dataset, augmentation expanded the original 1,000 images to 6,035 images, significantly enhancing model generalization [2].
CNN Architecture Selection
Multiple architectural approaches have demonstrated efficacy in sperm analysis:
- Custom CNN Architectures: Tailored networks with convolutional, pooling, and fully connected layers [2]
- ResNet-50: Proven effective for sperm motility classification using optical flow representations [16]
- Advanced Meta-Learning: HSHM-CMA algorithm utilizing contrastive meta-learning with auxiliary tasks for improved cross-domain generalization [43]
Implementation Framework
- Programming Language: Python 3.8 [2]
- Deep Learning Framework: TensorFlow/Keras [16]
- Optimization Algorithm: Adam optimizer with learning rate of 0.0004 [16]
- Loss Function: Mean Absolute Error (MAE) for regression tasks, categorical cross-entropy for classification [16]
Experimental Protocols
Sample Preparation and Image Acquisition
Protocol: Sperm Sample Preparation for Unstained Analysis
- Sample Collection: Obtain semen samples after informed consent from patients undergoing fertility evaluation [2]
- Inclusion Criteria: Select samples with sperm concentration ≥5 million/mL and varying morphological profiles [2]
- Exclusion Criteria: Exclude samples with concentration >200 million/mL to prevent image overlap [2]
- Smear Preparation: Prepare smears according to WHO manual guidelines without staining [2]
- Image Acquisition: Use MMC CASA system with bright field mode, oil immersion 100× objective [2]
- Capture Parameters: Acquire 37±5 images per sample depending on density and distribution [2]
- Single-Sperm Isolation: Ensure each image contains a single spermatozoon with head, midpiece, and tail [2]
Expert Annotation and Ground Truth Establishment
Protocol: Multi-Expert Morphological Classification
- Expert Selection: Engage three experienced andrology technicians [2]
- Classification System: Utilize modified David classification with 12 defect categories [2]
- Independent Assessment: Each expert classifies images independently without consultation [2]
- Agreement Assessment: Categorize annotations as NA (no agreement), PA (partial agreement: 2/3), or TA (total agreement: 3/3) [2]
- Statistical Analysis: Use Fisher's exact test with significance at p<0.05 to evaluate inter-expert differences [2]
- Ground Truth Compilation: Create comprehensive file with image name, expert classifications, and morphometric data [2]
Model Training and Evaluation
Protocol: CNN Training and Validation
- Data Partitioning: Random split with 80% for training and 20% for testing [2]
- Cross-Validation: Implement k-fold cross-validation (k=10) to ensure robustness [16]
- Early Stopping: Monitor validation loss with patience of 15 epochs to prevent overfitting [16]
- Performance Metrics:
- Comparative Analysis: Benchmark against conventional methods (SVM, decision trees) and inter-expert variability [20]
Performance Analysis and Benchmarking
Quantitative Performance Metrics
Table 2: Performance Comparison of Sperm Analysis Algorithms
| Model/Approach | Task | Performance Metrics | Dataset | Limitations |
|---|---|---|---|---|
| Custom CNN [2] | Morphology Classification | Accuracy: 55-92% | SMD/MSS (6,035 images) | Wide accuracy range depending on morphological class |
| ResNet-50 [16] | Motility Classification | MAE: 0.05 (3-category), 0.07 (4-category) | 65 video recordings | Limited to motility assessment |
| MotionFlow + DNN [27] | Motility & Morphology | MAE: 6.842% (motility), 4.148% (morphology) | VISEM | Novel motion representation required |
| HSHM-CMA [43] | Head Morphology | Accuracy: 65.83%, 81.42%, 60.13% (cross-domain) | Multiple HSHM datasets | Focused exclusively on sperm head |
| SVM Classifier [20] | Head Classification | AUC-ROC: 88.59%, Precision: >90% | 1,400 sperm cells | Limited to head morphology only |
| Bayesian Density [20] | Head Shape Classification | Accuracy: 90% | Four morphological categories | Handcrafted features required |
Comparative Analysis with Conventional Methods
Conventional machine learning approaches for sperm morphology analysis typically rely on handcrafted feature extraction followed by classification algorithms such as Support Vector Machines (SVM), k-means clustering, or decision trees [20]. These methods have demonstrated limitations in generalization across datasets and dependency on manual feature engineering [20]. For instance, while some SVM implementations achieved >90% precision for sperm head classification, they often fail to comprehensively address the full spectrum of morphological abnormalities across head, midpiece, and tail compartments [20].
Deep learning approaches offer the advantage of automated feature learning, potentially capturing subtle morphological patterns missed by manual feature engineering. However, CNNs require substantially larger datasets and computational resources compared to conventional methods [20].
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Essential Research Materials for CNN-based Sperm Analysis
| Item | Specification | Function/Application | Example Use Case |
|---|---|---|---|
| CASA System | MMC CASA with digital camera | Image acquisition from sperm smears | Standardized image capture [2] |
| Optical Microscope | Bright field with 100× oil immersion objective | High-resolution sperm imaging | Visualization of morphological details [2] |
| Temperature Control System | 37°C microscope stage | Maintain physiological temperature | Motility analysis [16] |
| Staining Kit | RAL Diagnostics (for stained samples) | Sperm morphology enhancement | Comparative studies with unstained samples [2] |
| Data Augmentation Tools | Python libraries (TensorFlow, Keras) | Dataset expansion | Addressing class imbalance [2] |
| Video Recording System | 400× magnification, 30 fps | Capture sperm motility | Optical flow analysis [16] |
| Annotation Software | Custom Excel templates or specialized tools | Expert classification documentation | Ground truth establishment [2] |
The implementation of CNNs for unstained live sperm analysis represents a significant advancement in male fertility assessment. The SMD/MSS dataset with modified David classification provides a valuable resource for training models on comprehensive morphological defects [2]. Current research demonstrates promising results, with accuracy ranging from 55% to 92% depending on morphological classes, approaching expert-level performance for specific tasks [2].
Critical challenges remain in dataset standardization, annotation consistency, and model generalization across diverse populations. Future research directions should focus on developing larger, more diverse datasets, integrating multimodal data (morphology, motility, concentration), and advancing explainable AI techniques to build clinician trust in automated classification systems.
The transition from research validation to clinical implementation will require extensive multicenter trials, regulatory approval pathways, and standardization of imaging protocols across laboratories. Nevertheless, deep learning approaches hold tremendous potential for revolutionizing semen analysis by enhancing objectivity, reproducibility, and diagnostic accuracy in male infertility evaluation.
In the field of human sperm morphology classification research, Convolutional Neural Networks (CNNs) have demonstrated significant potential to overcome the limitations of manual analysis, which is subjective and labor-intensive [9] [20]. However, simply using CNNs as black-box classifiers is insufficient for clinical adoption and biological insight. This document outlines application notes and experimental protocols for going beyond classification by integrating two powerful families of techniques: Variational Autoencoders (VAEs) for unsupervised feature learning and data augmentation, and Class Activation Mapping (CAM) methods for model interpretability. These approaches enable researchers to discover latent morphological patterns, generate synthetic data, and visualize the decisive morphological features identified by deep learning models, thereby building trust and deepening understanding of sperm quality biomarkers.
Application Notes
The Role of Feature Extraction and Visualization in Sperm Morphology Analysis
The analysis of sperm morphology presents unique computational challenges. The inherent class imbalance in morphological datasets, where normal sperm are outnumbered by various abnormal types, complicates the training of robust classifiers [2]. Furthermore, the clinical utility of a model is contingent upon its interpretability; a predicting system must be able to justify its decisions to gain the trust of clinicians [44] [45].
VAEs for Latent Feature Discovery and Augmentation: VAEs learn a compressed, probabilistic latent representation of input images. Within the context of sperm morphology, this latent space can be structured using priors like the Gaussian Mixture Model (GMM) or the more flexible Gamma Mixture Model (GamMM) to automatically discover and cluster distinct morphological phenotypes in an unsupervised manner [46] [47]. For instance, a Conditional GMM-VAE (CGMVAE) can model complex, non-linear variations in cognitive profiles, a approach that can be directly translated to modeling the nuanced spectrum of sperm morphological defects [47]. Furthermore, by sampling from the latent space of a specific morphological class, VAEs can generate high-quality synthetic sperm images, effectively balancing datasets and improving model generalization [2] [46].
CAMs for Model Interpretability and Verification: CAM techniques, such as Grad-CAM, address the "black box" problem by producing heatmaps that highlight the image regions most influential to the model's classification decision [44] [45]. In sperm morphology analysis, this allows researchers and clinicians to verify that a model is focusing on biologically relevant structures, such as the acrosome, vacuoles in the head, or the integrity of the midpiece and tail, rather than artifacts or irrelevant noise [9] [45]. This pixel-level explainability is crucial for validating the model's decision-making process against expert knowledge.
The following table summarizes key performance metrics reported in recent studies applying deep learning to medical image analysis, which provide a benchmark for expectations in sperm morphology research.
Table 1: Performance Metrics of Deep Learning Models in Medical Image Analysis
| Model / Technique | Application Area | Key Metric | Reported Performance | Reference |
|---|---|---|---|---|
| Deep CNN (Sperm) | Sperm Morphology Classification | Accuracy | 55% to 92% | [2] |
| CNN-Radiomics Fusion | Periapical Lesion Detection | Accuracy / AUC | 97.16% / 0.9914 | [45] |
| MotionFlow + CNN | Sperm Motility & Morphology | Mean Absolute Error | 6.842% (Motility), 4.148% (Morphology) | [27] |
| CGMVAE | Cognitive Profile Clustering | Cluster Separation | Identified 10 nuanced profiles | [47] |
| Grad-CAM | Organ-at-Risk Segmentation | Qualitative Agreement | High agreement with expert reasoning | [44] |
Experimental Protocols
Protocol 1: Data Preparation and Augmentation with VAE
Objective: To create a balanced and expanded dataset of sperm morphology images for robust model training using a VAE.
Materials:
- Sperm Image Datasets: Publicly available datasets such as SVIA (125,000+ annotations) [9], VISEM-Tracking (656,000+ objects) [9], or SMD/MSS (1,000 images, expandable to 6,035 via augmentation) [2].
- Computing Environment: Python 3.8+, with libraries: PyTorch/TensorFlow, NumPy, OpenCV, scikit-learn.
Methodology:
- Image Pre-processing:
- Cleaning: Handle missing values and outliers. Manually review a subset to exclude images with significant debris or overlapping cells [9] [2].
- Normalization: Resize all images to a consistent dimensions (e.g., 80x80 pixels for single-sperm images [2]). Convert to grayscale and normalize pixel values to a [0, 1] range.
- Denoising: Apply filters (e.g., Gaussian blur) to reduce noise from insufficient staining or microscope lighting [2].
VAE Training for Latent Space Formation:
- Architecture: Implement a VAE with an encoder (convolutional layers) and a decoder (deconvolutional layers). The encoder outputs parameters (μ, σ) for the latent distribution.
- Loss Function: Minimize the combined loss:
Loss = Reconstruction Loss (Binary Cross-Entropy) + β * KL-Divergence Loss, where β controls the weight of the latent space constraint. - Structured Latent Space (GamMM-VAE): For clustering, replace the standard Gaussian prior with a Gamma Mixture Model prior. The training objective incorporates the GamMM log-likelihood into the evidence lower bound (ELBO) [46].
Data Augmentation via Latent Sampling:
- For under-represented morphological classes, sample latent vectors (
z) from the corresponding mixture component of the trained GamMM-VAE. - Decode these sampled vectors to generate novel, high-quality synthetic sperm images that reflect the variations within the target class [46].
- For under-represented morphological classes, sample latent vectors (
Figure 1: Workflow for data augmentation using a VAE with a mixture model prior.
Protocol 2: CNN Model Interpretation with Grad-CAM
Objective: To generate visual explanations for a trained sperm morphology CNN classifier using Grad-CAM, verifying that it focuses on clinically relevant features.
Materials:
- Trained CNN Classifier: A model (e.g., based on Xception, EfficientNet) trained for normal/abnormal sperm classification or specific defect categorization [45].
- Input: Pre-processed sperm images from the test set.
Methodology:
- Model Inference and Feature Extraction:
- Forward-pass a test image through the trained CNN.
- Extract the final convolutional layer's feature maps and the corresponding output gradient for the target class (e.g., "amorphous head").
Grad-CAM Heatmap Calculation:
- Compute the neuron importance weights (
α_c^k) by global average pooling the gradients flowing back from the target class. - Perform a weighted combination of the feature maps using the calculated
α_c^kweights. - Apply a ReLU activation to the combined map to focus on features that have a positive influence on the class of interest.
L_{Grad-CAM}^c = ReLU(∑_k α_c^k A^k)
- Compute the neuron importance weights (
Visualization and Overlay:
- Upsample the calculated Grad-CAM heatmap to the original input image size.
- Overlay the heatmap onto the original sperm image using a color jet scheme (e.g., red for high-activation regions, blue for low).
- The resulting visualization highlights the spatial regions (e.g., head, midpiece) that the model deemed most important for its classification decision [44] [45].
Figure 2: Process for generating model explanations with Grad-CAM.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Computational Materials and Datasets for Sperm Morphology AI Research
| Item Name | Type | Function / Application | Example / Reference |
|---|---|---|---|
| SVIA Dataset | Public Dataset | Large-scale dataset with 125k+ instances for detection, segmentation, and classification tasks. | [9] |
| VISEM-Tracking | Public Dataset | Multi-modal dataset with videos & 656k+ annotated objects for tracking and motion analysis. | [9] [27] |
| SMD/MSS Dataset | Public Dataset | A dataset of 1k images (expandable) classified per modified David criteria, covering head, midpiece, and tail defects. | [2] |
| RAL Staining Kit | Wet Lab Reagent | Standardized staining of sperm smears to enhance morphological feature contrast for image acquisition. | [2] |
| MMC CASA System | Hardware/Software | Computer-Assisted Semen Analysis system for automated image acquisition and initial morphometric analysis. | [2] |
| Grad-CAM | Software Algorithm | Generates visual explanations for CNN decisions, critical for model validation and trust. | [44] [45] |
| GammaMM-VAE | Software Algorithm | A deep clustering model for unsupervised discovery of morphological subgroups in the latent space. | [46] |
| PyRadiomics | Software Library | Extracts handcrafted radiomic features (texture, shape) which can be fused with deep features for enriched analysis. | [45] |
Overcoming Challenges: Data, Model, and Computational Optimization
Addressing Data Scarcity and Class Imbalance in Medical Imaging
The implementation of Convolutional Neural Networks (CNNs) for human sperm morphology classification represents a paradigm shift in male fertility diagnostics. Traditional manual assessment is highly subjective, time-consuming, and suffers from significant inter-observer variability, with reported disagreement rates of up to 40% between expert evaluators [20] [5]. CNNs offer the potential for automated, standardized, and rapid analysis [48]. However, the development of robust, generalizable models is critically hampered by two interconnected challenges: medical data scarcity, arising from difficulties in acquiring and annotating specialized medical images, and class imbalance, inherent in medical datasets where abnormal findings are often underrepresented compared to normal cases [49] [20]. This document provides detailed application notes and experimental protocols to address these challenges within the context of CNN-based sperm morphology research.
Application Notes
Understanding the Data Landscape in Sperm Morphology Analysis
The core challenge in developing automated sperm morphology analysis systems lies in creating models that can accurately segment and classify sperm components (head, midpiece, tail) across a wide range of morphological anomalies [20]. The 2025 WHO guidelines require the analysis of over 200 sperm per sample, classifying them into categories including normal, head defects, neck/midpiece defects, tail defects, and excess residual cytoplasm [35]. This task is complicated by significant inter-expert disagreement during the annotation phase, with one study reporting only partial agreement (2/3 experts) on labels for a significant portion of data [2].
Table 1: Summary of Key Sperm Morphology Datasets for CNN Research
| Dataset Name | Sample Size (Images) | Classes/Categories | Notable Features | Reported Model Performance |
|---|---|---|---|---|
| SMD/MSS [2] | 1,000 (extended to 6,035 via augmentation) | 12 classes (Head, Midpiece, Tail defects) based on modified David classification | Expert annotations from 3 embryologists; comprehensive defect coverage | Accuracy: 55% - 92% (varies by class) |
| MHSMA [20] | 1,540 | Focus on acrosome, head shape, vacuoles | Non-stained, lower-resolution images | F0.5 Scores: Acrosome (84.74%), Head (83.86%), Vacuoles (94.65%) |
| SVIA [20] | 125,000 annotated instances | Object detection, segmentation, classification | Large-scale, multi-task dataset | Detailed metrics not fully reported |
| SMIDS [5] | 3,000 | 3-class | Used for benchmarking DL models | Test Accuracy: 96.08% with CBAM-ResNet50 + Feature Engineering |
| HuSHeM [5] | 216 | 4-class | Public benchmark for sperm head morphology | Test Accuracy: 96.77% with CBAM-ResNet50 + Feature Engineering |
Technical Strategies for Data Scarcity and Class Imbalance
The following technical strategies have demonstrated efficacy in mitigating data-related challenges in medical imaging.
Data-Centric Strategies
- Data Augmentation: Fundamental for combating data scarcity. Geometric transformations (rotation, flipping, scaling) and photometric adjustments (brightness, contrast) are standard. In sperm morphology analysis, augmenting a dataset from 1,000 to 6,035 images has been successfully used to train a CNN model, enabling it to learn invariant features [2].
- Advanced Data Synthesis: For complex class imbalances, particularly with rare morphological defects, generative models such as Generative Adversarial Networks (GANs) can create synthetic, high-fidelity sperm images to balance the representation of rare anomaly classes [48].
- Cross-Domain Generalization: Meta-learning algorithms, such as the Contrastive Meta-Learning with Auxiliary Tasks (HSHM-CMA), learn invariant features across multiple tasks or datasets. This approach improves a model's ability to adapt to new data categories and different imaging conditions, achieving accuracies up to 81.42% in cross-dataset evaluations [43].
Algorithmic and Model-Centric Strategies
- Bi-Level Class Balancing (BLCB): This novel pipeline addresses imbalance at multiple levels. Level-I performs inter-class balancing (e.g., vessel vs. non-vessel pixels in retinal images, analogous to sperm vs. background), while Level-II performs intra-class balancing (e.g., thin vs. thick vessels, analogous to subtle vs. obvious sperm defects) [50]. This hierarchical approach significantly improves sensitivity for underrepresented classes.
- Attention Mechanisms: Integrating Convolutional Block Attention Modules (CBAM) into CNN architectures like ResNet50 forces the model to focus computational resources on the most informative spatial regions and feature channels, such as specific sperm head defects or tail anomalies. This leads to more efficient learning from limited data and improves classification accuracy [5].
- Hybrid Deep Feature Engineering (DFE): This strategy combines the strength of deep learning with traditional machine learning. Features are extracted using a pre-trained CNN (e.g., ResNet50-CBAM), followed by dimensionality reduction (e.g., PCA) and classical classifiers (e.g., SVM). This method has been shown to boost baseline CNN accuracy by over 8-10% on sperm morphology datasets [5].
- Loss Function Engineering: Using tailored loss functions like Focal Loss helps down-weight the contribution of easy-to-classify majority classes (e.g., normal sperm), forcing the model to focus training on hard-to-classify minority classes (e.g., specific rare defects) [50].
Experimental Protocols
Protocol 1: Building a Balanced Dataset for Sperm Morphology
Objective: To acquire and pre-process sperm images and systematically address class imbalance through data augmentation before model training.
Materials:
- Microscope with camera (e.g., MMC CASA system) [2]
- Stained semen smears (e.g., RAL Diagnostics staining kit) [2]
- Image annotation software (e.g., Roboflow) [35]
Workflow:
- Image Acquisition: Capture at least 200 sperm images per patient sample using a 100x oil immersion objective lens to ensure sufficient detail of head, midpiece, and tail [2] [35].
- Expert Annotation: Have a minimum of three trained embryologists independently classify each spermatozoon according to a standardized classification system (e.g., modified David or WHO). Resolve discrepancies through consensus review. Compile a ground truth file [2].
- Data Pre-processing:
- Cleaning: Remove images with overlapping sperm, debris, or only partial sperm structures [20].
- Normalization: Resize all images to a uniform size (e.g., 80x80 pixels). Apply grayscale conversion and normalization to scale pixel values to a common range (e.g., 0-1) [2].
- Contrast Enhancement: Apply techniques like Contrast Limited Adaptive Histogram Equalization (CLAHE) to improve feature visibility [50].
- Data Augmentation: For underrepresented morphological classes, apply a pipeline of augmentation operations to increase their sample count. The following diagram illustrates this sequential process.
Protocol 2: Implementing a CNN with Class Imbalance Mitigation
Objective: To train a CNN model for multi-class sperm morphology classification using a combination of data-level and algorithm-level techniques to handle imbalanced data.
Materials:
- High-performance computing workstation with GPU.
- Python 3.8+ with deep learning frameworks (TensorFlow, PyTorch).
- Public or institutionally curated sperm morphology dataset (e.g., from Table 1).
Workflow:
- Dataset Partitioning: Split the annotated and augmented dataset into training (80%), validation (10%), and hold-out test (10%) sets. Ensure stratification to maintain class distribution across splits [2].
- Model Architecture Design:
- Configure Imbalance Handling:
- Option A (Loss Function): Use a weighted loss function (e.g., Weighted Cross-Entropy, Focal Loss) where class weights are inversely proportional to their frequency in the training set [50].
- Option B (Hybrid DFE): Replace the model's classification layer with a feature extractor. Use the extracted deep features to train a classic classifier (e.g., SVM with RBF kernel) following dimensionality reduction with PCA [5].
- Model Training and Evaluation:
- Train the model using the Adam optimizer, monitoring validation loss for early stopping.
- Evaluate on the held-out test set using a comprehensive suite of metrics: Accuracy, Precision, Recall (Sensitivity), Specificity, and F1-Score. The confusion matrix should be analyzed to pinpoint specific class-wise performance drops.
The logical relationship and flow of these algorithmic strategies are summarized in the following diagram.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials and Reagents for Sperm Morphology CNN Research
| Item Name | Function/Application | Specification/Example |
|---|---|---|
| Optical Microscope with Camera | Image acquisition from semen smears. | MMC CASA system with 100x oil immersion objective [2]. |
| Sperm Staining Kit | Provides contrast for morphological feature visualization. | RAL Diagnostics kit [2]. |
| Image Annotation Software | Platform for expert labeling of sperm images; crucial for ground truth creation. | Roboflow [35]. |
| Trumorph System | Dye-free fixation of sperm for morphology evaluation using pressure and temperature [35]. | Standardized preparation for consistent imaging. |
| High-Performance Computing Unit | Training and evaluating complex CNN models. | Workstation with high-end GPU (e.g., NVIDIA RTX series). |
| Deep Learning Framework | Software environment for building and training CNN models. | Python with TensorFlow/PyTorch libraries [2] [5]. |
| Public Benchmark Datasets | For model benchmarking, validation, and transfer learning. | SMIDS, HuSHeM, SMD-MSS [2] [20] [5]. |
Mitigating Overfitting in Deep Networks with Limited Data
The application of Convolutional Neural Networks (CNNs) for human sperm morphology classification presents a classic challenge in medical artificial intelligence (AI): achieving high model accuracy with limited training data. In this context, overfitting occurs when a model learns the training data too well, including its noise and random fluctuations, but fails to generalize to new, unseen data [51] [52]. This undesirable machine learning behavior manifests when a model gives accurate predictions for training data but not for novel data, significantly limiting its clinical utility [51].
The problem is particularly pronounced in sperm morphology analysis, where the creation of large, standardized, high-quality annotated datasets remains fundamentally challenging [20]. Expert annotation is time-consuming, requires specialized expertise, and suffers from inter-observer variability [2] [20]. Furthermore, sperm morphology assessment requires simultaneous evaluation of head, vacuoles, midpiece, and tail abnormalities, substantially increasing annotation complexity [20]. These constraints make overfitting mitigation strategies not merely beneficial but essential for developing robust, clinically applicable deep learning models for reproductive medicine.
Quantitative Landscape: Data and Performance in Sperm Morphology Studies
Table 1: Performance of Deep Learning Models in Sperm Morphology Analysis
| Study/Dataset | Original Sample Size | Augmented Sample Size | Reported Accuracy Range | Key Morphological Classes |
|---|---|---|---|---|
| SMD/MSS Dataset [2] | 1,000 images | 6,035 images | 55% to 92% | 12 classes (7 head defects, 2 midpiece defects, 3 tail defects) |
| MHSMA Dataset [20] | 1,540 images | Not specified | Not specified | Acrosome, head shape, vacuoles |
| SVIA Dataset [20] | 125,000 annotated instances | Not applicable | Not specified | Comprehensive detection, segmentation, and classification |
Table 2: Overfitting Mitigation Techniques and Their Relative Effectiveness
| Technique Category | Specific Methods | Reported Effectiveness | Implementation Complexity |
|---|---|---|---|
| Data-Level Strategies | Data Augmentation, Adding Noise to Input/Output [52] | High | Low |
| Model Architecture Strategies | Dropout, DropConnect, Simplified Models, Transition Modules [53] [54] [55] | Medium to High | Medium |
| Training Process Strategies | Early Stopping, K-fold Cross-Validation, Regularization (L1/L2) [51] [53] [52] | Medium | Low to Medium |
| Advanced Learning Paradigms | Contrastive Meta-Learning with Auxiliary Tasks (HSHM-CMA) [43] | High (65.83% to 81.42% across generalization tasks) | High |
The quantitative evidence reveals both the challenges and opportunities in this domain. The SMD/MSS dataset demonstrates how data augmentation can expand a limited dataset six-fold, from 1,000 to 6,035 images, enabling more robust model training [2]. However, the wide accuracy range (55% to 92%) highlights the significant performance variability that can result from different methodological approaches and the fundamental difficulty of the classification task itself [2].
Recent advances in specialized learning algorithms show particular promise for addressing the generalization challenge. The HSHM-CMA (Contrastive Meta-learning with Auxiliary Tasks) algorithm has demonstrated robust performance across multiple testing objectives, achieving 65.83% accuracy on the same dataset with different sperm head morphology categories, 81.42% on different datasets with the same categories, and 60.13% on different datasets with different categories [43]. This cross-domain generalization capability is particularly valuable for clinical applications where model deployment conditions often differ from training environments.
Experimental Protocols for Overfitting Mitigation
Data Augmentation and Preprocessing Protocol
A critical first step in addressing limited data is the implementation of a comprehensive data augmentation strategy. The following protocol, adapted from successful implementations in sperm morphology analysis, provides a systematic approach to expanding training datasets:
Image Acquisition and Preprocessing: Acquire sperm images using a CASA (Computer-Assisted Semen Analysis) system with bright field mode and an oil immersion x100 objective [2]. Convert all images to grayscale and resize to 80×80 pixels using linear interpolation to normalize dimensions and reduce computational complexity [2].
Data Cleaning and Normalization: Identify and handle missing values, outliers, or inconsistencies in the dataset. Normalize or standardize numerical features to bring them to a common scale, ensuring no particular feature dominates the learning process due to magnitude differences [2].
Augmentation Implementation: Apply multiple transformation techniques to the training dataset:
Dataset Partitioning: Split the augmented dataset into training (80%), validation (10%), and testing (10%) subsets, ensuring representative distribution of morphological classes across partitions [2].
This augmentation protocol systematically increases dataset diversity and size, providing the model with varied examples of each morphological class and reducing the risk of learning dataset-specific artifacts.
CNN Architecture with Integrated Regularization
This protocol details the implementation of a CNN architecture with built-in regularization mechanisms specifically designed for sperm morphology classification:
Base Network Configuration:
- Implement a CNN architecture using Python 3.8 and deep learning frameworks (e.g., TensorFlow, PyTorch) [2] [53]
- Design with 3-5 convolutional layers, progressively increasing filters (32, 64, 128)
- Use 3×3 kernels throughout with ReLU activation functions
- Incorporate max-pooling layers (2×2) after each convolutional block
Transition Module Integration:
- Replace the standard connection between final convolutional and fully connected layers with a transition module [54]
- Implement parallel convolutional layers with varying filter sizes (1×1, 3×3, 5×5)
- Apply global average pooling to each filter pathway to reduce spatial dimensions
- Concatenate outputs before passing to fully connected layers [54]
Regularization Implementation:
Training Configuration:
- Use Adam or RMSProp optimizers with learning rate 0.001
- Implement early stopping by monitoring validation loss with patience of 10-15 epochs
- Train for maximum 100 epochs with batch size 32-64 [2]
This architectural approach incorporates multiple regularization strategies at different levels of the network, creating a robust framework resistant to overfitting even with limited data.
CNN Architecture with Transition Module for Overfitting Mitigation
Advanced Meta-Learning Protocol
For researchers facing extreme data limitations or needing cross-domain generalization, this protocol outlines the implementation of contrastive meta-learning with auxiliary tasks:
Task Formulation:
- Define a set of meta-training tasks from available sperm morphology data
- Separate tasks into primary (main classification objectives) and auxiliary tasks (related but distinct learning objectives) [43]
- Ensure task diversity to encourage learning of invariant features
HSHM-CMA Algorithm Implementation:
Training and Validation:
- Meta-train across diverse task distributions
- Validate using three testing objectives:
- Same dataset with different HSHM categories
- Different datasets with same HSHM categories
- Different datasets with different HSHM categories [43]
- Fine-tune on target tasks with limited samples
This advanced approach moves beyond conventional regularization, fundamentally restructuring the learning process to excel in data-scarce environments through knowledge transfer across related tasks.
Table 3: Essential Research Reagents and Computational Resources
| Resource Category | Specific Tool/Platform | Function in Research | Implementation Example |
|---|---|---|---|
| Data Acquisition | MMC CASA System [2] | Automated sperm image capture and morphometric analysis | Acquire 1000+ individual sperm images per study |
| Staining Reagents | RAL Diagnostics Staining Kit [2] | Sperm staining for morphological visualization | Prepare semen smears according to WHO guidelines |
| Annotation Framework | Modified David Classification [2] | Standardized morphological defect categorization | Classify 12 defect types across head, midpiece, and tail |
| Computational Environment | Python 3.8 [2] | Algorithm development and implementation | Build CNN architectures with TensorFlow/PyTorch |
| Cloud Platforms | Amazon SageMaker [51], Tencent Cloud AI Platform [53] | Managed training with automatic overfitting detection | Enable early stopping and real-time training metrics |
| Validation Framework | K-fold Cross-Validation [51] [52] | Robust performance assessment and overfitting detection | Partition data into 5-10 folds for iterative validation |
The mitigation of overfitting in deep networks for sperm morphology classification requires a multifaceted approach that addresses data limitations, model architecture, and training methodologies. The strategies outlined in this document—from fundamental data augmentation to advanced meta-learning techniques—provide a comprehensive framework for developing robust models capable of generalizing well to clinical data.
Future research directions should focus on the development of more sophisticated domain adaptation techniques, federated learning approaches to leverage distributed data while maintaining privacy, and explainable AI methods to build clinician trust in model predictions. As these technologies mature, they hold significant promise for standardizing sperm morphology analysis, reducing inter-laboratory variability, and ultimately improving diagnostic accuracy in male fertility assessment worldwide.
Comprehensive Workflow for Mitigating Overfitting
The implementation of Convolutional Neural Networks (CNNs) for human sperm morphology classification within clinical environments presents significant challenges regarding processing speed and hardware constraints. Clinical laboratories require solutions that deliver high diagnostic accuracy while operating within practical timeframes and using computationally efficient hardware. This document provides detailed application notes and protocols for optimizing CNN-based sperm analysis systems to meet these clinical demands, ensuring robust performance without compromising diagnostic reliability.
Performance Benchmarks and Computational Requirements
Quantitative Performance of CNN Architectures
Table 1: Performance comparison of deep learning models for sperm analysis tasks
| Model Architecture | Task | Accuracy/Performance Metrics | Dataset | Computational Notes |
|---|---|---|---|---|
| EfficientNetV2 Ensemble (Feature-level + Decision-level fusion) | Sperm morphology classification (18 classes) | 67.70% accuracy | Hi-LabSpermMorpho (18,456 images) | Combines multiple EfficientNetV2 variants; feature fusion with SVM, RF, and MLP-Attention classifiers [32] |
| VGG16 with Transfer Learning | Sperm head classification | High accuracy (exact % not specified) | HuSHeM and SCIAN datasets | Retrained on ImageNet weights; avoids excessive computation compared to dictionary learning methods [33] |
| Mask R-CNN | Multi-part sperm segmentation | Highest IoU for head, nucleus, acrosome | Live unstained human sperm dataset | Robust for smaller, regular structures; two-stage architecture demands more resources [30] |
| U-Net | Sperm tail segmentation | Highest IoU for morphologically complex tail | Live unstained human sperm dataset | Global perception and multi-scale feature extraction; efficient for elongated structures [30] |
| YOLOv8 | Multi-part sperm segmentation | Comparable/slightly better than Mask R-CNN for neck segmentation | Live unstained human sperm dataset | Single-stage model with faster inference times [30] |
Processing Speed Considerations
For clinical workflows, the inference time per image is a critical metric. While specific frame-per-second rates for sperm analysis are not provided in the search results, comparative architectural insights inform selection criteria:
- Ensemble methods (as used with EfficientNetV2) typically increase processing time proportional to the number of base models but can be optimized through parallel processing [32].
- Single-stage detectors (YOLOv8, YOLO11) generally provide faster inference compared to two-stage detectors (Mask R-CNN), making them suitable for real-time applications [30].
- U-Net architectures offer efficient segmentation for complex structures like sperm tails, balancing accuracy and computational demand [30].
Experimental Protocols for Clinical Implementation
Protocol 1: Implementing Ensemble CNN for Morphology Classification
Objective: To achieve high-accuracy sperm morphology classification using an ensemble approach while optimizing for clinical processing speeds.
Materials:
- Hi-LabSpermMorpho dataset or comparable clinical dataset
- Python 3.8+ with TensorFlow 2.0+ or PyTorch
- GPU-enabled workstation (minimum NVIDIA Titan RTX or equivalent)
- Pre-trained EfficientNetV2 models (S, M, L variants)
Procedure:
- Data Preprocessing:
- Resize all images to uniform dimensions (e.g., 224×224 or 384×384 pixels)
- Apply normalization (pixel values 0-1 or standardized using dataset statistics)
- Implement data augmentation: random rotation (±15°), horizontal/vertical flipping, brightness/contrast variation (±10%)
Feature Extraction:
- Load multiple pre-trained EfficientNetV2 models without classification heads
- Extract features from penultimate layers for all training images
- Apply dimensionality reduction (PCA or dense layer transformation) to manage computational load
Classifier Training:
- Implement feature-level fusion by concatenating reduced features from all models
- Train multiple classifiers (SVM, Random Forest, MLP-Attention) on fused features
- Optimize hyperparameters using grid search or Bayesian optimization
Decision-Level Fusion:
- Generate prediction probabilities from all trained classifiers
- Implement soft voting mechanism to combine probabilities
- Select final classification based on highest consensus probability
Performance Validation:
- Evaluate on holdout test set using accuracy, precision, recall, and F1-score
- Measure average processing time per image across hardware configurations
- Compare against individual classifiers to quantify ensemble improvement [32]
Protocol 2: Multi-Part Sperm Segmentation for Clinical Deployment
Objective: To accurately segment sperm components (head, acrosome, nucleus, neck, tail) with optimized inference speed for clinical use.
Materials:
- Annotated dataset of live unstained human sperm
- GPU with minimum 8GB VRAM (NVIDIA RTX 3000+ series recommended)
- Deep learning framework (TensorFlow or PyTorch) with Detectron2 for Mask R-CNN
Procedure:
- Model Selection Strategy:
- For head, acrosome, and nucleus segmentation: Implement Mask R-CNN for highest IoU
- For tail segmentation: Implement U-Net for optimal performance on elongated structures
- For balanced speed/accuracy across all components: Implement YOLOv8 or YOLO11
Training Configuration:
- Initialize with pre-trained weights on COCO or ImageNet datasets
- Set initial learning rate: 0.001 with cosine decay scheduling
- Batch size: 8-16 (adjust based on GPU memory)
- Data augmentation: random scaling, rotation, color jittering
Inference Optimization:
- Implement model quantization (FP16) for faster inference
- Use TensorRT or OpenVINO for hardware-specific acceleration
- Optimize input pipeline to minimize preprocessing overhead
Validation Metrics:
- Calculate IoU, Dice coefficient, Precision, Recall for each sperm component
- Benchmark inference time per image on target clinical hardware
- Compare against manual segmentation by embryologists [30]
Implementation Workflow
The following diagram illustrates the optimized clinical workflow for CNN-based sperm morphology analysis:
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential materials and computational resources for CNN-based sperm analysis
| Item | Function/Application | Implementation Notes |
|---|---|---|
| Hi-LabSpermMorpho Dataset | Training and validation of morphology classification models | Contains 18 distinct morphology classes with 18,456 image samples; addresses class imbalance [32] |
| Live Unstained Human Sperm Dataset | Segmentation model development and validation | Enables clinically relevant segmentation without staining artifacts [30] |
| EfficientNetV2 Pre-trained Models | Feature extraction backbone | Multiple variants (S, M, L) balance accuracy and computational efficiency [32] |
| Mask R-CNN Framework | Instance segmentation of sperm components | Superior for head, nucleus, acrosome segmentation; Detectron2 implementation recommended [30] |
| U-Net Architecture | Semantic segmentation of complex structures | Optimal for tail segmentation; global perception capabilities [30] |
| SVM, Random Forest, MLP-Attention | Ensemble classification | Combined via feature-level and decision-level fusion for improved accuracy [32] |
| TensorFlow/PyTorch | Deep learning framework | GPU-accelerated model training and inference [58] |
| NVIDIA GPU (8GB+ VRAM) | Model training and inference | Titan RTX or RTX 3000+ series recommended for efficient processing [59] |
Hardware Optimization Strategies
Clinical Deployment Configurations
Table 3: Hardware recommendations based on clinical requirements
| Clinical Scenario | Recommended Hardware | Expected Performance | Implementation Considerations |
|---|---|---|---|
| High-throughput fertility clinic | Multiple NVIDIA RTX 4090 or A100 GPUs | Real-time processing (<1 second per image) | Batch processing capabilities; parallel model inference |
| Medium-sized laboratory | Single NVIDIA RTX 3080/4080 (12-16GB VRAM) | Near real-time (2-3 seconds per image) | Model quantization; optimized inference pipelines |
| Point-of-care or remote clinic | NVIDIA Jetson Orin or consumer-grade GPU (RTX 3060) | Acceptable processing (5-10 seconds per image) | Pruned models; INT8 quantization; cloud offloading options |
Speed-Accuracy Tradeoff Optimization
- Model Pruning: Remove redundant weights from trained networks to reduce computational load while preserving accuracy
- Quantization: Implement FP16 or INT8 precision for inference to accelerate processing on supported hardware
- Knowledge Distillation: Train smaller "student" models to mimic larger "teacher" ensemble models
- Hardware-Specific Optimization: Utilize TensorRT (NVIDIA) or OpenVINO (Intel) for deployment acceleration
Validation Protocol for Clinical Deployment
Objective: To ensure optimized models maintain diagnostic reliability in clinical settings.
Procedure:
- Performance Baseline:
- Compare model performance against manual assessment by multiple embryologists
- Establish minimum acceptable accuracy thresholds for each morphology class
Speed Validation:
- Measure end-to-end processing time for batch of 100 images
- Verify processing meets clinical workflow requirements (<5 minutes for full analysis)
Hardware Stress Testing:
- Evaluate performance under sustained load (8+ hours continuous operation)
- Monitor thermal throttling and memory utilization patterns
Clinical Validation:
These protocols provide a comprehensive framework for implementing CNN-based sperm morphology analysis systems that balance computational efficiency with clinical diagnostic requirements, enabling seamless integration into diverse laboratory environments.
Ensuring Generalizability Across Different Imaging Setups and Populations
The implementation of Convolutional Neural Networks (CNNs) for human sperm morphology classification represents a significant advancement in male fertility research and diagnostic medicine. However, a critical challenge persists: model generalizability across diverse imaging setups and patient populations. The performance of deep learning models can be substantially compromised by variations in staining protocols, microscope configurations, imaging conditions, and population-specific characteristics [62] [20]. This application note addresses these challenges by providing detailed protocols and methodologies specifically designed to enhance model robustness and ensure reliable performance across different clinical environments and demographic groups. Establishing generalizable models is paramount for clinical adoption, as it ensures consistent diagnostic accuracy regardless of the specific imaging setup or patient population being analyzed, ultimately leading to more reliable male fertility assessments worldwide.
The foundation of any generalizable CNN model lies in comprehensive, diverse, and well-characterized datasets. Significant variability exists across publicly available and research-specific datasets for sperm morphology analysis, impacting model performance when applied to new populations or imaging setups.
Table 1: Comparison of Key Sperm Morphology Datasets Highlighting Sources of Variability
| Dataset Name | Sample Size | Classes/Categories | Staining Method(s) | Imaging Setup | Reported Model Performance (Accuracy) |
|---|---|---|---|---|---|
| SMD/MSS [2] | 1,000 (extended to 6,035 with augmentation) | 12 classes (Modified David classification) | RAL Diagnostics staining kit | MMC CASA system, bright field, 100x oil objective | 55% to 92% (Deep Learning Model) |
| Hi-LabSpermMorpho [62] | 18 categories across 3 staining types | Head, neck, and tail abnormalities | Three Diff-Quick techniques (BesLab, Histoplus, GBL) | Bright-field microscopy with mobile phone camera | 68.41% to 71.34% (Two-Stage Ensemble Model) |
| MHSMA [20] | 1,540 images | Focus on acrosome, head shape, vacuoles | Information Not Specified | Information Not Specified | Information Not Specified |
| HuSHeM [12] | Information Not Specified | Information Not Specified | Information Not Specified | Information Not Specified | 97.78% (DenseNet169) |
| SCIAN-Morpho [12] | Information Not Specified | Information Not Specified | Information Not Specified | Information Not Specified | 78.79% (DenseNet169) |
| SVIA [20] | 125,000 annotated instances | Object detection, segmentation, classification | Information Not Specified | Information Not Specified | Information Not Specified |
The variability in datasets leads directly to performance discrepancies. For instance, the same DenseNet169 architecture showed a significant performance drop from 97.78% on the HuSHeM dataset to 78.79% on the SCIAN-Morpho dataset, underscoring the profound effect of dataset-specific characteristics on model generalizability [12]. Furthermore, staining protocol differences alone can cause accuracy variations of approximately 3% as observed in the Hi-LabSpermMorpho dataset evaluations [62]. These discrepancies highlight the necessity of standardized protocols and robust methodological approaches to overcome dataset-specific biases.
Experimental Protocols for Enhanced Generalizability
Protocol: Multi-Staining Model Training and Validation
Objective: To develop a CNN model robust to variations in sperm staining protocols, specifically addressing the challenges posed by different chemical compositions and color profiles.
Materials:
- Hi-LabSpermMorpho dataset or equivalent containing images from at least three staining protocols (e.g., BesLab, Histoplus, GBL) [62]
- Computational framework: Python with deep learning libraries (TensorFlow, PyTorch)
- Pre-trained CNN architectures (NFNet, Vision Transformer, DenseNet)
Procedure:
- Data Partitioning: Split each staining-specific dataset (BesLab, Histoplus, GBL) independently into training (80%), validation (10%), and test (10%) sets, ensuring no patient overlap between sets.
- Staining-Balanced Training: During each training batch, sample an equal number of images from each staining protocol to prevent model bias toward any single staining type.
- Data Augmentation: Apply stain-specific and stain-agnostic augmentations:
- Stain-Agnostic: Random rotations (±15°), flips, brightness/contrast variations (±10%), Gaussian noise
- Stain-Specific: Color space transformations in LAB space to simulate staining intensity variations
- Model Architecture:
- Implement a two-stage ensemble framework [62]
- Stage 1 - Splitting: Train a classifier to categorize input images into two major groups: (1) head and neck abnormalities, and (2) normal morphology with tail abnormalities
- Stage 2 - Specialized Ensembles: Route images to staining-specific ensemble models incorporating multiple architectures (NFNet, ViT)
- Validation: Employ k-fold cross-validation (k=5) within each staining protocol and report performance metrics separately for each protocol and overall.
Quality Control:
- Compute inter-observer agreement metrics between expert annotations for each staining type
- Establish a minimum acceptable accuracy threshold for each staining protocol (e.g., >65% based on reported results [62])
Protocol: Cross-Dataset Validation Framework
Objective: To objectively evaluate model performance across different populations and imaging setups through rigorous cross-dataset validation.
Materials:
- At least two distinct sperm morphology datasets with different population sources (e.g., SMD/MSS, Hi-LabSpermMorpho, HuSHeM)
- A trained model from Protocol 3.1
Procedure:
- Dataset Harmonization:
- Resize all images to a uniform resolution (e.g., 80×80 pixels as in SMD/MSS [2])
- Convert all images to grayscale to reduce color variance from different staining protocols
- Apply histogram equalization to standardize contrast levels across datasets
- Performance Benchmarking:
- Train the model on the complete training set of Dataset A
- Evaluate the trained model on the test sets of both Dataset A and Dataset B
- Calculate the performance drop: ΔAccuracy = AccuracyDatasetA - AccuracyDatasetB
- Domain Adaptation (if ΔAccuracy > 10%):
- Implement a domain adaptation approach by fine-tuning the model with a small subset (10-15%) of Dataset B
- Compare performance before and after domain adaptation
Quality Control:
- Report not only overall accuracy but also class-specific F1 scores for each dataset
- Perform statistical testing (e.g., McNemar's test) to determine if performance differences are significant
Protocol: Assessment of Inter-Expert Agreement
Objective: To quantify and account for the inherent subjectivity in sperm morphology classification, establishing a reliability baseline for model training.
Materials:
- A curated set of at least 200 sperm images representing diverse morphological classes
- Three independent experts with extensive experience in semen analysis
Procedure:
- Blinded Annotation: Provide each expert with the same set of images in randomized order with no communication between experts during the annotation process.
- Annotation Schema: Use a detailed classification schema based on established criteria (e.g., modified David classification with 12 classes as in SMD/MSS [2]).
- Agreement Calculation:
- Calculate Total Agreement (TA): Percentage of images where all three experts assign identical labels across all categories
- Calculate Partial Agreement (PA): Percentage of images where two of three experts agree on the same label for at least one category
- Calculate No Agreement (NA): Percentage of images with no consensus among experts
- Statistical Analysis:
- Use IBM SPSS Statistics or equivalent for Fisher's exact test to evaluate statistical differences between experts for each morphology class (considered significant at p < 0.05)
- Compute Fleiss' Kappa to measure inter-rater reliability beyond chance agreement
Quality Control:
- Establish ground truth based on majority voting (at least 2/3 experts) for model training labels
- Exclude images falling into the "No Agreement" category from training data or flag them for expert review
Technical Solutions for Generalizability
Two-Stage Divide-and-Ensemble Architecture
The two-stage divide-and-ensemble framework represents a significant architectural innovation for enhancing generalizability [62]. This approach decomposes the complex task of fine-grained sperm classification into manageable sub-tasks, reducing the model's vulnerability to domain-specific variations.
This architectural approach demonstrated a 4.38% improvement in accuracy over conventional single-model approaches in cross-staining evaluations [62]. The structured multi-stage voting mechanism further enhances decision reliability by allowing models to use both primary and secondary votes, mitigating the influence of dominant classes and ensuring more balanced decision-making across different sperm abnormalities.
Advanced Data Augmentation and Preprocessing
Strategic data augmentation is crucial for simulating population and imaging variability. Beyond standard transformations (rotation, flipping), the following advanced techniques specifically address generalizability challenges in sperm morphology analysis:
- Stain Normalization: CycleGAN or similar approaches to transform staining profiles between different laboratory protocols
- Multi-Scale Feature Extraction: Employing atrous spatial pyramid pooling (ASPP) to capture morphological features at various scales
- Quantitative Phase Imaging (QPI) Integration: For subcellular feature enhancement beyond bright-field limitations [63]
- MotionFlow Representation: Capturing temporal motility characteristics alongside static morphology [27]
The SMD/MSS dataset successfully expanded their dataset from 1,000 to 6,035 images through comprehensive augmentation, enabling more robust model training [2]. Preprocessing should include image denoising to address insufficient lighting or poorly stained smears, followed by normalization to bring all images to a common scale, typically resizing to 80×80 pixel grayscale images with linear interpolation [2].
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Research Reagents and Materials for Generalizable Sperm Morphology Analysis
| Reagent/Material | Function/Application | Implementation Notes |
|---|---|---|
| Diff-Quick Staining Kits (BesLab, Histoplus, GBL) | Standardized staining for morphological feature enhancement | Use multiple staining protocols during training to improve model robustness to staining variations [62] |
| RAL Diagnostics Staining Kit | Sperm smear preparation and staining | Follow WHO manual guidelines for smear preparation consistency [2] |
| SonoVue Contrast Agent | For contrast-enhanced ultrasound applications | Used in developing complementary diagnostic models for reproductive health [64] |
| MMC CASA System | Computer-Assisted Semen Analysis for image acquisition | Enables sequential image acquisition with standardized morphometric tools [2] |
| Partially Spatially Coherent Digital Holographic Microscope (PSC-DHM) | Quantitative phase imaging for subcellular analysis | Provides nanometric sensitivity for detecting subtle morphological changes in head, midpiece, and tail [63] |
| Python 3.8 with Deep Learning Libraries | Model development and implementation | Core environment for CNN implementation; essential for reproducibility [2] |
| Pre-trained CNN Models (NFNet, ViT, DenseNet) | Transfer learning for limited data scenarios | DenseNet169 effectively addresses gradient vanishing and improves feature efficiency [12] |
Ensuring generalizability across different imaging setups and populations is not merely a technical challenge but a fundamental requirement for the clinical adoption of CNN-based sperm morphology classification systems. The protocols and methodologies presented herein—including multi-staining training, cross-dataset validation, inter-expert agreement assessment, and innovative two-stage architectures—provide a comprehensive framework for developing robust models. Implementation of these strategies will significantly advance the field toward reliable, population-agnostic sperm morphology analysis, ultimately enhancing male fertility assessment accuracy and consistency across diverse clinical environments and patient populations worldwide.
Fundamental Principles and Architecture
The Hybrid Morphological-Convolutional Neural Network (MCNN) represents a specialized machine learning architecture designed to overcome the significant challenge of limited dataset size in medical image analysis. Unlike conventional deep Convolutional Neural Networks (CNNs) that require extensive computational resources and large volumes of training data, the MCNN integrates mathematical morphology operations directly into a compact network structure. This hybrid approach enables effective learning from medical image datasets containing only a few hundred samples, which is a common constraint in clinical settings where data acquisition is complex, costly, and time-consuming [65] [66].
The MCNN architecture strategically combines the strengths of convolutional layers with morphological image processing to enhance feature extraction capabilities. While standard CNNs utilize small kernels to capture textural information, they often struggle with computational complexity and overfitting when trained on limited medical datasets. The MCNN addresses these limitations by incorporating morphological operations that excel at highlighting specific geometrical structures within images, making it particularly suitable for medical conditions where diagnosis relies on recognizing distinct morphological features such as the relationship between optic disc and cup sizes in glaucoma or irregular borders and color variability in melanoma [65]. This architectural innovation provides a practical solution for researchers and clinicians working with constrained medical image datasets across various diagnostic applications.
MCNN Protocol for Medical Image Analysis
System Architecture and Workflow
The following diagram illustrates the comprehensive workflow of the Hybrid Morphological-Convolutional Neural Network (MCNN) system for medical image analysis:
Figure 1: MCNN System Architecture for Medical Image Analysis
Experimental Implementation Protocol
Data Preparation and Preprocessing
- Image Acquisition: Source medical images according to clinical standards. For sperm morphology analysis, this may involve bright-field microscopy images following WHO staining protocols [9].
- Dataset Partitioning: Divide the available dataset into training, validation, and testing subsets. Given the small dataset constraint typical of medical applications, employ cross-validation techniques with recommended ratios of 70:25:5 for optimal performance [12].
- Color Channel Separation: Decompose each medical image into its constituent red, green, and blue color channels for independent processing through parallel neural networks [65].
Network Configuration and Training
- Independent Channel Networks: Implement three separate neural networks, each dedicated to processing one color channel. These networks employ Extreme Learning Machine (ELM) algorithm for efficient training [65].
- Morphological Operations Integration: Incorporate morphological layers alongside standard convolutional operations to enhance feature extraction of geometrically significant structures.
- Feature Fusion: Combine probability outputs from all three channel networks into a consolidated probability matrix measuring 3×n dimensions, where n represents the number of target classes [65].
Classification and Validation
- Random Forest Classification: Feed the fused probability matrix into a Random Forest classifier for final diagnostic decision-making [65].
- Performance Validation: Evaluate system performance using appropriate statistical measures including Area Under Curve (AUC), sensitivity, specificity, and overall accuracy with confidence intervals [65].
Performance Analysis and Comparative Evaluation
Quantitative Performance Metrics
Table 1: MCNN Performance Across Medical Imaging Applications
| Medical Application | Dataset | Performance Metrics | Comparative CNN Architectures | Key Findings |
|---|---|---|---|---|
| Melanoma Classification | ISIC Dataset | AUC: 0.94 (95% CI: 0.91 to 0.97) [65] | ResNet-18, ShuffleNet-V2, MobileNet-V2 | MCNN outperformed all popular CNN architectures [65] |
| Glaucoma Classification | ORIGA Dataset | AUC: 0.65 (95% CI: 0.53 to 0.74) [65] | ResNet-18, ShuffleNet-V2, MobileNet-V2 | Performance similar to popular CNN architectures [65] |
| Sperm Morphology Classification | HuSHeM Dataset | Accuracy: Up to 97.78% with DenseNet169 [12] | Various CNN and hybrid approaches | Demonstrates potential for male infertility diagnosis [12] |
Advantages Over Conventional Deep Learning Approaches
The MCNN architecture provides several distinct advantages for medical image analysis compared to conventional deep learning approaches:
- Data Efficiency: MCNN achieves effective performance with medical image datasets limited to a few hundred samples, unlike deep CNNs which typically require thousands of training images [65].
- Computational Economy: The compact architecture reduces memory requirements and computational resources while maintaining diagnostic accuracy [65].
- Clinical Interpretability: The integration of morphological operations aligns with clinical reasoning processes that often focus on structural characteristics, potentially enhancing trust among medical professionals [65].
- Architectural Flexibility: The MCNN framework can be adapted to various medical imaging modalities by selecting appropriate morphological operations targeted to specific diagnostic features [65].
Application to Human Sperm Morphology Classification
Integration with Male Infertility Diagnostics
The application of MCNN to human sperm morphology classification addresses critical challenges in male infertility diagnostics. Traditional semen analysis suffers from significant inter-observer variability, with studies reporting up to 40% disagreement between expert evaluators and kappa values as low as 0.05-0.15, highlighting substantial diagnostic inconsistency [5]. Conventional manual sperm morphology assessment is time-intensive, requiring 30-45 minutes per sample, and is influenced by technician subjectivity [5]. MCNN offers a solution to these limitations through automated, objective analysis that can significantly reduce evaluation time to less than one minute per sample while improving standardization across laboratories [5].
Implementation Considerations for Sperm Morphology Analysis
Successful implementation of MCNN for sperm morphology classification requires addressing several domain-specific challenges:
- Dataset Limitations: Current publicly available sperm image datasets (e.g., HuSHeM, SMIDS, VISEM-Tracking) often face limitations in sample size, resolution, and annotation quality [9]. The MCNN's efficiency with small datasets makes it particularly suitable for this application.
- Morphological Feature Selection: For sperm morphology analysis, morphological operations should target specific structural components including head shape (oval, tapered, pyriform, amorphous), acrosome integrity, neck structure, and tail configuration [9] [5].
- Annotation Standards: Implement consistent annotation protocols following WHO guidelines, which categorize sperm morphology into head, neck, and tail compartments with 26 types of abnormal morphology [9].
Research Reagent Solutions and Materials
Table 2: Essential Research Materials for Sperm Morphology Analysis Implementation
| Reagent/Material | Specification | Function/Application | Implementation Notes |
|---|---|---|---|
| Staining Solutions | WHO-compliant stains (e.g., Diff-Quik, Papanicolaou) [9] | Enhances morphological contrast for imaging | Critical for highlighting acrosome, nucleus, and tail structures |
| Imaging Systems | Bright-field microscopy with standardized magnification [9] | Digital image acquisition | Ensure consistent resolution and lighting conditions |
| Public Datasets | HuSHeM (216 images), SMIDS (3,000 images), VISEM-Tracking (656,334 objects) [9] | Algorithm training and validation | Addresses data scarcity through standardized benchmarks |
| Annotation Tools | Specialized software for sperm structure labeling [9] | Ground truth generation | Requires expert andrologist input for reliability |
| Computational Framework | MCNN with morphological operations [65] | Feature extraction and classification | Optimized for limited data environments |
Future Directions and Development Opportunities
The implementation of MCNN for sperm morphology classification presents several promising research directions. Future work should focus on developing more comprehensive, high-quality annotated datasets with standardized preparation, staining, and imaging protocols to enhance model generalizability [9]. Additionally, exploring domain-specific morphological operations tailored to sperm structural abnormalities could further improve diagnostic accuracy. Transfer learning approaches combining pre-trained networks with MCNN architecture may enhance performance while maintaining computational efficiency. Clinical validation studies are essential to establish diagnostic reliability and facilitate integration into routine fertility assessment protocols, ultimately advancing personalized treatment strategies in reproductive medicine [9] [24].
Measuring Success: Model Validation, Performance, and Clinical Correlation
In the implementation of Convolutional Neural Networks (CNNs) for human sperm morphology classification, the model's performance is fundamentally constrained by the quality of its training data. The "ground truth" labels used for supervised learning must accurately reflect biological reality, a challenge in a domain characterized by high subjectivity and inter-laboratory variability. Manual sperm morphology assessment remains a challenging parameter to standardize due to its subjective nature, often reliant on the operator's expertise [2]. This application note details protocols for establishing robust, defensible ground truth through expert agreement, contextualized within a broader CNN research framework for reproductive biology.
The creation of a high-quality dataset is a prerequisite for effective CNN model training. Recent studies have developed specialized datasets and reported performance metrics that highlight both the challenges and potential of deep learning in this field.
Table 1: Recent Deep Learning Datasets for Sperm Morphology Classification
| Dataset Name | Initial/ Final Image Count | Classification Standard | Reported DL Model Accuracy | Key Annotated Features |
|---|---|---|---|---|
| SMD/MSS [2] | 1,000 / 6,035 (after augmentation) | Modified David Classification (12 classes) | 55% to 92% | Head (7 defect types), midpiece (2 defect types), tail (3 defect types) |
| HSHM-CMA Dataset [43] | Confidential | Sperm Head Morphology | 60.13% - 81.42% (across 3 generalization tests) | Sperm head morphology categories |
| MHSMA [20] | 1,540 | Not Specified | Performance metrics not explicitly stated | Acrosome, head shape, vacuoles |
| SVIA Dataset [20] | 125,000 objects | Not Specified | Performance metrics not explicitly stated | 26,000 segmentation masks; objects for detection & classification |
Table 2: Inter-Expert Agreement Analysis in Sperm Morphology Classification
| Agreement Scenario | Description | Implication for Ground Truth Quality |
|---|---|---|
| No Agreement (NA) | 0 out of 3 experts agree on a label for a given sperm image. | Image is ambiguous or experts lack shared mental model; requires exclusion or senior adjudicator. |
| Partial Agreement (PA) | 2 out of 3 experts agree on the same label for at least one category. | The majority label can be used provisionally; common trigger for adjudication. |
| Total Agreement (TA) | 3 out of 3 experts agree on the same label for all categories. | Represents the highest confidence ground truth; ideal for core training set. |
Experimental Protocols for Ground Truth Establishment
Protocol 1: Three-Expert Asynchronous Annotation with Adjudication
This protocol balances rigor with operational efficiency and is widely applicable for creating CNN training sets [2] [67].
Workflow Overview:
Title: Three-Expert Asynchronous Annotation with Adjudication
Detailed Methodology:
- Image Acquisition and Preparation: Acquire images of individual spermatozoa using a system like the MMC CASA system with a 100x oil immersion objective in bright-field mode [2]. Prepare smears from semen samples stained with a RAL Diagnostics kit, following WHO guidelines. Include samples with varying morphological profiles but exclude those with very high concentrations (>200 million/mL) to avoid overlapping sperm images.
- Expert Selection and Calibration: Engage three experts, each possessing extensive experience in semen analysis. Prior to the annotation task, conduct a calibration session using a shared set of reference images (not part of the study) to align classification criteria based on the chosen standard (e.g., modified David classification).
- Blinded Independent Annotation: Provide each expert with the same set of images in a randomized order. Use an independent data logging system (e.g., a dedicated Excel spreadsheet for each expert) where they document the morphological class for each part of the spermatozoon (head, midpiece, tail) without seeing others' annotations [2].
- Automated Consensus Calculation: Compile results from all three experts. Implement a simple rule: if at least two experts assign the same label for a given sperm component, that label is provisionally accepted [67].
- Adjudication of Discordant Cases: For images where there is no 2-of-3 majority agreement (the "No Agreement" scenario from Table 2), a fourth, senior expert (the adjudicator) performs a blinded review. The adjudicator's decision on the label is considered final [67].
- Ground Truth Compilation: Create a final ground truth file containing the image name and the adjudicated morphological label, which will be used for CNN training and validation.
Protocol 2: Quantitative Analysis of Inter-Expert Agreement
This protocol is used to validate the quality of the annotation process itself and understand the inherent complexity of the classification task [2].
Workflow Overview:
Title: Inter-Expert Agreement Analysis Workflow
Detailed Methodology:
- Data Collection: Use the annotated data from Protocol 1, Step 3.
- Agreement Categorization: For each sperm image, classify the level of inter-expert agreement into one of three categories: Total Agreement (TA), Partial Agreement (PA), or No Agreement (NA), as defined in Table 2 [2].
- Statistical Analysis: Use statistical software like IBM SPSS Statistics to assess the level of agreement among the three experts. Apply Fisher's exact test to evaluate if there are statistically significant differences (p < 0.05) between the experts' classifications within each morphology class [2].
- Interpretation: A high percentage of TA indicates a reliable dataset and clear class definitions. A high percentage of PA or NA for specific morphological classes (e.g., "abnormal acrosome" or "bent midpiece") highlights areas of high subjectivity. These classes may require redefinition of criteria, additional expert training, or may be candidates for exclusion from a preliminary model to improve overall CNN accuracy.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials and Reagents for CNN-based Sperm Morphology Studies
| Item Name | Function/Application | Example Specification/Supplier |
|---|---|---|
| RAL Diagnostics Stain | Staining semen smears to visualize sperm structure for microscopy. | Standardized staining kit as used in the SMD/MSS dataset creation [2]. |
| MMC CASA System | Automated image acquisition from stained smears; provides morphometric data. | Microscope with digital camera, 100x oil immersion objective, bright-field mode [2]. |
| DF12 Culture Medium | In vitro cultivation of reference cells (e.g., for model validation). | Dulbecco’s Modified Eagle’s Medium/Ham’s F-12, supplemented with FBS [68]. |
| FITC-CD73 / PE-CD90 Antibodies | Flow cytometry validation of cell surface markers for functional correlation. | Antibodies at 10 μg/mL for flow cytometric analysis, as a functional benchmark [68]. |
| Python with Deep Learning Libraries | Implementation of CNN models for image classification and segmentation. | Python 3.8 with libraries like TensorFlow/PyTorch for algorithm development [2]. |
| Statistical Analysis Software | Quantitative analysis of inter-expert agreement and model performance. | IBM SPSS Statistics for Fisher's exact test and agreement analysis [2]. |
In the field of male fertility research, the implementation of Convolutional Neural Networks (CNN) for human sperm morphology classification has emerged as a transformative approach to address the significant limitations of manual analysis. Traditional manual sperm morphology assessment is characterized by substantial inter-observer variability, with studies reporting up to 40% disagreement between expert evaluators and kappa values as low as 0.05–0.15, highlighting profound diagnostic inconsistency [5]. This manual process is not only labor-intensive but also time-consuming, requiring 30–45 minutes per sample analysis [5]. Deep learning models offer a solution through automated, objective classification that can reduce analysis time to less than one minute per sample while providing standardized, reproducible assessments [5].
The performance of these CNN models is quantitatively evaluated through key metrics that provide distinct insights into different aspects of model effectiveness. Accuracy, precision, recall, Area Under the Curve (AUC), and Mean Absolute Error (MAE) each measure critical dimensions of model performance, from overall correctness to specific class-wise discrimination capabilities and regression accuracy. These metrics are particularly crucial in medical diagnostics like sperm morphology classification, where clinical decisions depend on reliable, interpretable model outputs. The selection and interpretation of these metrics directly impact the clinical applicability of CNN models for male fertility assessment and drug development research.
Metric Definitions and Clinical Interpretations
Core Performance Metrics Table
Table 1: Definitions and clinical interpretations of key performance metrics in sperm morphology classification.
| Metric | Mathematical Definition | Clinical Interpretation in Sperm Morphology | Optimal Range |
|---|---|---|---|
| Accuracy | (TP + TN) / (TP + TN + FP + FN) | Overall correctness in classifying normal/abnormal sperm; crucial for diagnostic reliability | >90% [5] |
| Precision | TP / (TP + FP) | Proportion of correctly identified abnormal sperm among all predicted abnormal; minimizes false alarms | >88% [20] |
| Recall (Sensitivity) | TP / (TP + FN) | Ability to identify truly abnormal sperm; critical for avoiding missed diagnoses | 88-95% [25] |
| AUC-ROC | Area under ROC curve | Overall diagnostic ability across all classification thresholds; balances sensitivity vs. specificity | >0.885 [20] |
| MAE | Σ|ypred - ytrue| / n | Average magnitude of errors in continuous measures (e.g., sperm head dimensions) | Context-dependent |
Metric Interactions and Trade-offs
In clinical practice, these metrics reveal critical trade-offs that must be balanced for optimal diagnostic performance. High precision ensures that when a model flags sperm as abnormal, it is likely correct—this is vital for avoiding unnecessary treatments and patient anxiety. High recall (sensitivity) ensures that truly abnormal sperm are not missed—critical for preventing false assurances in fertility assessments [25]. The AUC-ROC provides a comprehensive view of model performance across all possible classification thresholds, with one study reporting an AUC-ROC of 88.59% for sperm head classification [20].
The F1-score, representing the harmonic mean of precision and recall, has emerged as a particularly valuable metric in sperm morphology classification due to its ability to balance both concerns. Recent research has demonstrated exceptional F1-scores of 96.73%, 98.55%, and 99.31% for morphological classification at 20x, 40x, and 60x magnifications, respectively [28]. For acrosome health detection, even higher F1-scores of 99.8% have been achieved at 60x magnification, highlighting the potential for label-free detection of subtle morphological variations [28].
Quantitative Performance of CNN Models
Comparative Performance Across Architectures
Table 2: Reported performance metrics of deep learning models for sperm morphology classification.
| Model Architecture | Dataset | Accuracy | Precision | Recall | F1-Score | AUC-ROC |
|---|---|---|---|---|---|---|
| CBAM-enhanced ResNet50 | SMIDS (3000 images, 3-class) | 96.08% ± 1.2% [5] | - | - | - | - |
| CBAM-enhanced ResNet50 | HuSHeM (216 images, 4-class) | 96.77% ± 0.8% [5] | - | - | - | - |
| CNN (Boar Sperm) | Custom (10,000 images) | - | - | - | 96.73% (20x) [28] | - |
| CNN (Boar Sperm) | Custom (10,000 images) | - | - | - | 98.55% (40x) [28] | - |
| CNN (Boar Sperm) | Custom (10,000 images) | - | - | - | 99.31% (60x) [28] | - |
| Specialized CNN | SCIAN dataset | - | - | 88% [25] | - | - |
| Specialized CNN | HuSHeM dataset | - | - | 95% [25] | - | - |
| SVM Classifier | Custom (1,400 cells) | - | >90% [20] | - | - | 88.59% [20] |
Impact of Dataset Characteristics on Performance
The quantitative performance of CNN models is significantly influenced by dataset characteristics including size, quality, and annotation consistency. Models trained on larger datasets (e.g., 10,000 spermatozoa images) demonstrate superior performance, with F1-scores exceeding 99% at higher magnifications [28]. The implementation of advanced deep feature engineering (DFE) techniques with CBAM-enhanced ResNet50 architecture has shown statistically significant improvements of 8.08% and 10.41% on SMIDS and HuSHeM datasets respectively over baseline CNN performance [5]. These improvements highlight the importance of architectural optimization beyond basic CNN implementations for achieving clinically viable performance.
Dataset quality issues including low resolution, limited sample size, insufficient morphological categories, and inter-expert annotation variability fundamentally limit model performance [20]. Publicly available datasets such as HSMA-DS, MHSMA, VISEM-Tracking, and SVIA present varying levels of quality, with the SVIA dataset offering 125,000 annotated instances for object detection, 26,000 segmentation masks, and 125,880 cropped image objects for classification tasks [20]. Models trained on more comprehensively annotated datasets demonstrate corresponding improvements in key performance metrics, particularly precision and recall.
Experimental Protocols for Metric Evaluation
CNN Training and Validation Protocol
Protocol Title: End-to-End CNN Training for Sperm Morphology Classification Primary Focus: Multiclass classification of sperm head abnormalities Experimental Workflow:
Dataset Preparation
- Acquire annotated sperm image datasets (e.g., SMIDS, HuSHeM, SCIAN)
- Apply data augmentation: rotation, flipping, brightness adjustment, elastic transformations
- Partition data into training (70%), validation (15%), and test (15%) sets
- Implement class balancing through oversampling or weighted loss functions
Model Configuration
- Select backbone architecture (ResNet50, Xception, or custom CNN)
- Integrate attention mechanisms (CBAM) for focus on morphologically significant regions
- Configure output layer with softmax activation for multiclass classification
- Initialize with pretrained weights (ImageNet) and fine-tune on sperm data
Training Procedure
- Set batch size (16-32 based on GPU memory)
- Use Adam optimizer with learning rate 1e-4, reduced on plateau
- Employ categorical cross-entropy loss function
- Train for 100-200 epochs with early stopping patience of 15-20 epochs
Performance Validation
- Calculate accuracy, precision, recall, F1-score for each morphological class
- Compute macro-averages for overall performance assessment
- Generate confusion matrices to identify specific misclassification patterns
- Perform 5-fold cross-validation to ensure result stability [5]
Diagram 1: CNN training workflow for sperm morphology classification. Total width: 700px.
Cross-Validation and Statistical Testing Protocol
Protocol Title: Robust Model Evaluation with Statistical Validation Primary Focus: Ensuring reliable performance estimation and clinical significance Experimental Workflow:
Cross-Validation Setup
- Implement 5-fold or 10-fold cross-validation
- Ensure stratified sampling to maintain class distribution across folds
- Use consistent preprocessing and augmentation within each fold
Performance Metric Calculation
- Compute metrics per fold: Accuracy, Precision, Recall, F1, AUC-ROC
- Calculate mean and standard deviation across all folds
- Perform per-class metric analysis to identify specific weaknesses
Statistical Significance Testing
- Apply McNemar's test for comparing model performances [5]
- Use p-value threshold of 0.05 for statistical significance
- Compute confidence intervals for key performance metrics
Clinical Validation
- Compare model performance against expert embryologist assessments
- Evaluate on diverse datasets to test generalization capability
- Assess computational efficiency for clinical workflow integration
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential research reagents and materials for CNN-based sperm morphology analysis.
| Reagent/Material | Specification | Research Function | Example Application |
|---|---|---|---|
| Annotated Datasets | SMIDS (3,000 images, 3-class) [5] | Model training and validation | Benchmarking architecture performance |
| Annotated Datasets | HuSHeM (216 images, 4-class) [5] | Model training and validation | Specialized sperm head classification |
| Annotated Datasets | SCIAN (1,854 images) [25] | Model training and validation | Multiclass abnormality detection |
| Image Acquisition | ImageStreamX Mark II IBFC [28] | High-throughput sperm imaging | Acquisition at 20x, 40x, 60x magnifications |
| Staining Reagents | Hematoxylin and Eosin [28] | Sperm morphology contrast enhancement | Bright-field whole-cell staining |
| Fixation Solution | 2% Formaldehyde [28] | Sperm sample preservation | Sample preparation for IBFC |
| Software Framework | Amnis AI (AAI) Software [28] | Deep learning implementation | CNN training and deployment |
Performance Optimization Strategy
Integrated Workflow for Metric Enhancement
Diagram 2: Performance optimization workflow for metric enhancement. Total width: 700px.
Advanced Techniques for Metric Improvement
Several advanced techniques have demonstrated significant improvements in key performance metrics for sperm morphology classification. The integration of Convolutional Block Attention Module (CBAM) with ResNet50 architecture has achieved state-of-the-art performance with test accuracies of 96.08% on SMIDS dataset and 96.77% on HuSHeM dataset, representing improvements of 8.08% and 10.41% respectively over baseline CNN performance [5]. This attention mechanism enables the network to focus on the most relevant sperm features (e.g., head shape, acrosome size, tail defects) while suppressing background or noise.
Deep feature engineering (DFE) represents another powerful approach, combining the representational power of deep neural networks with classical feature selection and machine learning methods. This hybrid approach enables automatic discovery of meaningful representations while maintaining interpretability benefits crucial for medical applications [5]. The best configuration (GAP + PCA + SVM RBF) has demonstrated superior performance compared to existing state-of-the-art approaches, including recent Vision Transformer and ensemble methods [5].
The choice of magnification significantly impacts model performance, with research showing F1-scores improving from 96.73% at 20x to 99.31% at 60x magnification for morphological classification [28]. For detecting subtle acrosome health variations, 60x magnification achieved remarkable F1-scores of 99.8%, enabling label-free detection of fertility biomarkers without costly biochemical staining [28]. These findings highlight the importance of image acquisition parameters in achieving optimal performance across all key metrics.
Benchmarking CNN Performance Against Expert Embryologists and CASA Systems
The morphological evaluation of human sperm remains a cornerstone of male fertility assessment, providing critical prognostic information for natural conception and assisted reproductive technology (ART) outcomes. For decades, this analysis has relied on manual assessment by trained embryologists or automated Computer-Assisted Sperm Analysis (CASA) systems. However, both methods are hampered by significant limitations; manual analysis is inherently subjective, time-consuming, and suffers from substantial inter-observer variability, while CASA systems often demonstrate poor agreement with expert consensus, particularly for morphology classification [20] [69]. The emergence of Convolutional Neural Networks (CNNs) offers a paradigm shift, promising automated, objective, and highly accurate sperm classification. This Application Note provides a detailed protocol and benchmark data for implementing CNN-based models to classify human sperm morphology, directly comparing their performance against expert embryologists and commercial CASA systems within the context of academic and clinical research.
Performance Benchmarking
Quantitative Performance Comparison
A synthesis of recent studies reveals that deep learning models, particularly CNNs, have achieved performance metrics that meet or exceed those of expert embryologists and significantly outperform traditional CASA systems in sperm morphology classification.
Table 1: Benchmarking CNN Performance Against Experts and CASA Systems
| Assessment Method | Reported Accuracy/Performance Metrics | Key Strengths | Key Limitations |
|---|---|---|---|
| Expert Embryologists | High inter-observer variability (κ values 0.05–0.15); Analysis time: 30–45 minutes per sample [5]. | Gold standard; integrates complex morphological expertise. | Subjective; time-intensive; suffers from significant diagnostic disagreement [20] [5]. |
| CASA Systems | Morphology analysis not consistent with manual results (ICC: 0.160–0.261) [69]. | Automated; provides quantitative motility and concentration data. | Limited accuracy in morphology; poor distinction of midpiece/tail defects; results in skewed IVF/ICSI treatment allocation [2] [69]. |
| CNN-Based Models | |||
| ― CBAM-enhanced ResNet50 with Feature Engineering | 96.08% accuracy (SMIDS); 96.77% accuracy (HuSHeM) [5]. | High accuracy; attention mechanisms improve interpretability; processes samples in <1 minute [5]. | Requires computational resources and technical expertise for model development. |
| ― Stacked Ensemble of CNNs | 98.2% accuracy (HuSHeM) [70]. | Leverages multiple architectures for superior performance. | Computationally complex. |
| ― VGG16 with Transfer Learning | 94.1% true positive rate (HuSHeM) [11]. | Utilizes pre-trained networks for efficient learning. | Performance dependent on quality of fine-tuning. |
| ― Custom CNN on SMD/MSS Dataset | Accuracy ranging from 55% to 92% [2]. | Demonstrates application on a dataset built with David's classification. | Wide performance range indicates impact of dataset and model architecture. |
Clinical and Operational Impact
The transition from conventional methods to CNN-based analysis has profound implications for clinical workflow and diagnostic reliability. The most significant impact lies in the drastic reduction of analysis time, from 30–45 minutes per sample for a manual assessment to under one minute for an AI-based system, offering substantial efficiency gains [5]. Furthermore, CNNs introduce a level of standardization and objectivity that is unattainable through manual methods, effectively eliminating the high inter-observer variability (reported kappa values as low as 0.05–0.15) that plagues traditional morphology assessment [5]. This enhanced reproducibility ensures consistent results across different laboratories and technicians. From a clinical decision-making perspective, while one study found that CASA systems could lead to skewed IVF/ICSI treatment allocation due to inaccurate morphology readings [69], the high-accuracy classification provided by robust CNN models promises to deliver more reliable morphological data, thereby supporting more appropriate and effective treatment planning.
Experimental Protocols
Protocol 1: CNN Model Training for Sperm Morphology Classification
This protocol describes the procedure for developing a CNN-based sperm classifier, from dataset preparation to model evaluation, based on established methodologies [2] [5] [11].
Research Reagent Solutions:
- Public Datasets: HuSHeM (216 images, 4-class), SCIAN-MorphoSpermGS, SMIDS (3000 images, 3-class), SVIA (125,000+ annotations) [20] [5] [11].
- Staining Reagents: RAL Diagnostics staining kit for preparing semen smears [2].
- Software & Libraries: Python 3.8+, TensorFlow/Keras or PyTorch, Scikit-learn, OpenCV, ITK-SNAP (for segmentation) [2] [71] [72].
Procedure:
- Sample Preparation & Image Acquisition:
- Prepare semen smears from samples with a concentration of at least 5 million/mL according to WHO guidelines and stain using a standardized kit (e.g., RAL Diagnostics) [2].
- Acquire images of individual spermatozoa using a microscope with a 100x oil immersion objective, ideally coupled with a CASA system's camera for sequential capture [2].
- Expert Annotation & Ground Truth Establishment:
- Have a minimum of three experienced embryologists classify each sperm image independently based on a standardized classification system (e.g., WHO strict criteria or modified David classification) [2].
- Resolve disagreements through consensus or a senior expert. Compile a ground truth file detailing the image name, expert classifications, and agreed label for each spermatozoon [2].
- Data Preprocessing & Augmentation:
- Clean images by handling missing values and outliers. Resize all images to a uniform dimension (e.g., 80x80 pixels) and normalize pixel values [2].
- Apply data augmentation techniques to balance classes and increase dataset size. Common operations include rotation, horizontal/vertical flipping, and zooming [2] [72]. The SMD/MSS dataset, for instance, was expanded from 1,000 to 6,035 images via augmentation [2].
- Model Architecture & Training:
- Option A (Transfer Learning): Fine-tune a pre-trained CNN (e.g., VGG16, ResNet50) [11]. Replace the final classification layer with a new one matching the number of sperm morphology classes. Unfreeze and fine-tune the final few layers of the base model.
- Option B (Custom Model): Design a custom CNN architecture with convolutional, pooling, and fully connected layers. Incorporating attention modules like CBAM can enhance performance by helping the network focus on salient sperm features [5].
- Partition the dataset into training (80%), validation (10%), and test (10%) sets. Train the model using an optimizer (e.g., Adam) and a loss function (e.g., categorical cross-entropy) appropriate for multi-class classification.
- Model Evaluation:
- Evaluate the final model on the held-out test set. Report standard metrics including accuracy, precision, recall, F1-score, and area under the ROC curve (AUC-ROC) [5].
- Use techniques like Grad-CAM or attention visualization to generate heatmaps that highlight the image regions most influential in the model's decision, thereby providing interpretability [73] [5].
- Sample Preparation & Image Acquisition:
Protocol 2: Benchmarking Against Experts and CASA
This protocol outlines the experimental design for a head-to-head comparison of a trained CNN model against human experts and CASA systems.
- Procedure:
- Selection of Benchmark Cohort:
- Execution of Comparative Analysis:
- CNN Model: Run the pre-trained CNN model on the test set to generate morphology classifications.
- Expert Embryologists: Have a panel of embryologists (blinded to the ground truth and each other's results) classify the same test set images.
- CASA Systems: Process the same samples or images using one or more commercial CASA systems (e.g., Hamilton-Thorne CEROS II, LensHooke X1 Pro) according to manufacturer instructions to obtain automated morphology classifications [69].
- Statistical Analysis:
- Calculate the agreement between each method (CNN, each expert, CASA) and the consensus ground truth using metrics like accuracy and Cohen's Kappa (κ).
- Assess inter-observer variability between experts and the CNN using κ statistics [5].
- For CASA vs. manual/ground truth, compute the Intraclass Correlation Coefficient (ICC) for continuous data or κ for categorical data, as applicable [69]. Bland-Altman plots can be used to visualize the agreement between CASA and manual results for parameters like concentration [69].
Workflow and Conceptual Diagrams
Experimental Workflow for Benchmarking CNN Performance
The following diagram illustrates the end-to-end experimental workflow for developing a CNN model and benchmarking its performance against traditional methods.
Diagram Title: Workflow for CNN Benchmarking in Sperm Analysis
Performance Comparison Logic
This diagram conceptualizes the performance hierarchy identified in the benchmark data, illustrating the relative positioning of CNN models, expert embryologists, and CASA systems.
Diagram Title: Performance Hierarchy of Assessment Methods
The Scientist's Toolkit
Table 2: Essential Research Reagents and Computational Resources
| Item Name | Function/Application | Specifications/Examples |
|---|---|---|
| HuSHeM Dataset | Public benchmark dataset for training and validating sperm head classification models. | 216 images, 4-class (Normal, Tapered, Pyriform, Small) [11]. |
| SMIDS Dataset | Public dataset for multi-class sperm morphology classification. | 3000 images, 3-class structure [5]. |
| SVIA Dataset | Large-scale public dataset for object detection, segmentation, and classification. | 125,000+ annotated instances; 26,000 segmentation masks [20]. |
| RAL Diagnostics Stain | Staining kit for preparing semen smears for morphological evaluation. | Used for creating standardized smears for image acquisition [2]. |
| Pre-trained CNN Models | Base models for transfer learning, significantly reducing development time and computational cost. | VGG16, ResNet50, DenseNet121, Vision Transformer [71] [11]. |
| Attention Modules (CBAM) | Enhances CNN feature extraction by focusing the model on relevant morphological structures (head, midpiece). | Convolutional Block Attention Module [5]. |
| TensorFlow / PyTorch | Primary software frameworks for building, training, and evaluating deep learning models. | Open-source libraries with extensive community support [72]. |
Comparative Analysis of State-of-the-Art Deep Learning Models
Deep learning has revolutionized the analysis of complex biomedical data, offering powerful tools for automating and standardizing tasks that were previously reliant on subjective human assessment. Within the specific domain of human sperm morphology classification—a critical procedure in male infertility diagnosis—deep learning models demonstrate particular promise for overcoming challenges such as inter-expert variability and the labor-intensive nature of manual analysis [20]. This review provides a comparative analysis of state-of-the-art deep learning architectures, evaluating their applicability, performance, and implementation protocols for sperm morphology classification. By framing this analysis within the context of a broader thesis on implementing convolutional neural networks (CNNs) for this purpose, we aim to provide researchers and drug development professionals with a practical roadmap for selecting, adapting, and deploying these advanced computational techniques in reproductive medicine.
State-of-the-Art Deep Learning Architectures
Foundational Architectures for Image Classification
The evolution of deep learning has produced several foundational architectures that form the basis for many specialized applications in medical image analysis. While newer architectures continue to emerge, several have established strong performance records across diverse domains.
Convolutional Neural Networks (CNNs) represent a fundamental architecture where multiple layers process input data to automatically learn hierarchical feature representations [74]. The VGG architecture, particularly VGG16, has demonstrated exceptional transfer learning capabilities for sperm head classification, achieving true positive rates of 94.1% on benchmark datasets when pre-trained on ImageNet and fine-tuned with sperm images [11]. The ResNet (Residual Network) architecture, specifically ResNet-50, introduces skip connections that enable the training of much deeper networks by mitigating the vanishing gradient problem, making it particularly valuable for complex visual pattern recognition in sperm motility assessment [16].
More recent advancements include EfficientNet, which utilizes a compound scaling method to systematically balance network depth, width, and resolution, achieving state-of-the-art efficiency and accuracy trade-offs [75]. ConvNeXt V2 modernizes the classic CNN design by incorporating ideas from Vision Transformers while maintaining the computational efficiency of pure convolutional architectures [75].
Transformer-Based and Hybrid Architectures
The success of transformer architectures in natural language processing has inspired their adaptation to computer vision tasks, leading to the development of several powerful models.
Vision Transformers (ViTs) process images by dividing them into patches and treating them as sequences, leveraging self-attention mechanisms to capture global contextual information [75]. DaViT (Dual Attention Vision Transformer) enhances this approach by incorporating two complementary self-attention mechanisms: spatial attention that processes tokens along the spatial dimension, and channel attention that processes tokens along the channel dimension [75]. This dual approach enables more comprehensive feature learning, with DaViT models achieving up to 90.4% top-1 accuracy on ImageNet-1K when fine-tuned [75].
CoCa (Contrastive Captioners) represents a multimodal architecture that combines contrastive learning and generative captioning in a unified framework [75]. While originally designed for vision-language tasks, its dual-purpose encoder-decoder architecture shows promise for medical image analysis tasks that require both classification and explanatory output.
Table 1: Performance Comparison of State-of-the-Art Deep Learning Models on Image Classification Benchmarks
| Model | Architecture Type | Parameters | ImageNet Top-1 Accuracy (Fine-tuned) | Key Strengths |
|---|---|---|---|---|
| CoCa | Multimodal Transformer | 2.1B | 91.0% | Excellent zero-shot capabilities, combines contrastive and generative learning |
| DaViT-Giant | Dual Attention Transformer | 1.4B | 90.4% | Complementary spatial and channel attention mechanisms |
| ConvNeXt V2 | Modernized CNN | Varies by size | ~88.7% | Pure convolutional efficiency with modern design |
| EfficientNet | Scaled CNN | Varies by size | ~88.9% | Optimal balance of depth, width, and resolution |
| VGG16 | Classical CNN | 138M | 94.1%* | Proven transfer learning capability, extensive community support |
| ResNet-50 | Residual CNN | 25.6M | High* | Skip connections enable deep networks, strong feature extraction |
Note: VGG16 and ResNet-50 ImageNet accuracy not explicitly stated in search results; VGG16 achieved 94.1% true positive rate on HuSHeM sperm dataset [11]; ResNet-50 demonstrated strong performance on sperm motility assessment [16].
Application to Human Sperm Morphology Classification
Performance Metrics and Comparative Analysis
The application of deep learning to sperm morphology classification has yielded impressive results, with several studies demonstrating performance comparable to or exceeding human expert assessment.
Table 2: Deep Learning Performance in Sperm Morphology and Motility Analysis
| Study | Model Architecture | Dataset | Key Performance Metrics | Classification Categories |
|---|---|---|---|---|
| Mohd Noor et al., 2025 [2] | Custom CNN | SMD/MSS (6,035 images) | Accuracy: 55-92% | 12 morphological classes (David classification) |
| Riordon et al., 2019 [11] | VGG16 (Transfer Learning) | HuSHeM & SCIAN | True Positive Rate: 94.1% (HuSHeM), 62% (SCIAN) | 5 WHO categories: Normal, Tapered, Pyriform, Small, Amorphous |
| Leanderson et al., 2023 [16] | ResNet-50 | ESHRE-SIGA EQA (65 videos) | MAE: 0.05 (3-category), 0.07 (4-category); Pearson's r=0.88 (progressive motility) | Motility categories: Progressive, Non-progressive, Immotile |
| BMC Urology Review, 2025 [20] | Multiple DL Models | SVIA (125,000 instances) | Various: High accuracy for segmentation and classification | Head, neck, and tail abnormalities |
The variation in reported accuracy (55-92% [2] versus 94.1% true positive rate [11]) highlights the significant impact of dataset quality, annotation consistency, and the specific classification schema employed. Models trained on datasets with expert consensus labels typically achieve higher performance metrics, underscoring the critical importance of ground truth quality in model development.
Technical Implementation Considerations
Successful implementation of deep learning for sperm morphology classification requires careful consideration of several technical factors. Transfer learning has emerged as a particularly effective strategy, where models pre-trained on large-scale natural image datasets (e.g., ImageNet) are fine-tuned on specialized sperm image datasets [11]. This approach leverages generalized feature extraction capabilities while adapting to domain-specific characteristics, significantly reducing data requirements and training time.
Data augmentation represents another critical component, with techniques such as rotation, flipping, color adjustment, and scaling employed to artificially expand dataset size and diversity [2]. In one study, the initial dataset of 1,000 images was expanded to 6,035 images through augmentation, substantially improving model robustness and generalization [2].
The attention mechanism, a cornerstone of transformer architectures, enables models to focus on the most relevant image regions for making classification decisions [75]. In sperm morphology analysis, this could translate to prioritized processing of head morphology features over background elements, potentially mirroring the analytical approach of human experts.
Experimental Protocols
Dataset Preparation and Augmentation Protocol
Objective: To create a robust, balanced dataset for training deep learning models in sperm morphology classification.
Materials:
- Microscope with digital camera (100x oil immersion objective recommended)
- Stained semen smears (RAL Diagnostics staining kit or equivalent)
- Computer with adequate storage capacity
- Data augmentation software (e.g., Python with OpenCV, Albumentations, or TensorFlow Data Augmentation modules)
Procedure:
- Sample Preparation and Image Acquisition:
- Prepare semen smears according to WHO standard procedures [2] [20]
- Capture images of individual spermatozoa using MMC CASA system or equivalent [2]
- Ensure each image contains a single spermatozoon with clear visualization of head, midpiece, and tail
- Save images in high-resolution format (e.g., PNG or TIFF to minimize compression artifacts)
Expert Annotation and Ground Truth Establishment:
- Engage multiple experienced andrologists for independent classification [2]
- Utilize standardized classification systems (WHO criteria or David's modified classification) [2] [20]
- Resolve discrepant classifications through consensus meetings or majority voting
- Compile ground truth file linking image names with classification labels and expert agreement levels [2]
Data Preprocessing:
- Resize images to consistent dimensions (e.g., 80×80 pixels for sperm head classification) [2]
- Convert to grayscale if color information is not diagnostically relevant
- Apply normalization to scale pixel values to standard range (e.g., 0-1)
- Implement noise reduction algorithms to minimize staining and optical artifacts [2]
Data Augmentation:
- Apply geometric transformations: rotation (±15°), horizontal/vertical flipping, slight scaling (±10%)
- Implement photometric adjustments: brightness (±20%), contrast (±15%), gamma correction
- Use advanced techniques: elastic deformations, grid distortion, random erasing
- Ensure augmentation preserves diagnostic features and morphological characteristics
Dataset Partitioning:
Dataset Preparation Workflow: Protocol for creating training data for sperm morphology classification models.
Model Training and Transfer Learning Protocol
Objective: To implement and fine-tune state-of-the-art deep learning models for sperm morphology classification using transfer learning.
Materials:
- GPU-enabled computational environment (NVIDIA GPUs with CUDA support recommended)
- Deep learning frameworks (TensorFlow, PyTorch, or Keras)
- Pre-trained model weights (ImageNet pre-trained models typically used)
- Prepared sperm morphology dataset (from Protocol 4.1)
Procedure:
- Model Selection and Initialization:
- Select appropriate base architecture (VGG16, ResNet-50, or EfficientNet for balance of performance and efficiency) [11] [16]
- Load pre-trained weights (ImageNet initialization typically used)
- Remove original classification head (final fully connected layers)
- Initialize new classification head with number of classes specific to sperm morphology task
Progressive Fine-tuning:
- Phase 1: Freeze all base model layers, train only the new classification head
- Use moderate learning rate (0.001-0.01)
- Train for sufficient epochs until validation performance plateaus
- Phase 2: Unfreeze all layers for end-to-end fine-tuning
- Use lower learning rate (0.0001-0.001) to avoid overwriting useful pre-trained features
- Employ differential learning rates if supported (lower rates for earlier layers)
- Monitor closely for overfitting using validation set performance
- Phase 1: Freeze all base model layers, train only the new classification head
Training Configuration:
- Loss function: Categorical cross-entropy for multi-class classification
- Optimizer: Adam (β₁=0.9, β₂=0.999) with gradient clipping if necessary [16]
- Batch size: Maximize within GPU memory constraints (typically 16-64)
- Early stopping: Monitor validation loss with patience of 10-20 epochs
- Learning rate scheduling: Reduce on plateau when validation performance stagnates
Validation and Evaluation:
- Evaluate on held-out test set only after model selection and training complete
- Report comprehensive metrics: accuracy, precision, recall, F1-score, confusion matrix
- For regression tasks (motility assessment): MAE, Pearson correlation coefficients [16]
- Compare against baseline methods and inter-expert variability
Transfer Learning Protocol: Two-phase approach for adapting pre-trained models to sperm classification.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Essential Research Reagents and Computational Tools for Deep Learning in Sperm Morphology Analysis
| Category | Item/Reagent | Specification/Function | Application in Research |
|---|---|---|---|
| Wet Lab Supplies | RAL Diagnostics Staining Kit | Provides differential staining of sperm components | Enhances contrast for morphological features in microscopic imaging [2] |
| Microscope Slides and Coverslips | Standardized dimensions and thickness | Ensures consistent imaging conditions and prevents specimen deformation | |
| Immersion Oil | High-quality, non-drying formula | Maintains optical clarity for high-resolution (100x) microscopy [2] | |
| Imaging Equipment | CASA System with Digital Camera | Integrated computer-assisted semen analysis | Standardized image acquisition with calibrated magnification [2] [16] |
| Phase-Contrast Microscope | Enhanced contrast for unstained specimens | Optional for motility analysis without staining artifacts [16] | |
| Temperature-Controlled Stage | Maintains 37°C during imaging | Preserves sperm viability and motility characteristics during assessment [16] | |
| Computational Resources | GPU Workstations | NVIDIA RTX series or equivalent with CUDA support | Accelerates model training and inference through parallel processing [11] |
| Deep Learning Frameworks | TensorFlow, PyTorch, or Keras | Provides pre-built components for model development and training [11] [16] | |
| Data Augmentation Libraries | Albumentations, Imgaug, or TensorFlow Addons | Expands effective dataset size and diversity through transformations [2] | |
| Reference Materials | WHO Laboratory Manual | Standardized procedures for semen examination | Establishes consistent classification criteria and protocols [20] [11] |
| Public Benchmark Datasets | HuSHeM, SCIAN, SVIA, or SMD/MSS | Provides baseline comparisons and additional training data [2] [20] [11] |
The comparative analysis presented herein demonstrates that state-of-the-art deep learning models offer viable, high-performance solutions for automating human sperm morphology classification. Architectures such as VGG16 and ResNet-50 have already demonstrated compelling performance in research settings, with true positive rates exceeding 94% in some studies [11]. Meanwhile, emerging transformer-based models like DaViT and CoCa present promising avenues for future research, potentially capturing more complex morphological patterns through advanced attention mechanisms [75].
The successful implementation of these technologies requires meticulous attention to dataset quality, appropriate application of transfer learning methodologies, and comprehensive performance validation against established clinical standards. As these computational approaches continue to mature, they hold significant potential for standardizing sperm morphology assessment across clinical laboratories, reducing inter-expert variability, and ultimately improving diagnostic accuracy in male infertility evaluation. Future research directions should focus on integrating multimodal assessment (combining morphology, motility, and clinical parameters), developing explainable AI approaches to enhance clinical trust, and creating larger, more diverse datasets to improve model generalization across heterogeneous patient populations.
Correlating AI Classification with Clinical Outcomes like Time to Pregnancy
The assessment of sperm morphology is a cornerstone of male fertility evaluation, providing critical insights into reproductive potential. Traditionally, this analysis has been a manual process, subject to significant inter-observer variability and reproducibility challenges [9]. The integration of Convolutional Neural Networks (CNNs) and deep learning represents a paradigm shift, offering a path toward automated, objective, and highly accurate sperm morphology classification [30] [76]. However, the ultimate validation of any diagnostic tool lies in its ability to predict clinically relevant endpoints. This document outlines detailed application notes and protocols for developing and validating CNN-based sperm morphology classifiers, with a specific focus on the critical step of correlating algorithmic outputs with clinical outcomes, most importantly, time to pregnancy.
Background and Literature Synthesis
Male factors contribute to approximately 50% of infertility cases globally, with sperm morphological quality showing a declining trend [9]. The World Health Organization (WHO) mandates the classification of over 200 sperm cells into normal or various abnormal categories (e.g., head, neck/midpiece, tail defects), a process that is both time-consuming and subjective [9] [10]. Deep learning models, particularly CNNs, have demonstrated superior performance in automating this task. Studies have successfully employed models like DenseNet169 for classification, Mask R-CNN and U-Net for precise segmentation of sperm components (head, acrosome, nucleus, tail), and YOLOv7/v8 for real-time detection and classification [10] [30] [76]. These models learn hierarchical feature representations directly from image data, overcoming the limitations of manual feature extraction in conventional machine learning [9].
A significant challenge in the field is the reproducibility crisis in machine learning research. Inconsistent reporting of data splits, hyperparameters, and evaluation metrics hinders the independent validation of models [77] [78] [79]. Furthermore, the "black box" nature of complex models can impede clinical adoption, as understanding how a model arrives at a decision is often as important as the decision itself [79]. Therefore, a rigorous framework that prioritizes reproducibility, interpretability, and, ultimately, clinical correlation is essential for translating algorithmic advances into reliable diagnostic tools.
Table 1: Key Deep Learning Models for Sperm Morphology Analysis
| Model Name | Primary Application | Reported Performance | Key Advantage |
|---|---|---|---|
| DenseNet169 [76] | Sperm head morphology classification | High accuracy (specific metrics not provided in source) | Mitigates vanishing gradient problem, improves feature efficiency. |
| Mask R-CNN [30] | Multi-part segmentation (head, acrosome, nucleus, tail) | High IoU for smaller, regular structures (head, nucleus) | Robust instance segmentation; excels at delineating fine structures. |
| U-Net [30] | Segmentation, particularly of morphologically complex parts | Highest IoU for tail segmentation | Superior global perception and multi-scale feature extraction. |
| YOLOv8 [30] | Segmentation and real-time detection | Comparable to Mask R-CNN for neck segmentation | Single-stage model offering a balance of speed and accuracy. |
| YOLOv7 [10] | Object detection and classification of sperm abnormalities | mAP@50: 0.73, Precision: 0.75, Recall: 0.71 | Balanced tradeoff between accuracy and computational efficiency. |
Experimental Protocols
Dataset Curation and Preprocessing Protocol
The foundation of a robust model is a high-quality, well-annotated dataset.
- Data Acquisition: Capture sperm images using standardized microscopy protocols. For clinical correlation, samples must be linked to de-identified patient records containing outcome data (e.g., time to pregnancy, success of IVF/ICSI). Ethical approval and informed consent for data use are mandatory [10].
- Data Annotation: Annotate images according to WHO guidelines [9]. Labels should include:
- Data Preprocessing: Implement a consistent pipeline:
- Resizing: Standardize image dimensions to the input size of the target CNN.
- Normalization: Scale pixel intensities to a [0, 1] or [-1, 1] range.
- Augmentation: Apply random transformations (rotation, flipping, brightness/contrast adjustments) to increase dataset diversity and improve model generalization [9].
Table 2: Publicly Available Sperm Image Datasets for Model Development
| Dataset Name | Key Characteristics | Content | Annotations |
|---|---|---|---|
| SVIA [9] [30] | Large-scale, low-resolution, unstained sperm | 125,000 annotated instances; 26,000 segmentation masks | Detection, Segmentation, Classification |
| VISEM-Tracking [9] | Multi-modal, includes videos | 656,334 annotated objects with tracking details | Detection, Tracking, Regression |
| MHSMA [9] | Non-stained, grayscale sperm head images | 1,540 sperm head images | Classification |
| HuSHeM [9] | Stained sperm head images, higher resolution | 725 sperm head images (216 publicly available) | Classification |
| SCIAN-MorphoSpermGS [9] | Stained sperm images | 1,854 sperm images across five classes | Classification |
CNN Model Development and Training Protocol
This protocol details the process of building and training a morphology classifier.
- Model Selection: Choose a model architecture based on the task (e.g., Mask R-CNN or U-Net for segmentation; DenseNet or YOLO for classification) [30] [76].
- Transfer Learning: Initialize the model with weights pre-trained on a large dataset like ImageNet. Fine-tune the final layers, and potentially intermediate layers, on the curated sperm morphology dataset [76].
- Hyperparameter Tuning: Systematically optimize hyperparameters, which significantly impact model accuracy and reproducibility [77]. Use a validation set for this purpose.
- Model Training: Split data into training (70-80%), validation (10-15%), and a held-out test set (10-15%). Train the model using the training set and monitor performance on the validation set to prevent overfitting.
Protocol for Correlating AI Output with Time to Pregnancy
This is the core protocol for establishing clinical validity.
- Cohort Definition: Assemble a retrospective cohort of patients for whom sperm morphology images and confirmed time-to-pregnancy (TTP) data are available. TTP is defined as the number of menstrual cycles required to achieve a pregnancy.
- AI-Based Phenotyping: Process the sperm images from this cohort through the trained CNN model. For each patient, extract one or more of the following quantitative biomarkers:
Morphology Score: The percentage of sperm classified as "normal" by the AI.Defect-Specific Scores: The prevalence of specific abnormalities (e.g., % head defects, % tail defects).Advanced Morphological Descriptors: Continuous measures derived from segmentation masks (e.g., head ellipticity, acrosome area ratio, tail length).
- Statistical Analysis:
- Spearman Correlation: Calculate the non-parametric correlation between the continuous AI-derived morphology score and TTP.
- Kaplan-Meier Analysis: Categorize patients into groups based on AI morphology score percentiles (e.g., low/normal/high abnormality). Plot TTP curves for each group and compare them using the Log-rank test.
- Cox Proportional-Hazards Regression: Perform multivariate analysis to determine if the AI morphology score is an independent predictor of TTP, after adjusting for confounders such as female partner's age, hormone levels, and lifestyle factors. Report the Hazard Ratio (HR) with confidence intervals.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials and Tools for AI-Based Sperm Morphology Research
| Item Name | Function/Application | Specification/Example |
|---|---|---|
| Phase-Contrast Microscope | High-quality image acquisition of live, unstained sperm. | Model: Optika B-383Phi [10]. |
| Sperm Fixation System | Immobilizes sperm for morphology analysis without dye. | System: Trumorph [10]. |
| Annotation Software | For precise labeling of sperm images for model training. | Software: Roboflow [10]. |
| Deep Learning Framework | Platform for building and training CNN models. | Frameworks: TensorFlow, PyTorch. |
| Pre-trained CNN Models | Base architectures for transfer learning. | Models: DenseNet169, ResNet-50, YOLOv8, Mask R-CNN [30] [76]. |
| Statistical Software | For performing correlation and survival analysis. | Software: R, Python (with lifelines, scikit-survival packages). |
Ensuring Reproducible and Interpretable Research
To combat the reproducibility crisis and build trust in AI models, adhere to the following guidelines:
- Documentation: Follow the CRISP-DM framework or checklists like MI-CLAIM to ensure all aspects of the research are reported [80] [81] [79]. This includes detailed descriptions of data provenance, model architectures, hyperparameters, and evaluation metrics.
- Code and Data Sharing: Where possible, share analysis code and, if ethically permissible, de-identified data or synthetic datasets to allow for independent verification (R1 reproducibility) [78] [79].
- Interpretability: Use techniques like Gradient-weighted Class Activation Mapping (Grad-CAM) to generate heatmaps showing which image regions the model focused on for its classification. This helps validate the model's decision against biological knowledge and builds clinical confidence [79].
The integration of CNN-based sperm morphology classification with clinical outcome data represents the frontier of male fertility assessment. By implementing the detailed protocols for dataset curation, model development, and clinical correlation outlined in this document, researchers can generate robust, reproducible, and clinically meaningful evidence. This approach moves beyond simple automation of a manual task and toward the development of powerful, AI-driven prognostic biomarkers that can ultimately improve patient counseling and treatment strategies for infertility.
Conclusion
The implementation of CNNs for sperm morphology classification represents a paradigm shift in andrology, offering a path toward unprecedented objectivity, standardization, and efficiency in male fertility assessment. The synthesis of research demonstrates that while challenges such as data scarcity and model generalizability persist, advanced approaches like data augmentation, transfer learning, and hybrid architectures yield models with accuracy rivaling or surpassing human experts. These AI tools not only automate a critical diagnostic step but also unlock deeper morphological insights through feature visualization. Future directions must focus on the development of larger, more diverse multi-center datasets, the clinical integration of these models into real-time assisted reproductive technologies like ICSI, and rigorous prospective trials to validate their impact on ultimate endpoints—successful pregnancy and live birth rates. For researchers and drug developers, this technology opens new avenues for high-throughput phenotypic screening and the objective assessment of therapeutic interventions on sperm quality.
References
- 1. Male Infertility - StatPearls - NCBI Bookshelf [https://www.ncbi.nlm.nih.gov/books/NBK562258/]
- 2. Deep-learning based model for sperm morphology ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12599357/]
- 3. Convolutional neural network [https://en.wikipedia.org/wiki/Convolutional_neural_network]
- 4. Deep Learning: Basics and Convolutional Neural Networks ... [https://www.ncbi.nlm.nih.gov/books/NBK597497/]
- 5. Deep feature engineering for accurate sperm morphology ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0330914]
- 6. The validity and reliability of computer-aided semen ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8350227/]
- 7. How to optimize the benefits of computer assisted sperm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3599994/]
- 8. Standardization of computer-assisted semen analysis ... [https://www.sciencedirect.com/science/article/abs/pii/S0093691X11001038]
- 9. Deep learning-based approach for sperm morphology analysis [https://pmc.ncbi.nlm.nih.gov/articles/PMC12538767/]
- 10. Design and Implementation of a Deep Learning System to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12567593/]
- 11. Deep learning for the classification of human sperm [https://www.sciencedirect.com/science/article/abs/pii/S0010482519302112]
- 12. Classification of Human Sperm Based on Morphology ... [https://www.atlantis-press.com/proceedings/icasmi-24/126011712]
- 13. An Expert Review From the French BLEFCO Group [https://pubmed.ncbi.nlm.nih.gov/41185394/]
- 14. Multi-model CNN fusion for sperm morphology analysis [https://www.sciencedirect.com/science/article/abs/pii/S0010482521005849]
- 15. Attention-enhanced deep learning for cervical cytology [https://pmc.ncbi.nlm.nih.gov/articles/PMC12550697/]
- 16. Sperm motility assessed by deep convolutional neural ... [https://www.nature.com/articles/s41598-023-41871-2]
- 17. Deep-learning based model for sperm morphology ... [https://pubmed.ncbi.nlm.nih.gov/41203404/]
- 18. Sperm Abnormality Detection Using Sequential Deep ... [https://www.mdpi.com/2227-7390/11/3/515?type=check_update&version=1]
- 19. VISEM-Tracking, a human spermatozoa tracking dataset [https://www.nature.com/articles/s41597-023-02173-4]
- 20. Deep learning-based approach for sperm morphology analysis [https://bmcurol.biomedcentral.com/articles/10.1186/s12894-025-01946-w]
- 21. Advancements in automated sperm morphology analysis [https://www.semanticscholar.org/paper/Advancements-in-automated-sperm-morphology-a-deep-Maalej-Abdelkefi/0c0c1d91a97509d1a998c3c14318a84b85b6bb80]
- 22. 3D+t Multifocal Imaging Dataset of Human Sperm [https://www.nature.com/articles/s41597-025-05177-4]
- 23. AndroGen: Open-source synthetic data generation for ... [https://www.sciencedirect.com/science/article/pii/S0169260725005486]
- 24. Revolutionizing Sperm Analysis with AI: A Review of ... [https://www.mdpi.com/2079-3197/13/6/132]
- 25. Deep Learning-Based Morphological Classification of ... [https://www.mdpi.com/2075-4418/10/5/325]
- 26. Sperm Abnormality Detection Using Sequential Deep ... [https://www.mdpi.com/2227-7390/11/3/515]
- 27. Deep learning-based sperm motility and morphology ... [https://www.sciencedirect.com/science/article/pii/S2352914824000157]
- 28. Deep learning classification method for boar sperm ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12368927/]
- 29. Human live spermatozoa morphology assessment using ... [https://www.nature.com/articles/s41598-022-08798-6]
- 30. Deep Learning Models for Multi-Part Morphological Segmentation and Evaluation of Live Unstained Human Sperm [https://www.mdpi.com/1424-8220/25/10/3093]
- 31. Democratising deep learning for microscopy with ... [https://www.nature.com/articles/s41467-021-22518-0]
- 32. Advanced Multi-Level Ensemble Learning Approaches for Comprehensive Sperm Morphology Assessment - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12192502/]
- 33. Deep learning for the classification of human sperm - PubMed [https://pubmed.ncbi.nlm.nih.gov/31279166/]
- 34. AI-Powered Microscopy Image Analysis: A Beginner's Guide [https://mediacy.com/blog/ai-powered-microscopy/]
- 35. Design and Implementation of a Deep Learning System to ... [https://www.mdpi.com/2306-7381/12/10/1015]
- 36. Enhancing the precision of male fertility diagnostics ... [https://www.nature.com/articles/s41598-025-25052-x]
- 37. Deep feature engineering for accurate sperm morphology ... [https://pubmed.ncbi.nlm.nih.gov/40929291/]
- 38. Automatic Microscopy Analysis with Transfer Learning for ... [https://www.mdpi.com/2076-3417/11/12/5369]
- 39. Transfer Learning in Medical Image Segmentation [https://pmc.ncbi.nlm.nih.gov/articles/PMC8164174/]
- 40. Optimizing CNN performance in medical imaging with ... [https://www.sciencedirect.com/science/article/pii/S2666521225000857]
- 41. Leveraging transfer learning-driven convolutional neural ... [https://www.nature.com/articles/s41598-024-81966-y]
- 42. How to improve the performance of CNN Model for a ... [https://stackoverflow.com/questions/70554413/how-to-improve-the-performance-of-cnn-model-for-a-specific-dataset-getting-low]
- 43. A generalized classification of human sperm head ... [https://www.sciencedirect.com/science/article/pii/S0925231225011269]
- 44. Interpreting convolutional neural network explainability for ... [https://www.sciencedirect.com/science/article/pii/S0010482525012776]
- 45. Explainable CNN–Radiomics Fusion and Ensemble ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12385016/]
- 46. Deep clustering analysis via variational autoencoder with ... [https://www.sciencedirect.com/science/article/pii/S0893608024009080]
- 47. Comparing factor mixture modeling and conditional ... [https://www.frontiersin.org/journals/psychology/articles/10.3389/fpsyg.2025.1474292/full]
- 48. Deep Convolutional Neural Networks in Medical Image ... [https://www.mdpi.com/2078-2489/16/3/195]
- 49. A review and systematic guide to counteracting medical ... [https://www.sciencedirect.com/science/article/pii/S266699002500045X]
- 50. A deep convolutional neural network-based novel class ... [https://www.nature.com/articles/s41598-025-04952-y]
- 51. Overfitting in Machine Learning Explained [https://aws.amazon.com/what-is/overfitting/]
- 52. What is Overfitting in Deep Learning [+10 Ways to Avoid It] [https://www.v7labs.com/blog/overfitting]
- 53. How to solve the overfitting problem in convolutional neural ... [https://www.tencentcloud.com/techpedia/106000]
- 54. a method for preventing overfitting in convolutional neural ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6561649/]
- 55. Mastering Dropout: The Ultimate Strategy to Prevent ... [https://www.lunartech.ai/blog/mastering-dropout-the-ultimate-strategy-to-prevent-overfitting-in-neural-networks]
- 56. deep learning - How to improve loss and avoid overfitting [https://datascience.stackexchange.com/questions/30097/how-to-improve-loss-and-avoid-overfitting]
- 57. Prevention of overfitting in convolutional layers of a CNN [https://stackoverflow.com/questions/36139980/prevention-of-overfitting-in-convolutional-layers-of-a-cnn]
- 58. Convolutional Neural Network (CNN) | TensorFlow Core [https://www.tensorflow.org/tutorials/images/cnn]
- 59. Deep Learning–based Detection of Intravenous Contrast ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9152686/]
- 60. Artificial intelligence in reproductive medicine - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6733338/]
- 61. Artificial Intelligence in Reproductive Medicine: Transforming ... [https://jivfww.scholasticahq.com/article/137620-artificial-intelligence-in-reproductive-medicine-transforming-assisted-reproductive-technologies]
- 62. Category-Aware Two-Stage Divide-and-Ensemble ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12428259/]
- 63. High spatially sensitive quantitative phase imaging ... [https://www.nature.com/articles/s41598-020-69857-4]
- 64. Deep Learning with Contrast-Enhanced Ultrasound for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12553386/]
- 65. Hybrid morphological-convolutional neural networks for ... [https://www.frontiersin.org/journals/artificial-intelligence/articles/10.3389/frai.2023.1253183/full]
- 66. Hybrid morphological-convolutional neural networks for ... [https://doaj.org/article/0b28415523b94f738b1a37cb0bbd4a10]
- 67. Definitive Guide to AI/ML SaMD Ground Truthing [https://innolitics.com/articles/adjudication-strategies/]
- 68. Utilization of convolutional neural networks to analyze ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11245879/]
- 69. ICSI/IVF treatments allocation using CASAs compared to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11813069/]
- 70. Ensembled deep learning for the classification of human ... [https://research.monash.edu/en/publications/ensembled-deep-learning-for-the-classification-of-human-sperm-hea/]
- 71. Deep Learning Radiopathomics Models Based on Contrast ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11966553/]
- 72. Deep learning approaches for classification tasks in medical X ... [https://bmcmedimaging.biomedcentral.com/articles/10.1186/s12880-025-01701-5]
- 73. Deep learning-based image classification for AI-assisted ... [https://www.frontiersin.org/journals/medicine/articles/10.3389/fmed.2025.1574514/full]
- 74. A Survey on State-of-the-art Deep Learning Applications ... [https://arxiv.org/html/2403.17561v9]
- 75. Image Classification: State-of-the-Art Models in 2025 [https://hiringnet.com/image-classification-state-of-the-art-models-in-2025]
- 76. Classification of Human Sperm Based on Morphology ... [https://www.semanticscholar.org/paper/Classification-of-Human-Sperm-Based-on-Morphology-Aristoteles-Syarif/ef744a44e797ac3f66ecfde1548d694c435ff9f7]
- 77. Evaluating the factors influencing accuracy, interpretability ... [https://www.nature.com/articles/s41598-025-00245-6]
- 78. 5 – Reproducibility [https://blog.ml.cmu.edu/2020/08/31/5-reproducibility/]
- 79. A critical moment in machine learning in medicine [https://pmc.ncbi.nlm.nih.gov/articles/PMC10791964/]
- 80. Reproducible machine learning research in mental ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11039816/]
- 81. Towards reproducible machine learning-based process ... [https://www.sciencedirect.com/science/article/pii/S0952197625022316]