Optimizing Sperm Epigenetic Clocks: A Roadmap for Accurate Biomarkers in Male Fertility and Offspring Health
This article provides a comprehensive guide for researchers and drug development professionals on enhancing the precision and clinical utility of sperm epigenetic clocks.
Optimizing Sperm Epigenetic Clocks: A Roadmap for Accurate Biomarkers in Male Fertility and Offspring Health
Abstract
This article provides a comprehensive guide for researchers and drug development professionals on enhancing the precision and clinical utility of sperm epigenetic clocks. It explores the fundamental principles distinguishing sperm from somatic epigenetic aging, details advanced methodological approaches for clock construction—including machine learning on large, diverse datasets—and addresses key challenges such as tissue specificity and environmental confounders. Furthermore, it outlines rigorous validation frameworks and comparative analyses with other biomarkers, establishing sperm epigenetic age (SEA) as a novel, independent indicator of male fecundity and reproductive outcomes. The synthesis aims to accelerate the development of robust, clinically applicable tools for assessing paternal reproductive health and its intergenerational impacts.
The Basis of Sperm Epigenetic Aging: From Fundamental Principles to Clinical Correlations
Core Concepts: What is Sperm Epigenetic Age?
Answer: Sperm Epigenetic Age (SEA) is an estimate of the biological age of male gametes derived from DNA methylation patterns at specific genomic sites [1] [2]. It is determined using a sperm-specific epigenetic clock, which is a statistical model built via machine learning that analyzes age-related changes in the sperm DNA methylome [2]. SEA represents the molecular aging of sperm, which can diverge from the donor's chronological age, providing insights into his reproductive biological age [1] [3].
Key Distinctions: How Does the Sperm Epigenetic Clock Differ from Somatic Clocks?
Answer: Sperm epigenetic clocks are fundamentally different from somatic epigenetic clocks in their underlying DNA methylation patterns and the genomic sites used for age prediction.
The following table summarizes the core distinctions:
Table 1: Key Differences Between Sperm and Somatic Epigenetic Clocks
| Feature | Sperm Epigenetic Clocks | Somatic Epigenetic Clocks (e.g., Horvath, Hannum) |
|---|---|---|
| Target Cell | Male germ cells (sperm) [2] [4] | Somatic tissues (blood, saliva, etc.) [5] |
| Methylation Dynamics | Exhibit unique, sperm-specific age-related methylation changes; many regions show hypomethylation with age [4] [6] | Predominantly based on methylation patterns common across somatic tissues [5] |
| Relevant CpG Sites | Use loci specific to spermatogenesis (e.g., in genes like FOLH1, SH2B2, EXOC3) [4] [7] | Use loci predictive in somatic tissues (e.g., the Horvath clock uses 353 CpGs) [5] |
| Cross-Tissue Application | Not applicable to somatic tissues [5] | Designed for broad (pan-tissue) or specific (blood) somatic application [5] |
| Primary Context | Research on male fertility, fecundability, and offspring health [1] [3] | Research on general health, mortality, and age-related diseases [5] |
The pan-tissue Horvath clock, for instance, which accurately predicts age in diverse somatic tissues, performs poorly and significantly underestimates age when applied to sperm cells [4] [5]. This is because the sperm epigenome is uniquely structured and undergoes different aging dynamics compared to somatic cells [3].
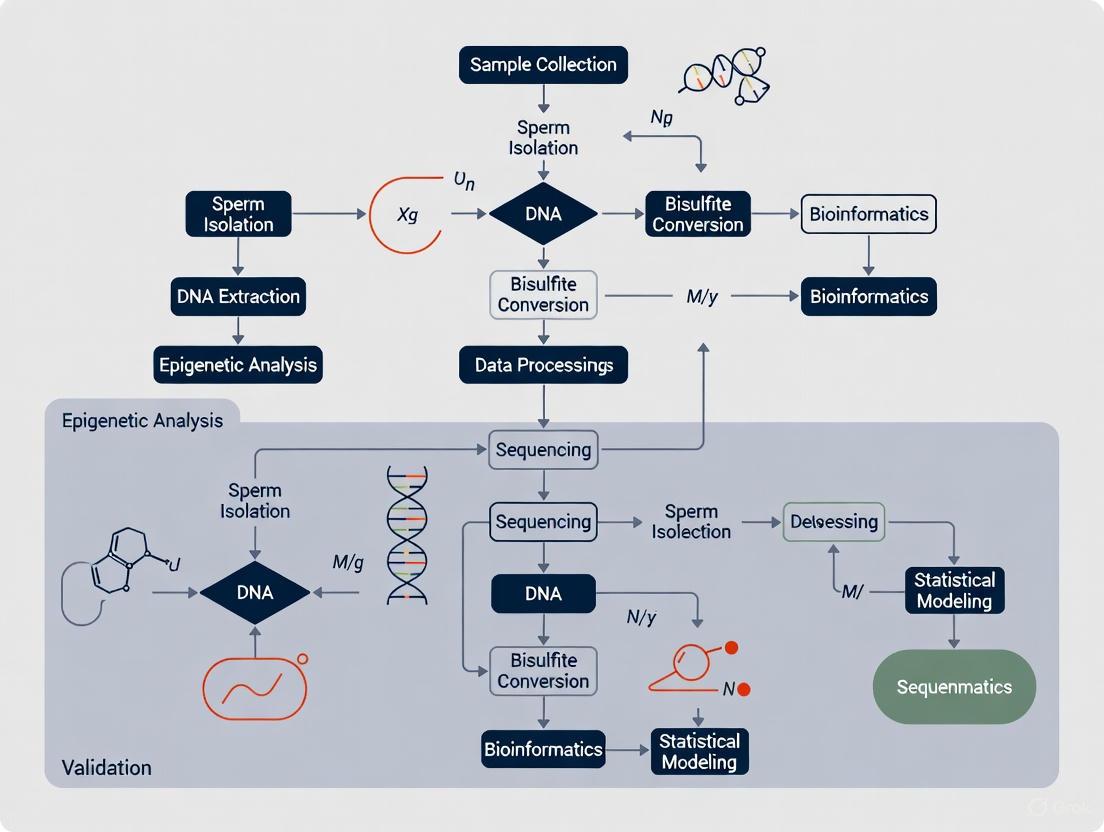
Experimental Protocols: How is SEA Measured?
Answer: Measuring SEA involves a multi-step process from semen sample collection to computational prediction. The workflow below outlines the key stages.
Detailed Methodology
Semen Sample Collection and Preparation:
- Cohorts: Studies often use both population-based cohorts (e.g., the Longitudinal Investigation of Fertility and Environment (LIFE) study) and clinical cohorts from fertility clinics (e.g., the Sperm Environmental Epigenetics and Development Study (SEEDS)) [1] [2].
- Collection: Samples are collected after a recommended period of ejaculatory abstinence (e.g., 2-3 days) [1] [8].
- Sperm Isolation: Sperm are isolated from seminal fluid using density gradient centrifugation to minimize somatic cell contamination [1] [8].
Sperm DNA Extraction:
DNA Methylation Profiling:
- Microarray-Based: The most common method uses the Illumina Infinium MethylationEPIC BeadChip, which Interrogates over 850,000 CpG sites across the genome [1] [2] [4].
- Sequencing-Based: For higher coverage and novel discovery, methods like reduced representation bisulfite sequencing (RRBS) or double-enzyme RRBS (dRRBS) are used. These are particularly valuable for identifying age-related CpG sites not covered by commercial arrays [6] [7]. For validation, Bisulfite Amplicon Sequencing (BSAS) is employed for targeted analysis of specific loci [4] [7].
Bioinformatic Processing and SEA Calculation:
- Quality Control (QC): Raw data undergoes normalization, dye bias correction, and removal of low-quality or cross-hybridizing probes. A key QC step is confirming minimal somatic cell contamination by checking methylation at imprinted genes like H19 and DLK1 [1] [2].
- Clock Application: A pre-trained sperm-specific epigenetic clock model is applied. These models are often built using ensemble machine learning algorithms (e.g., Super Learner) or penalized regressions that take the DNA methylation data from hundreds of samples as input to generate the SEA value [1] [2].
Table 2: Key Research Reagent Solutions for SEA Analysis
| Reagent / Material | Function / Application | Example & Notes |
|---|---|---|
| TCEP (Tris(2-carboxyethyl)phosphine) | Reducing agent for efficient sperm cell lysis and DNA extraction. | A stable alternative to DTT; used in rapid DNA extraction protocols [1] [8]. |
| Infinium MethylationEPIC BeadChip | Genome-wide DNA methylation profiling. | Covers >850,000 CpGs; standard for population studies [1] [4]. |
| dRRBS / RRBS Kits | Discovery of novel age-related CpG sites beyond microarray coverage. | Provides comprehensive, genome-wide methylation data; ideal for novel marker identification [6] [7]. |
| BSAS (Bisulfite Amplicon Sequencing) Reagents | Targeted validation of candidate age-related CpG sites. | Uses multiplex PCR and next-generation sequencing for high-sensitivity validation [4] [7]. |
| Sperm Isolation Kits (Density Gradient) | Purification of sperm cells from seminal plasma and somatic cells. | Critical for obtaining a pure sperm methylome signal [1] [8]. |
The Scientist's Toolkit: Troubleshooting Common SEA Experimental Challenges
Answer: Here are solutions to frequently encountered issues in SEA research.
FAQ 1: Our SEA predictions are inaccurate and inconsistent. What could be the cause?
- Somatic Cell Contamination: This is a primary concern. Somatic cells have vastly different methylation profiles. Always check for contamination by analyzing imprint control regions like H19/IGF2 [1] [6].
- Suboptimal DNA Extraction: Ensure your DNA extraction protocol is optimized for sperm, specifically including a robust reducing agent step to break protamine disulfide bonds [1].
- Inappropriate Epigenetic Clock: Verify that you are using a clock specifically trained on sperm DNA methylation data. Applying a somatic clock (like Horvath's) will yield erroneous results [4] [5].
FAQ 2: We have limited DNA from forensic or clinical samples. Which method should we use?
- For minimal DNA input, targeted approaches like Bisulfite Amplicon Sequencing (BSAS) are ideal. Studies have developed models with high accuracy (MAE ~3.3 years) using as few as 9 CpG sites, which is suitable for low-quantity and low-quality forensic DNA [7].
FAQ 3: Why are the age-related CpG sites in sperm different across studies?
- This is a common observation due to several factors:
- Technology: Different discovery platforms (450K vs. EPIC array vs. RRBS) cover different sets of CpGs [4] [7].
- Population Differences: Variations in ethnicity, geography, and lifestyle of the cohort can influence the specific loci identified [9].
- Statistical Power and Modeling: The choice of statistical models and algorithms can select different subsets of predictive CpGs from the highly correlated methylome [2] [6]. Despite this, the overall functional enrichment of these genes in developmental and neurological pathways is often consistent [6].
Data Presentation: Quantitative Associations of SEA
Answer: SEA shows specific associations with reproductive outcomes and morphological parameters, but not always with standard semen analysis.
Table 3: Documented Associations of Sperm Epigenetic Age from Research Studies
| Associated Factor | Association with SEA | Study Cohort & Citation |
|---|---|---|
| Time-to-Pregnancy (TTP) | Negative association. Advanced SEA linked to 17% lower probability of pregnancy within 12 months and longer TTP (FOR=0.83) [2]. | LIFE Study (General Population) [2] |
| Gestational Age at Birth | Negative association. Advanced SEA associated with shorter gestational age (-2.13 days) [2]. | LIFE Study (General Population) [2] |
| Sperm Head Morphology | Significant association. Higher SEA linked to increased head length and perimeter, more pyriform/tapered shapes, and lower elongation factor [1] [8]. | LIFE Study (General Population) [1] |
| Standard Semen Parameters | No significant association. SEA was not correlated with sperm count, concentration, or motility in clinical and non-clinical cohorts [1] [8]. | LIFE & SEEDS Cohorts [1] |
| Smoking | Positive association. Current smokers displayed advanced SEA [2]. | LIFE Study (General Population) [2] |
| Chronological Age | Strong positive correlation. Sperm clocks show high correlation with donor age (r = 0.91 in validation) [2] [4]. | Multiple Cohorts [2] [4] |
Visualizing the Impact of Advanced SEA
The diagram below synthesizes the documented biological and clinical associations of advanced Sperm Epigenetic Age, connecting molecular changes to potential phenotypic outcomes.
FAQs: Sperm Epigenetic Aging and Reproductive Outcomes
What is Sperm Epigenetic Aging? Sperm epigenetic aging refers to the biological age of sperm, which encapsulates cumulative genetic and environmental factors, rather than the father's chronological age. It is a novel biomarker that may better predict male reproductive contribution than conventional semen quality tests [10].
How does paternal age affect the genetic quality of sperm? As men age, harmful genetic changes in sperm become substantially more common. One landmark study found that while about 2% of sperm from men in their early 30s carried disease-causing mutations, this proportion rises to 3–5% in middle-aged and older men. By age 70, approximately 4.5% of sperm carry such mutations. This increase is driven not only by random DNA changes but also by a form of natural selection during sperm production that gives some harmful mutations a competitive edge [11].
What is the link between sperm epigenetic aging and time-to-pregnancy? Research has shown that higher sperm epigenetic aging is associated with a longer time to achieve pregnancy. One study reported a 17% lower cumulative probability of pregnancy after 12 months for couples where the male partner had older sperm epigenetic aging compared to those with younger epigenetic aging. This underscores the male partner's significant role in reproductive success [10].
What health implications for offspring are linked to older paternal age? Older paternal age is linked to an increased risk of passing on harmful genetic mutations. Researchers have identified 40 genes where certain DNA changes are favored during sperm production; many of these are linked to serious childhood diseases, severe neurodevelopmental disorders, and inherited cancer risk [11]. Furthermore, higher sperm epigenetic aging has been associated with shorter gestation periods in pregnancies that are achieved [10].
Troubleshooting Guides for Common Research Scenarios
Problem: Inconsistent Results in Sperm Epigenetic Clock Measurements
Description A researcher encounters high variability when measuring the sperm epigenetic age across different samples within the same study cohort, leading to unreliable data.
Solution Follow a systematic troubleshooting process to isolate and resolve the issue.
Understand the Problem:
- Ask: Review laboratory notebooks. Were there any changes in reagent lots, personnel, or equipment calibration around the time the inconsistency appeared?
- Gather Information: Compile all quality control metrics from the sample processing runs (e.g., DNA yield, purity ratios, bisulfite conversion efficiency). Check if the inconsistencies correlate with a specific sample batch, processing day, or technician.
- Reproduce the Issue: Re-run the epigenetic clock assay on a subset of samples with previously stable readings to see if the inconsistency persists.
Isolate the Issue:
- Remove Complexity: Simplify the workflow to identify the problematic stage.
- Test DNA Extraction: Process a control sample with a known epigenetic age using a fresh, certified reagent kit.
- Test Bisulfite Conversion: Run a control DNA with a known conversion rate to ensure this critical step is performing optimally.
- Change One Thing at a Time: If the problem persists, systematically test individual components of the PCR or sequencing reaction, such as primers or polymerase enzymes.
- Compare to a Working Version: Compare the entire workflow, from sample collection to data analysis, against the standard operating procedure established during earlier, successful experiments. Look for any unintentional deviations.
- Remove Complexity: Simplify the workflow to identify the problematic stage.
Find a Fix or Workaround:
- If the issue is traced to a specific reagent lot, discontinue its use and validate a new lot.
- If the problem is with a specific instrument, perform maintenance and re-calibration.
- Document the root cause and the solution in your lab's protocol to prevent future occurrences.
Problem: Low Statistical Power in Associating SEA with Time-to-Pregnancy
Description A research team finds that the association between Sperm Epigenetic Aging (SEA) and couple's time-to-pregnancy is not statistically significant, potentially due to study design limitations.
Solution
Understand the Problem:
- Ask: What is the current sample size? What is the effect size you are trying to detect? What is the prevalence of the outcome in your population?
- Gather Information: Perform a power analysis retrospectively to determine if the study was adequately powered from the outset. Examine the distribution of both SEA values and time-to-pregnancy data for anomalies.
Isolate the Issue:
- The core issue is often an insufficient number of participants for a relatively rare outcome or a small effect size.
- Check for confounding variables that were not controlled for, such as female partner's age, lifestyle factors (e.g., smoking status of the male partner, which is known to affect epigenetic aging [10]), or unaccounted fertility treatments.
Find a Fix or Workaround:
- The primary fix is to increase the sample size. Consider collaborating with other research institutions to create a larger, multi-center cohort.
- If increasing sample size is not feasible, consider refining the phenotype. For example, focus on couples with confirmed infertility or stratify the analysis based on the female partner's age or ovarian reserve.
- Ensure that diverse races and ethnicities are included, as initial findings were based on a largely Caucasian cohort and require confirmation in other groups [10].
Experimental Protocols from Key Studies
Protocol 1: Sperm Collection and DNA Methylation Analysis for Epigenetic Clock Construction
This methodology is adapted from the Wayne State University study that developed a novel measure of sperm epigenetic age [10].
1. Participant Recruitment and Sperm Sample Collection
- Recruit male partners from couples who have recently discontinued contraception for the purpose of becoming pregnant.
- Collect semen samples following standard clinical protocols. Record detailed participant metadata, including chronological age, smoking status, and medical history.
- Key Reagent: Standard semen collection kits.
2. Sperm DNA Extraction and Purification
- Isolate sperm cells from the seminal plasma using density gradient centrifugation.
- Extract genomic DNA using a commercial kit designed for sperm cells, which are notoriously resistant to lysis. Ensure high DNA purity (A260/A280 ratio ~1.8) and integrity (check via gel electrophoresis).
- Key Reagent: Sperm-specific DNA extraction kit (e.g., Qiagen QIAamp DNA Mini Kit with optimized lysis protocols).
3. Bisulfite Conversion and Microarray Analysis
- Treat extracted DNA with sodium bisulfite using a dedicated kit. This process converts unmethylated cytosines to uracils, while methylated cytosines remain unchanged.
- Hybridize the bisulfite-converted DNA to a genome-wide methylation microarray, such as the Illumina Infinium MethylationEPIC BeadChip.
- Key Reagent: Bisulfite conversion kit (e.g., Zymo Research EZ DNA Methylation Kit); Illumina MethylationEPIC BeadChip.
4. Computational Construction of the Epigenetic Clock
- Process raw microarray data using R packages like
minfifor normalization and background correction. - Use a penalized regression model (e.g., ElasticNet) to identify a subset of CpG sites whose methylation levels collectively predict chronological age. This model becomes the "epigenetic clock."
- Sperm epigenetic age is calculated by applying this model to new methylation data. The difference between epigenetic age and chronological age indicates biological aging (e.g., age acceleration).
Protocol 2: NanoSeq for Ultra-Accurate Mutation Detection in Sperm
This methodology is adapted from the landmark study that mapped harmful DNA changes in sperm with unprecedented precision [11].
1. Sperm Sample Preparation and DNA Sequencing
- Obtain sperm samples from a well-characterized cohort (e.g., a twin registry). Include men across a broad age range (e.g., 24-75 years).
- Extract sperm DNA as described in Protocol 1.
- Prepare sequencing libraries and perform deep duplex sequencing using the NanoSeq method. This technique sequences both strands of DNA independently, dramatically reducing sequencing error rates.
- Key Reagent: NanoSeq library preparation reagents; Illumina sequencing platforms.
2. Variant Calling and Filtering
- Align sequencing reads to the human reference genome.
- Identify single-nucleotide variants (SNVs) using variant callers optimized for duplex sequencing data. Stringent filters are applied to remove technical artifacts and retain only high-confidence mutations.
- Key Reagent: High-performance computing cluster with sufficient RAM and storage.
3. Analysis of Clonal Expansion and Selection
- Compare mutation spectra and burdens across different age groups.
- Identify "driver genes" by looking for genes that are mutated more frequently than expected by chance. This signals positive selection during sperm production.
- Correlate the presence of mutations in specific genes (e.g., those linked to childhood disorders or cancer) with the age of the donor.
The following tables consolidate key quantitative findings from the reviewed literature.
Table 1: Paternal Age and Mutation Burden in Sperm
| Metric | Men in Early 30s | Middle-Aged Men (43-58) | Older Men (59-74) | Age 70 | Source |
|---|---|---|---|---|---|
| Sperm carrying disease-causing mutations | ~2% | 3-5% | 3-5% | ~4.5% | [11] |
| Key Driver | Steady DNA change buildup | Natural selection in testes | Natural selection in testes | Natural selection in testes | [11] |
Table 2: Impact of Sperm Epigenetic Aging on Pregnancy Outcomes
| Metric | Finding | Impact / Notes | Source |
|---|---|---|---|
| Pregnancy Probability | 17% lower after 12 months | For couples with male partners in older vs. younger sperm epigenetic aging categories | [10] |
| Gestation Length | Associated with shorter gestation | Among couples that achieved pregnancy | [10] |
| Environmental Factor | Higher aging in men who smoked | Modifiable risk factor | [10] |
Research Reagent Solutions
Table 3: Essential Research Materials for Sperm Epigenetic Clock and Mutation Studies
| Item | Function | Example / Specification |
|---|---|---|
| Sperm DNA Extraction Kit | Isolves high-quality, intact genomic DNA from resilient sperm cells. | Qiagen QIAamp DNA Mini Kit (with protocol modifications for sperm) |
| Bisulfite Conversion Kit | Converts unmethylated cytosine to uracil for downstream methylation analysis. | Zymo Research EZ DNA Methylation Kit |
| DNA Methylation Microarray | Profiles genome-wide methylation levels at single-base resolution. | Illumina Infinium MethylationEPIC BeadChip |
| NanoSeq Library Prep Reagents | Enables ultra-accurate duplex sequencing by tracking both DNA strands. | As described in the Neville et al. Nature 2025 protocol [11] |
| CpG Site Validation Primers | Validates clock-associated CpG sites using targeted bisulfite pyrosequencing or PCR. | Custom-designed, HPLC-purified primers |
Experimental Workflow and Signaling Pathways
Workflow for Sperm Epigenetics and Mutational Analysis
FAQs: Sperm Epigenetic Aging (SEA) and Male Fertility
Q1: What is Sperm Epigenetic Age (SEA), and how does it differ from chronological age? Sperm Epigenetic Age (SEA) is a measure of the biological age of sperm cells, derived from specific patterns of DNA methylation at CpG sites across the genome. Unlike chronological age, which is simply the time since birth, SEA reflects the cumulative biological impacts of internal factors (like genetics) and external factors (such as environment and lifestyle) on sperm cells. Research shows that an advanced SEA is associated with a longer time for a couple to achieve pregnancy, independent of the man's chronological age [8] [12].
Q2: Is Sperm Epigenetic Age associated with standard semen analysis parameters? Interestingly, SEA has been found to be largely independent of standard semen parameters like sperm concentration, motility, and volume [8]. However, it shows significant associations with more specific, less routinely measured parameters. Specifically, an advanced SEA is linked to aberrations in sperm head morphology, including higher sperm head length and perimeter, the presence of pyriform and tapered sperm, and a lower sperm elongation factor [8].
Q3: How does lifestyle, particularly smoking, impact the sperm epigenome? Lifestyle choices have a measurable impact on sperm epigenetic age. Studies have consistently shown that smoking is associated with advanced SEA [12] [13]. Smokers exhibit a significantly higher sperm epigenetic age compared to non-smokers, highlighting the reversible yet impactful nature of epigenetic modifications on male reproductive health [14].
Q4: Can the biological aging of sperm be reversed? Epigenetic marks, including DNA methylation, are fundamentally reversible. This reversability suggests that interventions, potentially through lifestyle changes such as improved diet, cessation of smoking, or supplementation (e.g., with Zinc and Folic acid), could help "rejuvenate" the sperm epigenome and promote a younger sperm epigenetic age [13].
Troubleshooting Guides for SEA Research
Table 1: Common Experimental Challenges in Sperm Epigenetic Clock Research
| Challenge | Potential Cause | Solution |
|---|---|---|
| Low DNA yield from sperm samples | Inefficient cell lysis due to unique sperm chromatin packaging. | Implement a lysis buffer containing a reducing agent like Tris(2-carboxyethyl)phosphine (TCEP) to break down protamine-based packaging [8]. |
| Inaccurate epigenetic age prediction | Use of clocks designed for somatic cells, which have different methylation patterns. | Develop and use a sperm-specific epigenetic clock based on CpG sites identified from semen-derived DNA [15]. |
| Inconsistencies in sample processing | Differing density gradient centrifugation methods between clinical and research cohorts. | Standardize the sperm isolation protocol across all samples, ideally using a validated, multi-step density gradient centrifugation method [8]. |
| Confounding by cell composition | Age-related shifts in the composition of somatic cells within semen samples. | Isinate sperm cells from semen samples prior to DNA extraction to ensure the methylation profile is specific to sperm [8] [16]. |
Detailed Protocol: Sperm DNA Isolation for Methylation Analysis
The integrity of DNA methylation analysis is highly dependent on the quality of the initial DNA extraction. The following protocol is adapted from a method used in clinical and research cohorts [8].
Principle: Sperm DNA is packaged with protamines instead of histones, requiring a reducing agent for efficient lysis and DNA purification.
Reagents Needed:
- Lysis Buffer: Containing guanidine thiocyanate and 50 mM Tris(2-carboxyethyl)phosphine (TCEP)
- 0.2 mm steel beads
- Silica-based spin columns (e.g., from Qiagen or similar)
- Proteinase K (optional, for increased yield)
Procedure:
- Homogenization: Transfer the sperm sample to a tube containing 0.2 mm steel beads and the lysis buffer with TCEP.
- Lysis: Homogenize the mixture at room temperature for 5 minutes. The TCEP is a stable reducing agent that effectively disrupts protamine-DNA complexes.
- DNA Purification: Transfer the lysate to a silica-based spin column and proceed with the manufacturer's standard washing and elution steps. This method consistently yields over 90% high-quality DNA and avoids lengthy Proteinase K digestions [8].
Key Experimental Workflows and Signaling Pathways
Sperm Epigenetic Clock Development Workflow
The following diagram illustrates the key steps involved in creating a sperm-specific epigenetic clock, from sample collection to model validation.
DNA Methylation and Demethylation Pathway
This diagram outlines the core molecular mechanism of DNA methylation, a key process measured by epigenetic clocks.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Sperm Epigenetics Research
| Item | Function/Application in Research | Example Use Case |
|---|---|---|
| Infinium MethylationEPIC BeadChip | Genome-wide DNA methylation profiling of over 850,000 CpG sites. | Discovery of novel, age-correlated differentially methylated sites (DMSs) in sperm DNA [15]. |
| Tris(2-carboxyethyl)phosphine (TCEP) | Reducing agent for efficient lysis of protamine-packaged sperm DNA. | Key component in rapid, room-temperature sperm DNA extraction protocols [8]. |
| Sperm-Specific Epigenetic Clock Model | A predictive model using specific CpG sites to estimate biological age from sperm DNA. | Assessing the impact of environmental exposures or lifestyle on sperm biological age (SEA) [8] [15]. |
| Targeted Bisulfite MPS Panels | Validation and precise quantification of methylation levels at candidate CpGs. | Confirming age-correlation of DMSs discovered by microarray in an independent sample set [15]. |
| Computer-Assisted Semen Analysis (CASA) | Automated, detailed analysis of sperm concentration, motility, and morphology. | Correlating advanced SEA with specific defects in sperm head morphology [8]. |
Troubleshooting Guide: Frequently Asked Questions
Q1: Our lab's sperm morphology assessments show high variability between technicians. How can we improve consistency?
A1: High inter-technician variability is a common challenge, primarily due to the subjective nature of traditional morphology assessment [17]. A 2025 study demonstrated that without standardized training, novice morphologists showed high variation (Coefficient of Variation = 0.28) and accuracies as low as 53% when using a complex 25-category classification system [17].
- Solution: Implement a standardized digital training tool based on machine learning principles. One study used a "Sperm Morphology Assessment Standardisation Training Tool" with images classified by expert consensus ("ground truth") [17].
- Result: After four weeks of repeated training, accuracy significantly improved from 82% to 90%, classification speed increased, and variation between technicians was greatly reduced [17]. For the highest accuracy, use simpler classification systems (2-category: normal/abnormal) before moving to complex ones [17].
Q2: Are traditional sperm morphology parameters like "percent normal forms" clinically relevant for predicting ART outcomes?
A2: Recent expert guidelines have significantly shifted the answer to this question. The French BLEFCO Group's 2025 review recommends against using the percentage of normal forms as a prognostic tool for selecting between IUI, IVF, or ICSI [18]. They concluded that the overall level of evidence for the clinical value of this parameter is low [18].
- Solution: Focus morphology assessment on the detection of specific, monomorphic abnormalities, such as globozoospermia or macrocephalic spermatozoa syndrome, which have clear clinical implications [18]. The working group also gives a positive opinion on using qualified and validated automated systems based on cytological analysis after staining to reduce subjectivity [18].
Q3: How can environmental factors confound research on sperm epigenetics and morphology?
A3: Environmental toxicants are a major confounder in male fertility research. Exposure to endocrine-disrupting chemicals (EDCs), air pollution, and heavy metals can induce oxidative stress, leading to sperm DNA fragmentation, morphological alterations, and epigenetic changes [19] [20].
- Mechanism: Toxicants like particulate matter (PM2.5) and polycyclic aromatic hydrocarbons (PAHs) can generate reactive oxygen species (ROS), causing lipid peroxidation and DNA damage [19]. Furthermore, these chemicals can create DNA adducts and alter DNA methylation patterns, which are critical for the accuracy of sperm epigenetic clock research [19].
- Recommendation: Document and account for participants' environmental exposures (e.g., smoking, occupational hazards) as these can advanced sperm epigenetic aging, a biomarker associated with longer time-to-pregnancy [2] [20].
Q4: What functional sperm tests can we use to complement basic morphology in an epigenetic study?
A4: Moving beyond static morphology to functional and chromatin integrity assays provides a more comprehensive view for epigenetic research.
- Flow Cytometry: Use multiparametric flow cytometry to assess sperm viability, acrosomal integrity, membrane stability, and mitochondrial status [21]. This technique offers high-throughput, accurate analysis of sperm function parameters [21].
- Sperm Chromatin Integrity: Evaluate protamination, condensation, and DNA integrity [22]. The sperm epigenome is shaped by histone retention, DNA methylation, and RNAs; aberrant integrity can negatively impact reproductive success and is a key variable in embryo development [22].
Summarized Data Tables
| Classification System | Untrained User Accuracy (%) | Final Accuracy After Training (%) |
|---|---|---|
| 2-Category (Normal/Abnormal) | 81.0 ± 2.5 | 98.0 ± 0.4 |
| 5-Category (Head, Midpiece, etc.) | 68.0 ± 3.6 | 97.0 ± 0.6 |
| 8-Category (Cattle Industry) | 64.0 ± 3.5 | 96.0 ± 0.8 |
| 25-Category (Individual Defects) | 53.0 ± 3.7 | 90.0 ± 1.4 |
| Outcome Measure | Association with Advanced Sperm Epigenetic Aging | Study Details |
|---|---|---|
| Time-to-Pregnancy (TTP) | 17% lower cumulative probability at 12 months | FOR=0.83; 95% CI: 0.76, 0.90; P = 1.2×10⁻⁵ |
| Gestational Age | Shorter by 2.13 days | 95% CI: -3.67, -0.59; P = 0.007 (n=192) |
| Chronological Age | High predictive correlation (r = 0.91) | Population-based prospective cohort (n=379) |
Experimental Protocols
Protocol 1: Standardized Sperm Morphology Assessment Using a Training Tool
Objective: To minimize inter-technician variability and improve the accuracy of sperm morphology classification.
Materials: Standardized digital image library with expert-consensus "ground truth" labels, computer-based training tool [17].
Methodology:
- Baseline Testing: Have technicians perform an initial classification test on a set of images using the desired category system (e.g., 2-category, 5-category).
- Structured Training: Expose technicians to the training tool, which provides immediate feedback on their classifications against the expert consensus.
- Repeated Practice: Implement a schedule of repeated training and testing over several weeks (e.g., tests over 4 weeks).
- Proficiency Assessment: Monitor improvements in accuracy and reduction in time taken per classification. Continue training until a pre-defined accuracy threshold (e.g., >90% for the target category system) is consistently met [17].
Protocol 2: Sperm Functional Analysis by Flow Cytometry
Objective: To perform a multiparametric assessment of sperm function parameters, complementing morphology and epigenetic data.
Materials: Flow cytometer, fluorochromes, semen sample, specific stains for viability (e.g., SYBR Green/Propidium Iodide [23]), acrosomal status, mitochondrial membrane potential, and oxidative stress [21].
Methodology:
- Sample Preparation: Aliquot liquefied semen and stain with the appropriate combination of fluorescent probes.
- Instrument Setup: Calibrate the flow cytometer using appropriate controls. Adjust settings for forward scatter, side scatter, and fluorescence detectors based on the fluorochromes used [21].
- Data Acquisition: Acquire a minimum of 10,000 events per sample. Use a gate to exclude debris and aggregates, focusing on the single sperm population.
- Data Analysis: Analyze the fluorescence data to determine the percentage of viable sperm, sperm with intact acrosomes, stable membranes, and high mitochondrial membrane potential [21].
Workflow Diagrams
Sperm Quality & Epigenetics Research Pathway
Morphology Training & Standardization Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Sperm Quality and Epigenetics Research
| Reagent / Material | Primary Function in Research | Key Considerations |
|---|---|---|
| Fluorochrome Kits for Flow Cytometry [21] | Multiparametric assessment of sperm viability, acrosomal integrity, mitochondrial membrane potential, and oxidative stress. | Allows high-throughput, objective analysis of sperm function. |
| SYBR Green/Propidium Iodide [23] | Fluorescent live/dead staining for sperm viability assessment. Correlates well with motility. | Suitable for both conventional microscopy and CASA systems. |
| Methylation Microarray/Sequencing Kits [2] | Profiling sperm DNA methylation for constructing epigenetic clocks (SEA). | Machine learning algorithms are then applied to predict biological age from methylation data. |
| Standardized Digital Morphology Library [17] | Training and standardizing technicians to reduce subjective bias in morphology assessment. | Must be built on expert consensus ("ground truth") for reliable training. |
| Antioxidant Supplements (in vitro) | Mitigating oxidative stress induced by environmental toxicants during sample processing [19]. | Can help maintain sperm membrane and DNA integrity during assays. |
Building Superior Sperm Epigenetic Clocks: Methodologies, Machine Learning, and Model Training
Frequently Asked Questions (FAQs)
FAQ 1: What is the fundamental difference between a pan-tissue and a sperm-specific epigenetic clock?
Pan-tissue epigenetic clocks are designed to predict chronological age across multiple tissue types. They are trained on DNA methylation data from diverse tissues (e.g., blood, brain, liver) to identify age-related methylation patterns that are universal. The classic Horvath clock, which uses 353 CpG sites, is a prime example [24] [25]. In contrast, a sperm-specific clock would be trained exclusively on sperm samples to capture aging signals unique to the male germline. These signals may be linked to specialized biological processes like spermatogenesis and the unique epigenetic reprogramming that occurs in sperm [26].
FAQ 2: My research aims to link male biological aging to offspring health. Why should I consider a sperm-specific clock instead of a established pan-tissue clock?
Using a pan-tissue clock on sperm may miss or miscalibrate the specific aging processes of the male germline. Sperm cells have a unique epigenetic landscape, including widespread DNA hypomethylation in certain genomic regions. A pan-tissue clock, optimized for somatic tissues, may not be sensitive to the subtle, biologically critical age-related changes in sperm [24] [26]. Furthermore, advanced paternal age is associated with increased risk of neurodevelopmental disorders in offspring due to mutations in sperm [27]. A purpose-built sperm clock is more likely to detect such age-related deterioration relevant to reproductive outcomes, making it a more appropriate tool for your research on intergenerational health.
FAQ 3: What are the key technical challenges in developing an accurate sperm-specific epigenetic clock?
Key challenges include:
- Cellular Homogeneity: Sperm samples are more cellularly homogeneous than heterogeneous tissues like blood. While this reduces one confounding factor, it places greater demand on the accuracy of the methylation assay to detect small, true age-related changes [24].
- Magnitude of Change: Age-related methylation changes at individual CpG sites are often very small, with one large-scale analysis finding an average lifetime change of only ~1.5% [24]. This requires precise measurement and large sample sizes for robust clock development.
- Biological Interpretation: Even with an accurate clock, understanding whether the selected CpG sites are causal in aging processes or simply correlated with time is a significant challenge. The relationship between "accelerated" epigenetic aging in sperm and specific functional deficits is an active area of research [24] [25].
Troubleshooting Guides
Issue 1: Inconsistent age predictions from a pan-tissue clock when applied to sperm samples.
| Possible Cause | Solution |
|---|---|
| Fundamental Tissue Difference | This is the most likely cause. Pan-tissue clocks are calibrated for somatic tissues. The solution is to use or develop a clock trained specifically on sperm methylation data. |
| Inappropriate Control for Cellular Composition | While sperm is relatively homogeneous, contamination with somatic cells (e.g., white blood cells) can skew results. Purify sperm cells using a standardized density gradient isolation procedure before DNA extraction [26] [28]. |
| Technical Assay Variation | Ensure consistent and accurate DNA methylation measurement. Use high-quality bisulfite conversion methods and consider high-resolution platforms like the Illumina Infinium MethylationEPIC array for broader genomic coverage [29]. |
Issue 2: Weak association between epigenetic age acceleration in sperm and phenotypic outcomes (e.g., pregnancy success).
| Possible Cause | Solution |
|---|---|
| Clock Not Fit for Purpose | The clock you are using may be trained only on chronological age, not on the phenotype of interest. Consider developing a "second-generation" clock trained on phenotypic outcomes (e.g., sperm motility, DNA fragmentation) in addition to age [25]. |
| Confounding Factors | Factors like paternal abstinence time significantly influence standard semen quality parameters and sperm DNA fragmentation index (DFI) [28]. Control for and record these variables meticulously in your experimental design. A standardized abstinence period (e.g., 2-4 days) is recommended. |
| Insufficient Statistical Power | The effect size may be small. Increase your sample size. Large-scale analyses, such as one involving over 6,000 samples, are often needed to detect clear age-related trends in sperm parameters [30]. |
Table 1: Documented Effects of Male Aging on Sperm Parameters This table synthesizes findings from large-scale clinical studies on how advancing age affects measurable sperm quality and DNA integrity [30] [28].
| Parameter | Documented Change with Advancing Age | Clinical Context & Notes |
|---|---|---|
| Semen Volume | Significant decline [30] [28] | Associated with age-related changes in accessory gland function (e.g., prostate) [31]. |
| Sperm Motility (Progressive & Total) | Significant decline [30] | A key factor in reduced natural fertility potential with age [31]. |
| Sperm DNA Fragmentation Index (DFI) | Significant increase [30] [28] | A DFI >30% is linked to challenges in natural conception and embryo development [30]. |
| Incidence of Harmful Mutations | Increases from ~2% (age 30) to ~4.5% (age 70) [27] | These are de novo mutations in sperm, linked to neurodevelopmental disorders in offspring [27]. |
Table 2: Comparison of Epigenetic Clock Generations This table outlines the evolution of epigenetic clocks, which is critical for selecting the right tool for your research question [32] [25].
| Generation | Primary Training Target | Example Clocks | Utility for Sperm Research |
|---|---|---|---|
| First | Chronological Age | Horvath, Hannum | Useful for basic age prediction; may lack biological relevance to sperm function. |
| Second | Biomarkers & Mortality | PhenoAge, GrimAge | More likely to capture health-related aging processes; potential model for sperm clocks trained on sperm quality. |
| Third | Pace of Aging | DunedinPACE | Measures the rate of aging; concept could be applied to model the pace of sperm quality decline. |
| Fourth | Causality (via Mendelian randomization) | Causal Clocks | Aims to identify CpG sites causally involved in aging; the future goal for understanding sperm aging mechanisms. |
Detailed Experimental Protocols
Protocol 1: Standardized Sperm Collection, Purification, and DNA Methylation Analysis
This protocol is adapted from methodologies used in recent studies on sperm epigenetics [26] [28].
Participant Selection and Semen Collection:
- Recruit participants according to approved ethical guidelines, obtaining informed consent.
- Record relevant metadata: age, abstinence time, smoking status, BMI, and medical history.
- Collect semen samples by masturbation after a recommended abstinence period of 2-7 days [28]. Allow samples to liquefy for 30 minutes at room temperature.
Sperm Quality Analysis:
Sperm Purification:
- Isolate and purify sperm cells using a discontinuous density gradient centrifugation procedure [26].
- Layer 1 mL of semen over a gradient medium (e.g., 1 mL of 40% over 1 mL of 80% density-gradient medium).
- Centrifuge at 400 x g for 15 minutes. Discard the supernatant and resuspend the resulting sperm pellet in a suitable buffer (e.g., Phosphate-Buffered Saline or Ham's F10 medium). Repeat washing steps [26].
DNA Extraction and Bisulfite Conversion:
- Extract genomic DNA from the purified sperm pellet using a commercial kit (e.g., QIAamp DNA Blood Mini Kit) [29].
- Treat the DNA with bisulfite using a dedicated kit (e.g., EZ DNA Methylation Kit from Zymo Research) to convert unmethylated cytosines to uracils, while leaving methylated cytosines unchanged [29].
Genome-Wide Methylation Profiling:
- Analyze the bisulfite-converted DNA using a high-throughput platform such as the Illumina Infinium MethylationEPIC BeadChip, which interrogates over 850,000 CpG sites.
- Process the raw data in GenomeStudio or R to obtain beta-values (a measure of methylation level from 0 to 1) for each CpG site [29].
Protocol 2: Building a Sperm-Specific Epigenetic Clock
Data Collection and Preprocessing:
- Assemble a large dataset (n > 1000 recommended) of sperm methylation beta-values from donors across a wide age range (e.g., 20-60 years).
- Perform rigorous quality control and normalization of the methylation data. Correct for potential technical artifacts and batch effects.
Clock Training with Penalized Regression:
- Use a supervised machine learning method, such as elastic net regression, to train the clock model [24] [25].
- Input: The methylation levels of all high-quality CpG sites.
- Output: The chronological age of the donors.
- The elastic net algorithm will automatically select a sparse set of CpG sites whose weighted methylation levels best predict chronological age.
Validation and Phenotypic Association:
- Validate the clock's accuracy on a separate, independent set of sperm samples not used in the training.
- Calculate "Age Acceleration" (the residual from regressing epigenetic age on chronological age) for each sample [25] [29].
- Statistically test whether this age acceleration is correlated with phenotypic outcomes like sperm DFI, motility, or pregnancy success in ART cycles [30].
Workflow and Relationship Diagrams
Sperm Clock Development Workflow
Tissue Selection Decision Guide
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Sperm Epigenetic Clock Research
| Item | Function in the Protocol | Example Product / Specification |
|---|---|---|
| Density Gradient Medium | To isolate and purify viable sperm from semen and remove somatic cell contamination. | SilSelect (Fertipro), PureSperm (Nidacon) |
| DNA Extraction Kit | To obtain high-quality, high-molecular-weight genomic DNA from purified sperm cells. | QIAamp DNA Blood Mini Kit (QIAGEN) |
| Bisulfite Conversion Kit | To convert unmethylated cytosine to uracil for subsequent methylation analysis. | EZ DNA Methylation Kit (Zymo Research) |
| Methylation Array | For genome-wide, high-throughput quantification of DNA methylation levels at specific CpG sites. | Illumina Infinium MethylationEPIC BeadChip |
| Sperm DNA Integrity Assay Kit | To measure sperm DNA fragmentation, a key phenotypic correlate of sperm quality and aging. | Sperm Chromatin Structure Assay (SCSA) kit |
| Statistical Software | For data normalization, clock construction (elastic net regression), and statistical analysis. | R with glmnet package, SPSS |
In the specialized field of sperm epigenetic clock research, the volume and quality of training data are not merely technical details—they are fundamental determinants of predictive accuracy and clinical utility. Sperm epigenetic age (SEA) has emerged as a significant biomarker, demonstrating associations with time-to-pregnancy and specific sperm morphological factors, even when standard semen parameters appear normal [1]. Unlike somatic cells, sperm exhibit unique epigenetic aging patterns that require specialized prediction models [33] [34]. The construction of accurate epigenetic clocks relies on machine learning algorithms that identify age-associated DNA methylation patterns from training data. As these models are increasingly applied to assess male fertility potential and reproductive outcomes, understanding how training set size influences their performance becomes paramount for advancing both basic research and clinical applications.
Technical FAQs: Data Requirements for Sperm Epigenetic Clocks
How does training set size specifically affect sperm epigenetic age prediction accuracy?
The relationship between training set size and prediction accuracy follows a principle of diminishing returns. Initial increases in sample size yield substantial improvements in model precision, but these gains gradually plateau as the training set becomes more comprehensive.
Quantitative Evidence from Epigenetic Research: A 2024 study developing epigenetic clocks resistant to immune cell composition changes utilized a massive database of 14,601 DNA methylation samples from 71 datasets to ensure robust performance across cell types [16]. While this exemplifies the scale used for somatic clocks, sperm-specific models show that carefully selected markers can achieve reasonable accuracy with smaller, targeted datasets. For instance, one sperm epigenetic clock study utilized 379 men from a non-clinical cohort and 192 from a clinical cohort, demonstrating that SEA could be associated with sperm head morphology despite the moderate sample size [1].
Machine Learning Performance Patterns: General machine learning principles confirm that prediction performance typically scales as a power law with dataset size. One analysis found that across six datasets of varying sizes, training an XGBoost classifier on just 30% of the data could retain at least 95% of the performance achievable with the full dataset [35]. The following table summarizes how prediction performance typically evolves with expanding training sets:
Table: Relationship Between Training Set Size and Model Performance
| Training Set Size Range | Expected Impact on Sperm Epigenetic Clock | Typical Performance Metrics |
|---|---|---|
| Small (n < 100) | High variance, substantial risk of overfitting to donor-specific patterns | RMSE: ~5-10 years [34]; Limited generalizability |
| Moderate (n = 100-500) | Improved stability, better capture of population variation | RMSE: ~3-5 years; Beginning of plateau effect |
| Large (n > 500) | Diminishing returns, enhanced detection of subtle effects | RMSE: ~2-3 years [36]; More robust biological insights |
What constitutes a "sufficient" training set for sperm epigenetic clock development?
Sufficiency depends on multiple factors including the desired precision, population diversity, and biological complexity of the targeted aging process. For sperm epigenetic clocks, the longitudinal stability of methylomes within individuals means that between-donor variation far exceeds within-donor variation, necessitating careful sample selection [33].
Key Considerations for Determining Sample Size:
- Feature-to-Sample Ratio: Maintain a high ratio of samples to DNA methylation markers analyzed. Studies incorporating sex chromosomal markers alongside autosomal markers have utilized training sets of 860 whole blood samples to achieve RMSE of 2.54 years [36].
- Population Heterogeneity: Ensure representation across age ranges, ethnicities, and clinical statuses. One whole-genome bisulfite sequencing study of sperm used a longitudinal design with 10 donors sampled 10-18 years apart to control for individual variation [33].
- Validation Strategy: Allocate sufficient samples for hold-out testing. The 2024 sperm epigenetic age study used separate clinical (SEEDS) and non-clinical (LIFE) cohorts for validation, demonstrating consistent associations with sperm morphology across populations [1].
Why would increasing training data sometimes fail to improve predictions?
Despite the general principle that more data enhances accuracy, several scenarios can diminish or negate these benefits in sperm epigenetic clock research:
- Data Quality Issues: Low-quality methylation data from challenging genomic regions like sex chromosomes can introduce noise that outweighs the benefits of additional samples [36].
- Irrelevant Training Data: Adding samples that don't match the application context provides limited value. One voice recognition experiment found that a small, highly relevant dataset outperformed a much larger but less applicable one [37].
- Model Capacity Limitations: A simple model with limited parameters may be unable to capture additional patterns from increased data. Model complexity must align with dataset size [38].
- Platform Batch Effects: Technical artifacts from combining datasets across different methylation array batches or sequencing platforms can introduce confounding variation [33].
Troubleshooting Guides: Addressing Common Data-Related Challenges
Problem: Diminishing returns in prediction accuracy despite adding more training samples
Diagnosis: The model may have reached its performance plateau given current features and architecture.
Solution Strategy:
- Enhance Feature Quality Rather Than Quantity: Identify and focus on highly predictive methylation markers. A 2025 study achieved significant improvement (RMSE: 2.54 years) by combining just four X chromosomal markers with six autosomal markers, rather than using thousands of probes [36].
- Incorporate Complementary Data Types: Consider adding relevant clinical parameters or environmental exposure data that may explain residual variance in epigenetic aging [1].
- Optimize Model Architecture: Experiment with more sophisticated algorithms that can capture non-linear relationships in the data once sufficient samples are available.
Table: Research Reagent Solutions for Sperm Epigenetic Studies
| Reagent/Resource | Function in Sperm Epigenetic Research | Implementation Example |
|---|---|---|
| Illumina Infinium MethylationEPIC BeadChip | Genome-wide DNA methylation profiling | Analysis of ~850,000 CpG sites in sperm DNA [1] |
| Whole-Genome Bisulfite Sequencing (WGBS) | Comprehensive methylome analysis at single-base resolution | Longitudinal study of sperm methylome changes using T2T-CHM13 reference genome [33] |
| TCEP (tris(2-carboxyethyl)phosphine) | Reducing agent for sperm DNA extraction | Efficient protamine removal during DNA purification for methylation analysis [1] |
| NanoSeq Technology | Ultra-accurate DNA sequencing for mutation detection | Identification of age-related mutation patterns in sperm [11] |
Problem: Model fails to generalize to new populations or clinical cohorts
Diagnosis: The training data may lack sufficient diversity or contain population-specific biases.
Solution Strategy:
- Implement Cohort-Stratified Sampling: Ensure proportional representation across different populations in the training set. The 2024 SEA study validated findings across both a general population cohort (LIFE) and a fertility clinic cohort (SEEDS) [1].
- Apply Advanced Normalization Techniques: Use methods like Functional Normalization (preprocessFunnorm) to remove technical variation while preserving biological signals [36].
- Create Ensemble Models: Develop separate models for distinct subpopulations when consistent demographic or clinical factors affect epigenetic aging patterns.
Problem: Hardware limitations prevent training on full dataset
Diagnosis: Computational constraints are forcing suboptimal data utilization.
Solution Strategy:
- Employ Strategic Subsampling: Research indicates that randomly selecting 30% of a large dataset can often retain 95% of the performance while dramatically reducing computational requirements [35].
- Utilize Distributed Computing Frameworks: Implement Spark or other distributed systems for large-scale epigenetic data processing.
- Leverage Approximated Hyperparameter Search: Conduct initial hyperparameter optimization on data subsets to identify promising configurations before full training [35].
Experimental Protocols: Methodologies for Data Optimization
Protocol: Determining optimal training set size for sperm epigenetic clocks
Background: Systematically evaluate the relationship between sample size and prediction accuracy to allocate resources efficiently.
Workflow:
- Begin with a master dataset of sperm methylation samples with chronological age annotations.
- Generate progressively larger random subsets (e.g., 10%, 25%, 50%, 75%, 100%).
- Train identical model architectures on each subset using cross-validation.
- Evaluate performance on a fixed, independent test set.
- Identify the inflection point where additional samples yield negligible improvement.
Protocol: Cross-validation strategy for limited sperm methylation data
Background: Maximize model evaluation robustness when total samples are constrained.
Workflow:
- Perform stratified k-fold cross-validation (k=5 or 10) to maintain age distribution across folds.
- Implement nested cross-validation for hyperparameter tuning to prevent overfitting.
- Calculate performance metrics (RMSE, MAD, R²) for each fold and report mean ± standard deviation.
- Compare against a baseline model to establish practical significance of improvements.
The development of accurate sperm epigenetic clocks requires a strategic approach to training data collection that balances quantity with quality and relevance. While expanding training set size generally enhances prediction accuracy, researchers must consider the diminishing returns beyond certain thresholds and the critical importance of data quality and relevance. Future directions should focus on multi-center collaborations to assemble larger, more diverse sperm methylation datasets, development of efficient algorithms that maximize information extraction from limited samples, and integration of sperm-specific biological knowledge to guide feature selection. By applying these principles, researchers can build more robust epigenetic clocks that advance our understanding of male reproductive aging and its clinical implications.
Technical Troubleshooting Guides
Troubleshooting Guide: CpG Imputation from HM450 to EPIC Array
Problem: Low imputation accuracy when expanding coverage from HumanMethylation450 (HM450) to EPIC (HM850) BeadChip platforms.
| Problem Phenomenon | Potential Causes | Diagnostic Steps | Recommended Solutions |
|---|---|---|---|
| High Root-Mean-Square Error (RMSE) after imputation. | Inappropriate algorithm selection; tissue-specific methylation patterns not accounted for. | 1. Perform cross-validation within your specific tissue type (e.g., placenta, whole blood, semen).2. Check the correlation structure of neighboring CpG sites. | 1. Use the CUE (CpG impUtation Ensemble) framework, which combines multiple models.2. Ensure imputation is performed within the same tissue type, as patterns differ dramatically between tissues like blood and sperm [39] [15]. |
| Successful imputation rate below 85% (where success is defined as RMSE < 0.05 and accuracy > 95%). | Weak correlation between HM450 probes and target HM850-only CpGs; suboptimal model parameters. | 1. Filter out HM850-only CpG sites located far from any HM450 probes.2. Check the pre-trained model was built for your tissue of interest. | 1. Leverage a pre-trained CUE model from a relevant tissue. Pre-trained models for placenta and whole blood are available [39].2. For semen-specific studies, use models trained on sperm methylome data, as it differs significantly from somatic cells [15] [7]. |
| Model fails to converge or produces nonsensical values. | Singularity in the predictor matrix due to high dimensionality (p >> n). | Check the rank of the predictor matrix; it is likely less than the number of features (p). | Switch to penalized regression methods (Ridge, Lasso) via the glmnet package in R, which adds a penalty term to the estimating function to make the matrix invertible [40]. |
Troubleshooting Guide: Penalized Regression for High-Dimensional Methylation Data
Problem: Poor performance or instability when applying regression models for CpG selection and age prediction.
| Problem Phenomenon | Potential Causes | Diagnostic Steps | Recommended Solutions |
|---|---|---|---|
| Inability to compute coefficient estimates using ordinary least squares (OLS). | The (X^T * X) matrix is singular and not invertible because the number of CpG sites (p) exceeds the number of samples (n). |
Use the rankMatrix(X) function in R to confirm the rank is less than p. |
Use Ridge Regression, which solves β = (X^T * X + λ * I)^-1 * X^T * Y. The λ penalty makes the matrix full rank [40]. |
| Model does not generalize to independent test sets (overfitting). | The model is too complex and has learned noise from the training data. | Compare performance metrics (e.g., RMSE, MAE) between training and validation sets. | 1. Implement k-fold cross-validation (e.g., 10-fold) to find the optimal penalty parameter λ.2. Use the Lasso (Least Absolute Shrinkage and Selection Operator) to automatically perform feature selection by driving some coefficients to zero [40]. |
| Difficulty in interpreting the final model with thousands of CpGs. | The model includes a very large number of features with non-zero coefficients. | Examine the coefficient profile plot from a Lasso regression to see how the number of features changes with λ. |
1. For a more sparse model, use Lasso regression by setting alpha = 1 in the glmnet() function [40].2. For a compromise between Ridge and Lasso, use the Elastic Net (alpha between 0 and 1), which is useful when features are correlated [40]. |
Frequently Asked Questions (FAQs)
Q1: What is the most accurate method for imputing missing CpG methylation values from an HM450 to an EPIC array?
A: Based on cross-validation studies, an ensemble approach is most accurate. The CpG impUtation Ensemble (CUE) framework, which leverages multiple machine learning and statistical methods (KNN, logistic regression, penalized functional regression, random forest, XGBoost), has been shown to achieve the lowest RMSE and highest accuracy (e.g., 99.97% in one cohort) compared to any single method [39]. This ensemble is particularly valuable for increasing the coverage of the epigenomic landscape in existing HM450 datasets.
Q2: Why is my epigenetic age prediction model performing poorly in semen samples when it works well in blood?
A: Sperm cells exhibit very different age-related DNA methylation (DNAm) patterns compared to somatic cells. In sperm, DNAm often decreases with age in most genes, contrary to patterns in blood [15] [7]. Furthermore, the CpG sites most predictive of age in blood (e.g., in genes like ELOVL2) are often not predictive in sperm. Therefore, it is crucial to use semen-specific age-related CpG (AR-CpG) sites and prediction models trained exclusively on semen data [15] [7] [8].
Q3: How do I choose between Ridge, Lasso, and Elastic Net regression for my CpG selection problem?
A: The choice depends on your goal and the structure of your data.
- Ridge Regression (
alpha = 0): Use when you want to retain all features but shrink their coefficients. It is useful when you believe many CpG sites have a small but non-zero effect on the outcome [40]. - Lasso Regression (
alpha = 1): Use when you want a sparse model—that is, you want to select a small number of the most important CpG sites and set the coefficients of others to zero. This greatly aids interpretability [40]. - Elastic Net (
0 < alpha < 1): Use when you have many highly correlated CpG sites (e.g., sites located close to each other on the genome). Lasso might arbitrarily select one from a group, while Elastic Net can select groups of correlated features [40].
Q4: What is a realistic performance expectation for a sperm epigenetic clock model?
A: Performance varies based on the number and quality of CpGs and the modeling technique. Recent studies using genome-wide discovery and robust validation report:
- A 9-CpG Random Forest model achieved a Mean Absolute Error (MAE) of ~3.30 years [7].
- A 6-CpG linear model achieved an MAE of ~5.1 years [15].
- Earlier models with 3 CpGs reported MAEs of ~4.2-5.4 years [7]. These figures can serve as benchmarks for your own experiments.
Experimental Protocols & Workflows
Detailed Protocol: CUE Ensemble Imputation
This protocol is adapted from the CUE study for imputing HM850-only CpG sites using existing HM450 data [39].
1. Input Data Preparation:
- Format your HM450 beta-value matrix (samples x probes).
- Obtain a pre-trained CUE model or a reference dataset where the same samples have been profiled on both HM450 and HM850 arrays.
- The reference dataset should be from the same tissue type (e.g., whole blood, placenta, semen).
2. Model Training (If creating a new model):
- On the reference dataset, use the HM450 data as the predictor (X) and the HM850-only CpG values as the outcome (Y) for each target CpG.
- For each of the 339,014+ HM850-only CpGs, train the following five models within a cross-validation framework:
- k-Nearest Neighbours (KNN)
- Logistic Regression (with dichotomized beta values)
- Penalized Functional Regression (PFR)
- Random Forest (RF)
- XGBoost
- The CUE framework then ensembles the predictions from these models to produce a single, more accurate imputation.
3. Imputation and Quality Control:
- Apply the trained CUE model to your HM450 dataset to generate imputed values for all HM850-only CpGs.
- Filter out low-quality imputations. A recommended success metric is RMSE < 0.05 and accuracy > 95%, as calculated on a hold-out validation set [39]. In the original study, this successfully imputed 85.4% of target CpGs.
Detailed Protocol: Building a Sperm Epigenetic Clock with Penalized Regression
This protocol is based on recent studies that built accurate age prediction models for semen [7].
1. Genome-Wide Discovery of AR-CpGs:
- Use a comprehensive technique like double-enzyme Reduced Representation Bisulfite Sequencing (dRRBS) on semen samples from a discovery cohort (e.g., n=21) spanning a wide age range. This allows identification of AR-CpGs beyond the coverage of commercial arrays.
2. Targeted Validation:
- Design multiplex PCR panels for Bisulfite Amplicon Sequencing (BSAS). Include:
- Top AR-CpGs from your dRRBS discovery.
- Previously reported semen AR-CpGs (e.g., cg06304190, cg06979108, cg12837463) [7].
- Neighboring CpGs around the top hits, as they may also be predictive.
- Sequence a larger, independent validation cohort (e.g., n=125-247).
3. Model Building and Validation:
- Use a repeated nested cross-validation framework (e.g., 10-fold outer CV with 10-fold inner CV, repeated 10 times) to avoid overfitting.
- Train multiple algorithms, including:
- Multiple Linear Regression
- Random Forest (RF): Often shows superior accuracy for this task [7].
- Compare models based on Mean Absolute Error (MAE) and R-squared on the test sets.
Signaling Pathways & Workflow Visualizations
CUE Ensemble Imputation Workflow
CUE Ensemble Imputation Workflow: This diagram illustrates the process of using the CUE framework to impute missing HM850-only CpG sites from existing HM450 data, culminating in quality control checks.
Sperm Epigenetic Clock Development
Sperm Epigenetic Clock Development: This workflow outlines the key phases in creating a robust sperm epigenetic clock, from genome-wide discovery of age-related CpGs to model validation.
Research Reagent Solutions
Essential materials and computational tools used in the featured experiments and field.
| Item Name | Function / Application in Research | Specific Examples / Notes | ||
|---|---|---|---|---|
| Illumina BeadChip Arrays | Genome-wide DNA methylation profiling. | HumanMethylation450 (HM450): Covers ~485,000 probes. MethylationEPIC (EPIC/HM850): Covers ~850,000 probes. EPIC provides much more comprehensive coverage outside CpG islands [39]. | ||
| Bisulfite Amplicon Sequencing (BSAS) | Targeted, high-depth validation of candidate age-related CpG sites. | Used for robust, multiplex validation of dozens to hundreds of CpGs in large sample cohorts (e.g., n=247) [7]. | ||
| double-enzyme Reduced Representation Bisulfite Sequencing (dRRBS) | Cost-effective, genome-wide discovery of novel CpG sites beyond array coverage. | Identified >4 million CpG sites per sample in semen; revealed that >95% of shared CpGs were not on conventional arrays [7]. | ||
| CUE (CpG impUtation Ensemble) | R-based tool for imputing HM850-only CpG sites from HM450 data. | Pre-trained models for placenta and whole blood are available on GitHub: GangLiTarheel/CUE [39] [41]. |
||
glmnet R Package |
Fitting penalized regression models (Lasso, Ridge, Elastic Net). | Essential for dealing with high-dimensional data where the number of CpGs (p) exceeds samples (n). Used for feature selection and model regularization [40]. | ||
| Semen-Specific AR-CpG Database | A reference of pre-validated age-related CpG sites for sperm. | Provides a starting point for model building. Recent studies have compiled databases of 71+ AR-CpGs with | rho | > 0.50 [7]. |
Core Concepts and FAQs
FAQ 1: What is the fundamental difference between a first-generation and a second-generation epigenetic clock?
First-generation clocks, such as the Horvath and Hannum clocks, are predictive models trained using DNA methylation (DNAm) patterns that correlate strongly with an individual's chronological age. Their primary output is an estimate of chronological age [42] [43]. Second-generation clocks, such as PhenoAge and GrimAge, are trained to predict biological age or mortality risk by correlating DNAm patterns with clinical biomarkers, physical performance measures, or time-to-pregnancy (in the context of sperm). They are more powerful for predicting functional decline, age-related diseases, and other phenotypic outcomes [42] [8] [43].
FAQ 2: Why is developing sperm-specific epigenetic clocks particularly challenging?
Sperm cells exhibit very different patterns of age-related DNA methylation compared to somatic cells. While global DNA methylation decreases with age in many somatic tissues, sperm DNAm shows distinct, tissue-specific patterns of age-related change [15]. Furthermore, chronological age does not fully capture the biological aging of sperm, as intrinsic and extrinsic factors can cause sperm epigenetic age (SEA) to deviate from chronological age [8].
FAQ 3: My sperm epigenetic age (SEA) assessment shows acceleration. What does this mean for my research on male fertility?
Emerging evidence suggests that an advanced SEA is positively associated with the time taken for a couple to achieve pregnancy [8]. Crucially, SEA may not be associated with standard semen parameters like concentration or motility. Instead, it is significantly associated with more subtle defects in sperm head morphology (e.g., higher sperm head length and perimeter, presence of pyriform and tapered sperm, and a lower elongation factor) [8]. This indicates that SEA could be an independent biomarker of sperm quality and male fecundity that captures information beyond routine clinical assessments.
FAQ 4: Can an epigenetic clock be misled by cellular composition changes in a sample?
Yes, this is a critical technical consideration. Many epigenetic clocks are trained on bulk tissues, whose cellular composition changes with age. For example, in blood, the frequency of naïve CD8+ T cells decreases with age, while effector memory cells increase. Naïve T cells can exhibit an epigenetic age 15-20 years younger than effector memory T cells from the same individual. Therefore, a clock can be confounded by shifts in cell populations rather than purely measuring cell-intrinsic aging [16]. Using homogeneous cell populations or developing composition-resistant clocks like the IntrinClock is essential for precise measurement [16].
Troubleshooting Guide for Sperm Epigenetic Clock Development
Table 1: Common Experimental Challenges and Solutions
| Challenge | Potential Cause | Solution / Verification Step |
|---|---|---|
| Weak or No Correlation with Age | • Incorrect CpG marker selection• Somatic cell contamination | • Validate novel, sperm-specific DMSs (e.g., in SH2B2, EXOC3, IFITM2, GALR2, FOLH1B) [15]• Check for somatic contamination via DLK1 and H19 methylation analysis [8] |
| High Prediction Error (MAE) | • Suboptimal prediction model• Limited number of predictive CpGs | • Test various machine learning models (linear regression, elastic net)• Increase the number of age-correlated DMSs analyzed; a 6-CpG model achieved MAE=5.1 years, but more CpGs can improve accuracy [15] |
| Inconsistent Results Across Replicates | • Technical variation in DNA methylation measurement• Inconsistent sperm processing | • Use a consistent, reduced-bias DNA extraction protocol with a stable reducing agent like TCEP [8]• Standardize semen processing (e.g., density gradient centrifugation steps) across all samples [8] |
| Clock fails to predict phenotypic outcomes | • Clock may be capturing random drift or non-causal changes | • Focus on constructing clocks from methylation changes with a likely biological function, distinguishing between changes that cause damage (Type 1) and those that represent repair responses (Type 2) [44] |
| Poor Performance on Forensic Samples | • Low quantity/quality of input DNA• Inefficient bisulfite conversion | • Employ targeted MPS technologies for high-sensitivity analysis [15]• Implement strict quality control checks for bisulfite conversion efficiency [15] |
Table 2: Key Reagent Solutions for Sperm Epigenetics Research
| Research Reagent | Function in Experiment |
|---|---|
| Illumina Infinium MethylationEPIC BeadChip | Epigenome-wide discovery of age-correlated differentially methylated sites (DMSs) [8] [15]. |
| Tris(2-carboxyethyl)phosphine (TCEP) | A stable, room-temperature reducing agent used in sperm DNA lysis buffer to break down protamine-based packaging for efficient DNA purification [8]. |
| DNA Methylation Inhibitors (e.g., 5-aza-2'-deoxycytidine) | Tool compounds for functional validation of clock CpGs to test causality in aging pathways. |
| Targeted Bisulfite MPS Panels | Validating and quantifying DNAm levels at specific candidate CpG loci with high sensitivity, suitable for low-quality/quantity DNA [15]. |
| Positive Control Samples | Semen samples from donors of verified, diverse ages used to calibrate and validate prediction models [15]. |
Detailed Experimental Workflows
Workflow 1: Developing a Novel Sperm-Specific Epigenetic Clock
The following diagram outlines the key stages for building a predictive model for sperm biological age.
Diagram: Sperm Clock Development Workflow
Protocol Details:
- Cohort Selection: Assemble a cohort of male donors spanning a wide chronological age range. For the LIFE study, 379 men were included from a non-clinical population, while SEEDS included 192 men from a fertility clinic [8].
- Semen Sample Collection and Processing: Collect semen samples after a recommended period of ejaculatory abstinence. Process samples using density gradient centrifugation (e.g., one-step 50% gradient or two-step 40%/80% gradient) to isolate sperm [8].
- Sperm DNA Isolation: Extract DNA using a protocol designed for sperm chromatin. A recommended method involves homogenizing sperm in a lysis buffer containing guanidine thiocyanate and 50 mM TCEP, followed by purification on silica-based columns. This method avoids lengthy proteinase K digestions and works efficiently at room temperature [8].
- DNA Methylation Profiling: Analyze bisulfite-converted DNA using the Illumina Infinium MethylationEPIC BeadChip array, which covers over 850,000 CpG sites [8] [15].
- Bioinformatic Analysis:
- Preprocessing: Perform normalization, batch effect correction, and remove cross-hybridized or low-quality probes [8].
- DMS Discovery: Conduct correlation analysis to identify CpG sites whose methylation levels are significantly associated with chronological age. In one study, this identified 14,916 significant age-correlated DMSs [15].
- Predictive Model Building:
- Divide the cohort into a training set (e.g., 80%) and a validation set (20%) [42].
- Using the training set, train a machine learning model (e.g., generalized linear model with cross-validation) to predict age based on the methylation levels of the most significantly age-correlated DMSs. A model based on 6 CpGs from genes like SH2B2, EXOC3, and FOLH1B has been shown to predict age with a mean absolute error (MAE) of 5.1 years [15].
- Validation: Apply the final model to the held-out test set and independent cohorts to evaluate its prediction accuracy (e.g., correlation and MAE between predicted and chronological age) [42] [8].
Workflow 2: Troubleshooting a Failed Age Prediction Experiment
This workflow provides a logical sequence for diagnosing problems when experimental results are unexpected.
Diagram: Troubleshooting Logic Flow
Protocol Details:
- Repeat the Experiment: Before extensive troubleshooting, simply repeat the experiment to rule out simple human error, such as pipetting mistakes or incorrect sample labeling [45].
- Verify Experimental and Biological Controls:
- Technical Controls: Ensure that positive control samples (of known age and methylation profile) are included in the run and yield the expected results.
- Biological Controls: Confirm that your sample purity is high. Check for somatic cell contamination by analyzing imprinting control regions like DLK1 and H19 [8].
- Check Reagents and Equipment:
- Reagent Integrity: Molecular biology reagents can degrade. Confirm that all reagents, especially bisulfite conversion kits and enzymes, have been stored correctly and are not past their expiration dates [45].
- Equipment Calibration: Ensure that instrumentation, such as the scanner for methylation arrays or the sequencer for MPS, is properly calibrated and maintained.
- Systematically Change Variables: If the problem persists, identify and test key variables one at a time [46] [45].
- DNA Input Quality/Quantity: Test a range of DNA input amounts and assess DNA quality (e.g., via Bioanalyzer).
- Bisulfite Conversion Efficiency: Check the efficiency of the bisulfite conversion reaction, as incomplete conversion is a major source of error [15].
- CpG Panel: If using a targeted approach, verify that the selected CpG markers are performing well. Consider expanding the panel or re-optimizing the assay conditions for problematic loci.
- Document Everything: Meticulously record all steps, observations, and changes made during troubleshooting in a lab notebook. This is critical for identifying the root cause and ensuring reproducibility [45].
Overcoming Analytical Hurdles: Confounders, Standardization, and Technical Noise
Sperm epigenetic clocks are powerful tools for assessing male fertility and biological aging by measuring DNA methylation patterns in sperm. However, the accuracy of these clocks can be significantly compromised by technical and biological confounders. This guide provides targeted troubleshooting advice to help researchers identify, mitigate, and correct for the critical issues of cellular composition, batch effects, and donor biology in their experiments.
FAQs: Resolving Common Experimental Challenges
Q1: Our sperm epigenetic age (SEA) predictions vary significantly between different processing batches. How can we identify and correct for this?
Batch effects arise from technical variations between different experimental runs, laboratories, or operators. To address this:
- Prevention: Implement careful sample randomization across sequencing arrays or processing batches to minimize systematic technical bias [8].
- Detection: Use exploratory data analysis, such as Principal Component Analysis (PCA), to check if samples cluster by batch rather than by biological variables of interest.
- Correction: Employ batch correction methods designed for high-dimensional data. The Mutual Nearest Neighbors (MNN) method is particularly effective as it does not assume identical cell population composition across batches. It identifies pairs of cells from different batches that are most similar (mutual nearest neighbors) and uses them to estimate the batch effect, which is then subtracted from the data [47]. For complex integrations, newer methods like sysVI, a conditional variational autoencoder (cVAE) using VampPrior and cycle-consistency, show improved performance in integrating datasets with substantial batch effects, such as those from different protocols or systems [48].
Q2: We suspect non-sperm cells in our semen samples are contaminating our epigenetic analysis. How can we confirm and address this?
Somatic cell contamination is a critical confounder, as epigenetic clocks are highly cell-type-specific.
- Confirmation: Analyze methylation levels at imprinted control regions, such as the DLK1 and H19 loci, which serve as a quality control measure to confirm minimal somatic cell contamination in your sperm DNA samples [8].
- Mitigation: Ensure your sperm processing protocol includes robust somatic cell lysis or density gradient centrifugation steps designed to isolate a pure sperm cell population before DNA extraction [8].
Q3: Our epigenetic clock performs well in our primary cohort but fails to generalize to an independent cohort from a different study. What could be the cause?
This often results from unaccounted-for batch effects or differences in donor biology between cohorts.
- Cross-Study Batch Effects: Differences in sample collection, processing, and sequencing between studies can introduce large batch effects. Frameworks like SCCAF-D are specifically designed for this "cross-reference" setting. It integrates multiple datasets and selects a 'self-consistent' subset of cells to create an optimized reference, improving deconvolution and generalization accuracy [49].
- Biological Differences: The cohorts may differ in key biological variables. Always measure and adjust for critical covariates like donor age, BMI, and smoking status in your association models, as these factors are known to correlate with sperm epigenetic aging [2] [8].
Troubleshooting Guides
Issue 1: Inconsistent Results Across Replicates or Batches
Problem: High variability in SEA estimates when the same sample is processed in different batches or by different technicians.
Step-by-Step Resolution:
- Repeat the Experiment: Unless cost or time-prohibitive, first repeat the experiment to rule out a simple one-off error [45].
- Check Controls: Verify that positive and negative controls are performing as expected across all batches [45].
- Audit Reagents and Equipment: Check that all reagents have been stored correctly and have not expired. Visually inspect solutions for precipitates or cloudiness [45].
- Systematically Change Variables: If the problem persists, generate a list of variables that could differ between batches (e.g., bisulfite conversion efficiency, array hybridization time, technician). Change only one variable at a time to isolate the root cause [45].
- Apply Computational Correction: Once the source is identified, apply a suitable batch effect correction algorithm like MNN [47] or sysVI [48] to the finalized data.
Issue 2: Weak or No Association Between SEA and Clinical Outcomes
Problem: Your sperm epigenetic clock shows poor predictive power for reproductive outcomes like time-to-pregnancy (TTP).
Step-by-Step Resolution:
- Verify the Experiment: Revisit the scientific literature. A weak association with standard semen parameters (count, concentration, motility) is possible, as SEA may be more strongly linked to sperm head morphology (e.g., head length, perimeter, presence of pyriform/tapered shapes) [8]. Ensure you are investigating the correct phenotypic endpoints.
- Assess Covariates: Critically evaluate your statistical models. Are you adjusting for all known confounders? Failing to adjust for male smoking status, which is associated with advanced SEA, can obscure true biological relationships [2] [8].
- Investigate Cellular Composition: Rule out somatic cell contamination by checking control loci like DLK1 and H19 [8].
- Refine the Clock Model: The choice of epigenetic markers matters. Clocks based on individual CpGs (SEA~CpG~) may show stronger associations with TTP and gestational age than those based on differentially methylated regions (DMRs) [2]. Ensure you are using the most predictive marker set for your specific research question.
Data and Methodology Tables
Table 1: Key Sperm Epigenetic Age Associations from Clinical and Population Cohorts
| Cohort Type | Association with SEA | Effect Size / Summary | P-value | Citation |
|---|---|---|---|---|
| General Population (LIFE Study) | Time-to-Pregnancy (TTP) | FOR=0.83 (17% lower pregnancy probability per unit SEA increase) | 1.2×10⁻⁵ | [2] |
| General Population (LIFE Study) | Gestational Age | -2.13 days | 0.007 | [2] |
| General Population (LIFE Study) | Sperm Head Morphology | Associated with head length, perimeter, pyriform/tapered shapes | < 0.05 | [8] |
| Clinical (SEEDS - IVF) | Standard Semen Parameters | No significant associations found | > 0.05 | [8] |
Table 2: Comparison of Batch Effect Correction Methods for Genomic Data
| Method | Primary Principle | Key Advantage | Use Case |
|---|---|---|---|
| Mutual Nearest Neighbours (MNN) [47] | Identifies most similar cells across batches to estimate technical noise. | Does not require identical population composition across batches. | Correcting technical batch effects in scRNA-seq or methylation data. |
| SCCAF-D [49] | Integrates datasets and selects a 'self-consistent' reference via machine learning. | Achieves stable accuracy (PCC >0.75) in cross-reference settings. | Deconvolving bulk data or integrating single-cell references from different studies. |
| sysVI (VAMP + CYC) [48] | Conditional VAE with VampPrior and cycle-consistency constraints. | Improves integration of substantial batch effects (e.g., cross-species) while preserving biology. | Integrating datasets with strong technical/biological confounders (e.g., different protocols, species). |
Experimental Protocols
Protocol 1: Sperm DNA Extraction for Methylation Analysis
Objective: To isolate high-quality, contaminant-free DNA from semen samples for downstream epigenetic profiling.
Materials:
- Lysis buffer with guanidine thiocyanate
- Tris(2-carboxyethyl)phosphine (TCEP), a stable reducing agent
- 0.2 mm steel beads
- Silica-based spin columns (e.g., from Qiagen)
- Microcentrifuge
Method:
- Sperm Isolation: Isolate sperm from crude semen using a density gradient centrifugation step (e.g., 50% or 40%/80% gradients) to remove somatic cells and debris [8].
- Lysis and Reduction: Homogenize the purified sperm pellet with steel beads in a lysis buffer containing guanidine thiocyanate and 50 mM TCEP. Incubate at room temperature for 5 minutes. TCEP is critical for breaking protamine disulfide bonds and efficiently releasing sperm DNA [8].
- DNA Purification: Purify the DNA using a commercially available silica-based spin column, following the manufacturer's protocol. This method foregoes lengthy proteinase K digestions and works efficiently at room temperature [8].
- Quality Control: Quantify DNA and confirm purity via spectrophotometry. Assess bisulfite conversion efficiency and check for somatic cell contamination by analyzing methylation at imprinted loci like DLK1 and H19 [8].
Protocol 2: Validating Clock Performance in an Independent Cohort
Objective: To test the generalizability and clinical relevance of a sperm epigenetic clock.
Materials:
- Independent cohort with semen samples and phenotypic data (e.g., time-to-pregnancy, semen parameters).
- DNA methylation profiling platform (e.g., EPIC array, targeted bisulfite sequencing).
Method:
- Cohort Selection: Select an independent validation cohort. Ideal cohorts include both a general population group (e.g., couples trying to conceive, like the LIFE Study) and a clinical group (e.g., couples undergoing IVF, like the SEEDS study) [2] [8].
- Data Generation: Process the independent cohort's samples using your established sperm epigenetic clock protocol.
- Statistical Analysis:
- Evaluate the clock's basic performance by calculating the correlation (r) between predicted epigenetic age and chronological age. High-performance clocks show correlations of r > 0.90 [2].
- Use multivariable regression models to assess the association between SEA and reproductive outcomes. Crucially, adjust for covariates like male age, BMI, and smoking status [2] [8].
- Test associations with both standard semen parameters and, if available, detailed sperm morphology measures [8].
The Scientist's Toolkit
Table 3: Essential Research Reagents and Materials for Sperm Epigenetic Clock Development
| Item | Function / Application | Technical Notes |
|---|---|---|
| Infinium MethylationEPIC BeadChip | Genome-wide DNA methylation profiling. | Covers over 850,000 CpG sites. Standard for discovery phase [15] [8]. |
| Targeted Bisulfite MPS | Validating and applying clock markers. | More suitable for low-quality/quantity forensic DNA or for focused analysis of specific CpGs [15]. |
| Tris(2-carboxyethyl)phosphine (TCEP) | Reducing agent in sperm DNA extraction. | Preferable over DTT as it is stable at room temperature and efficiently breaks protamine bonds [8]. |
| SH2B2, EXOC3, IFITM2, GALR2, FOLH1B CpG Panels | Core markers for age prediction. | A 6-CpG model from these genes can predict age with a MAE of ~5.1 years [15]. |
Workflow and Relationship Diagrams
Diagram 1: A workflow for identifying and mitigating critical confounders in sperm epigenetic clock research.
Diagram 2: A logical troubleshooting guide for resolving weak associations in SEA analysis.
Frequently Asked Questions (FAQs)
1. What is Sperm Epigenetic Aging (SEA) and why is it important for male fertility research? Sperm Epigenetic Aging (SEA) refers to the biological age of sperm cells, estimated using epigenetic clocks based on DNA methylation patterns. Unlike chronological age, SEA can be accelerated or decelerated by various environmental and lifestyle factors. It is a crucial biomarker because an advanced SEA is associated with a 17% lower cumulative probability of pregnancy within 12 months and a longer time-to-pregnancy for couples, independent of the female partner's age [2]. This makes it a valuable metric for assessing male reproductive health and the impact of environmental exposures.
2. What is the critical window during which environmental exposures can affect the sperm epigenome? The process of spermatogenesis—the creation of mature sperm—takes approximately 74 days. This constitutes a critical window during which environmental exposures can significantly influence the final epigenetic patterns in sperm. Therefore, for optimal reproductive outcomes, men should focus on reducing harmful exposures for a minimum of three months prior to conception [50].
3. Which environmental exposures are most strongly linked to alterations in SEA? Research has consistently identified several key accelerants of SEA:
- Smoking/Tobacco: Significantly alters sperm DNA methylation patterns and is associated with advanced SEA [51] [2] [52].
- Toxicants: Exposure to endocrine-disrupting chemicals (EDCs) like phthalates and THC (the main psychoactive component in cannabis) is linked to disrupted sperm DNA methylation, particularly at genes important for neurodevelopment [50] [53] [54].
- Paternal Stress: Chronic stress in fathers is associated with epigenetic changes in sperm and an increased risk of metabolic dysfunction and stress sensitivity in offspring [53].
- Paternal Diet and Obesity: These factors are associated with epigenetic alterations in sperm that may increase the risk of metabolic disorders in the next generation [53].
4. My research shows altered global DNA methylation in sperm after nicotine exposure, but I am unsure how to validate its functional relevance. What are the next steps? Observing global changes is a starting point. The next step is to move from association to functional correlation. You should:
- Conduct Targeted Gene Expression Analysis: Correlate the methylation changes in specific genes of interest with their transcript levels in sperm, as methylation often regulates gene expression. For example, smoking has been shown to lead to the downregulation of genes like
PTPRN2andPGAM5, which are linked to sperm function [52]. - Investigate Offspring Outcomes: In model organisms, examine if the observed methylation changes are transmitted to the offspring and if they correlate with phenotypic outcomes, such as alterations in neurodevelopment or metabolic health [54].
- Assess Clinical Correlates: In human studies, correlate the specific methylation signatures with concrete clinical outcomes like fertilization rates, embryo quality, or time-to-pregnancy [2] [53].
Troubleshooting Guides
Issue: High Variability in SEA Measurements Within a Cohort
| Potential Cause | Diagnostic Steps | Recommended Solution |
|---|---|---|
| Inconsistent Sample Processing | Review protocols for somatic cell lysis and DNA extraction. Check methylation data for contamination using control loci (e.g., DLK1). |
Implement a standardized, stringent somatic cell lysis protocol [51] and use a column-based DNA extraction kit validated for sperm [51]. |
| Unaccounted Confounding Exposures | Administer detailed lifestyle questionnaires to participants (smoking, diet, occupation). Statistically control for these variables. | Use a comprehensive covariate model that includes smoking status, BMI, alcohol consumption, and age [2] [53]. |
| Technical Batch Effects | Perform Principal Component Analysis (PCA) on methylation data to check for batch effects. | Include batch as a covariate in analysis. Process cases and controls simultaneously and use normalization techniques like SWAN [51]. |
Issue: Correlating a Specific Exposure (e.g., Phthalates) with a Specific Offspring Phenotype
| Experimental Challenge | Solution & Workflow |
|---|---|
| Establishing Paternal Causality | 1. Controlled Animal Models: Expose only the male to the toxicant before mating with a naive female. This isolates the paternal contribution [50] [54].2. Human Cohort Studies: In prospective pregnancy cohorts, collect detailed paternal exposure data and sperm samples prior to conception [2]. |
| Identifying the Molecular Vector | Analyze multiple components of the sperm epigenome in the exposed father:• DNA Methylation: Use beadchip arrays (450K/EPIC) or RRBS [51] [2] [54].• sncRNA: Sequence sncRNAs from sperm and seminal plasma extracellular vesicles [50] [53]. |
| Linking Sperm Signature to Offspring Health | 1. Track Epigenetic Inheritance: Assess whether sperm DNA methylation or sncRNA changes are also present in offspring tissues [53].2. Functional Studies: Use techniques like zygotic microinjection of sperm sncRNAs from exposed males into control embryos to test for phenotype recapitulation. |
Table 1: Impact of Smoking on Sperm DNA Methylation and Gene Expression
| Metric | Exposed Group (Heavy Smokers) | Control Group (Non-Smokers) | P-value & Notes | Source |
|---|---|---|---|---|
| Differentially Methylated CpGs | 141 significant CpGs | Baseline | Genome-wide analysis | [51] |
| Methylation Variance | Increased genome-wide variance | Lower variance | Suggests stochastic epigenetic changes | [51] |
PGAM5 Expression |
Significant downregulation | Normal expression | p ≤ 0.03; associated with reduced motility, count | [52] |
PTPRN2 Expression |
Significant downregulation | Normal expression | p ≤ 0.01; associated with reduced normal form, vitality | [52] |
Table 2: Sperm Epigenetic Clock (SEA) Performance and Correlations
| Clock / Metric | Correlation with Chronological Age | Key Clinical Correlation | Source |
|---|---|---|---|
| SEACpG Clock | r = 0.91 | FOR for TTP = 0.83 (17% lower pregnancy probability per cycle) | [2] |
| SEA Acceleration (Smoking) | N/A | Current smokers displayed advanced SEACpG (P < 0.05) | [2] |
| SEA & Gestational Age | N/A | Advanced SEACpG associated with -2.13 days gestation (P = 0.007) | [2] |
Key Experimental Protocols
Protocol 1: Genome-Wide Sperm DNA Methylation Analysis Using BeadChip Arrays
This protocol is essential for constructing epigenetic clocks and identifying exposure-specific signatures [51] [2].
- Sperm Collection and Purification: Collect semen sample after 2-7 days of abstinence. Purify using a discontinuous density gradient.
- Somatic Cell Lysis: Incubate sperm pellet in somatic cell lysis buffer (0.1% SDS, 0.5% Triton X-100) for ≥60 minutes at 4°C to remove leukocyte contamination. Verify purity microscopically.
- DNA Isolation: Extract DNA using a column-based kit (e.g., DNeasy Kit, Qiagen) with sperm-specific modifications.
- Bisulfite Conversion: Treat DNA using a commercial bisulfite conversion kit (e.g., Zymo DNA Methylation-Gold Kit).
- Array Processing: Hybridize converted DNA to Infinium HumanMethylation450K or EPIC BeadChip per manufacturer's protocol.
- Data Processing & Normalization: Use pipelines like ChAMP to process raw data, perform quality control, filter probes, and apply normalization (e.g., SWAN). Logit-transform β-values to M-values for statistical analysis.
Protocol 2: Validating Exposure Effects in a Controlled Animal Model
This protocol is crucial for establishing causality, as demonstrated in THC and nicotine studies [54].
- Animal Exposure:
- THC: Administer via daily subcutaneous injection (e.g., 4 mg/kg) or oral gavage to model different consumption routes. Use vehicle-injected controls.
- Nicotine: Adminerate via subcutaneous injection or drinking water at doses relevant to human exposure.
- Duration: Continue exposure for a full spermatogenic cycle (~50-60 days in rodents).
- Sperm Collection: Collect sperm from cauda epididymis post-euthanasia.
- Targeted Methylation Analysis:
- DNA Extraction & Bisulfite Conversion: As in Protocol 1.
- Pyrosequencing: Design assays around CpG sites of interest identified from prior sequencing. Perform PCR on bisulfite-converted DNA and analyze on a pyrosequencing system for quantitative, high-resolution methylation data.
Pathway and Workflow Visualizations
Mechanism of Environmental Impact on Sperm Epigenome
Sperm DNA Methylation Analysis Workflow
The Scientist's Toolkit: Essential Research Reagents & Materials
| Item | Function / Application in SEA Research | Key Considerations |
|---|---|---|
| Infinium Methylation EPIC BeadChip | Genome-wide DNA methylation profiling for epigenetic clock development and exposure signature discovery. | Covers >850,000 CpG sites. Ideal for large cohort studies. Requires high-quality bisulfite-converted DNA [51] [2]. |
| Somatic Cell Lysis Buffer (0.1% SDS, 0.5% Triton X-100) | Critical for removing leukocyte contamination from sperm samples, ensuring methylation profiles are sperm-specific. | Post-lysis visual inspection and validation with control loci (e.g., DLK1) are mandatory [51]. |
| Zymo DNA Methylation-Gold Kit | Bisulfite conversion of unmethylated cytosines to uracils, while methylated cytosines remain unchanged. | High conversion efficiency is crucial for accurate downstream quantification of methylation levels [51] [54]. |
| Pyrosequencing System | Targeted, quantitative validation of DNA methylation levels at specific CpG sites identified from genome-wide screens. | Provides high accuracy and reproducibility for a small number of loci. Essential for validating findings from array or RRBS data [54] [52]. |
| PureSperm Density Gradient | Purification of motile, morphologically normal spermatozoa from seminal plasma and other cells. | Standardizes the sperm population being analyzed, reducing noise in epigenetic data [52]. |
FAQs: Core Concepts and Mechanisms
Q1: What is the proposed mechanistic link between the blood-testis barrier (BTB) and sperm epigenetic aging? The BTB, the tightest blood-tissue barrier in the body, creates a unique biochemical environment for spermatogenesis. Recent research identifies the mTOR pathway in Sertoli cells as a critical regulator of both BTB integrity and the rate of sperm epigenetic aging. The balance between two mTOR complexes is key: mTORC1 promotes BTB disassembly, while mTORC2 promotes its integrity. Environmental stressors like heat shock and cadmium disrupt this balance, increasing BTB permeability and accelerating age-related changes in sperm DNA methylation, a process termed sperm epigenetic aging [55] [56].
Q2: How do environmental factors like heat stress and cadmium exposure exploit this mechanism? Environmental factors accelerate sperm epigenetic aging via distinct, BTB-centric pathways, as demonstrated in mouse models [55]:
- Heat Stress (mTOR-dependent): Acts through a mechanism that involves the activation of mTOR complexes, leading to BTB disruption.
- Cadmium (mTOR-independent): Disrupts BTB integrity through pathways not directly involving mTOR, yet still results in accelerated epigenetic aging. Both stressors ultimately cause similar detrimental changes to sperm DNA methylation, affecting genes involved in embryonic and neurodevelopment [55].
Q3: Why is a sperm-specific epigenetic clock necessary, and how accurate are current models? Sperm cells have a very different pattern of age-related DNA methylation compared to somatic cells. Clocks designed for blood or other tissues perform poorly on semen samples [15] [7]. Sperm-specific clocks are therefore essential for accurate age prediction in andrology and forensic science. The table below summarizes the performance of recently developed models.
Table 1: Performance of Recent Sperm Epigenetic Clocks
| Model Description | Number of CpG Sites | Technology Used | Reported Mean Absolute Error (MAE) | Citation |
|---|---|---|---|---|
| Random Forest Model | 9 CpGs | Bisulfite Amplicon Sequencing (BSAS) | 3.30 years | [7] |
| Linear Model | 6 CpGs | Targeted MPS | 5.1 years | [15] |
| Methylation SNaPshot | 3 CpGs | SNaPshot / Microarray | ~4.2 - 5.4 years | [7] |
Q4: Does advanced paternal age directly impact fertility and offspring health? Yes. Epidemiological and animal model evidence links advanced paternal age to:
- Reduced Fecundity: Delayed time to pregnancy and decreased likelihood of achieving pregnancy [3].
- Offspring Health Risks: Increased risk of neuropsychiatric disorders such as schizophrenia and autism, as well as early development of cancer [3] [56]. These outcomes are mediated, at least in part, by age-dependent alterations in the sperm epigenome [3].
Troubleshooting Guides
Issue: Inconsistent Sperm Epigenetic Age (SEA) Predictions
Potential Causes and Solutions:
- Cause: Somatic Cell Contamination.
- Solution: Implement rigorous sperm purification protocols using density gradient centrifugation to remove leukocytes and immature germ cells, which have distinct DNA methylation patterns [7].
- Cause: Suboptimal DNA Extraction from Sperm.
- Solution: Use a specialized sperm DNA extraction method that includes a reducing agent like Tris(2-carboxyethyl)phosphine (TCEP) to break down protamine-based packaging efficiently [8].
- Cause: Choice of Epigenetic Clock Model.
Issue: Modeling Environmental Stress in Animal Experiments
Potential Causes and Solutions:
- Cause: Uncontrolled Stressor Exposure.
- Solution: Use precise, validated exposure protocols. For heat stress, an acute intermittent whole-body protocol (e.g., 31.5°C or 34.5°C) can mimic human heat waves. For cadmium, intraperitoneal injection of CdCl2 (e.g., 2 mg/kg body weight) has been shown to be effective [55].
- Cause: Lack of Functional BTB Integrity Assessment.
- Solution: Complement epigenetic analyses with a direct BTB integrity assay. The biotin tracer assay is a gold-standard method where biotin is injected into the testis interstitium, and its diffusion into the tubule lumen is visualized, quantifying BTB permeability [56].
Experimental Protocols
Protocol: Assessing BTB Integrity via Biotin Tracer Assay
This protocol is adapted from established methods in mouse models [56].
Principle: A small, membrane-impermeable biotin tracer is injected into the testis interstitium. In a healthy, intact BTB, the tracer is confined to the interstitial space. A compromised BTB allows the tracer to penetrate the adluminal compartment of the seminiferous tubules.
Procedure:
- Anesthetize the experimental animal (e.g., mouse) and expose the testis.
- Micro-inject 5-10 µL of a EZ-Link Sulfo-NHS-LC-Biotin solution (10 mg/mL in PBS) into the testis interstitium.
- Allow diffusion for 30 minutes.
- Collect and fix the testis in 4% paraformaldehyde for 4-6 hours.
- Cryo-section the testis into 10-µm thick sections.
- Stain the sections with fluorescently conjugated Streptavidin (e.g., Streptavidin-FITC, 1:100 dilution) and a counterstain for Sertoli cell nuclei (e.g., DAPI).
- Image and Analyze using fluorescence microscopy. A functional BTB will show biotin signal only in the interstitial spaces. Leakage of the signal into the tubule lumen indicates BTB disruption.
Protocol: Genome-Wide Discovery of Sperm AR-CpGs Using dRRBS
This protocol is for identifying novel age-related CpG sites with greater coverage than microarray platforms [7].
Workflow:
Key Steps:
- Sample Collection & Stratification: Collect semen samples from donors across a wide age range (e.g., 22-67 years). Stratifying into young, middle-aged, and older groups can enhance discovery power [7].
- DNA Extraction: Use a standardized, high-yield method for sperm DNA.
- dRRBS Library Preparation: Digest genomic DNA with the restriction enzymes MspI and MseI. This combination enriches for CpG-rich regions and provides more comprehensive genome-wide coverage compared to microarrays.
- Bisulfite Conversion & Sequencing: Treat the library with bisulfite to convert unmethylated cytosines to uracils, followed by high-throughput sequencing.
- Bioinformatic Analysis:
- Align sequences to a reference genome.
- Calculate methylation levels at each CpG site.
- Perform differential methylation and correlation analyses to identify CpG sites whose methylation status is strongly correlated with donor age (|rho| > 0.50 is a common threshold).
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Research Reagents and Models
| Reagent / Model | Function/Description | Application in BTB/Epigenetic Aging Research |
|---|---|---|
| CdCl₂ (Cadmium Chloride) | Heavy metal salt, environmental toxicant. | Used to induce mTOR-independent BTB disruption and model environmental acceleration of epigenetic aging [55]. |
| AMH-Cre Transgenic Mice | Mouse model expressing Cre recombinase specifically in Sertoli cells. | Enables cell-type-specific knockout of genes (e.g., Rptor or Rictor) to study mTOR pathway function in BTB regulation [56]. |
| Rptor / Rictor KO Mice | Models with knocked-out components of mTORC1 (Rptor) or mTORC2 (Rictor). | Critical for establishing the causal role of mTOR balance in Sertoli cells on sperm epigenetic aging and rejuvenation [56]. |
| Sulfo-NHS-LC-Biotin | A membrane-impermeable, water-soluble biotinylation reagent. | The active tracer used in the biotin tracer assay for functional assessment of BTB integrity [56]. |
| Infinium MethylationEPIC BeadChip | Microarray for analyzing DNA methylation at >850,000 CpG sites. | A standard tool for epigenome-wide association studies and for constructing epigenetic clocks [15]. |
| TCEP (Tris(2-carboxyethyl)phosphine) | A stable, reducing agent. | Essential for efficiently breaking protamine disulfide bonds during DNA extraction from mature sperm [8]. |
Key Signaling Pathways and Workflows
The mTOR/BTB Signaling Axis in Sperm Epigenetic Aging
The following diagram summarizes the core mechanistic pathway linking environmental stress to sperm epigenetic aging via the BTB.
Core Concepts: Sperm Epigenetic Clocks and Standardization
Sperm Epigenetic Age (SEA) is an estimate of the biological age of sperm based on DNA methylation patterns, which can differ from chronological age. It is derived from epigenetic clocks, which are statistical models trained to predict age using DNA methylation data from specific genomic sites [57] [2]. Advanced SEA has been significantly associated with a 17% lower cumulative probability of pregnancy within 12 months and a longer time-to-pregnancy (TTP), underscoring its clinical relevance [2]. Furthermore, SEA shows associations with specific sperm morphological defects, such as abnormal head shape, even when standard semen parameters appear normal [8].
The accuracy of SEA is highly dependent on the robustness of the underlying data. Standardized protocols from sample collection to data analysis are critical to minimize technical noise and ensure that measurements reflect true biological signals rather than experimental artifacts. This is essential for developing reliable biomarkers for male fecundity [57] [8].
Frequently Asked Questions (FAQs)
Q1: Why is a sperm-specific epigenetic clock necessary? Can't I use clocks developed for somatic tissues? The DNA methylation loci used in somatic tissue epigenetic clocks have shown no predictive value in male germ cells [2]. Sperm has a unique epigenetic landscape, including regions of hypermethylation and hypomethylation that differ from somatic cells. Therefore, specialized clocks trained on sperm DNA methylation data are required for accurate biological age estimation in this cell type [2] [8].
Q2: My DNA yield from sperm is low. How does this impact downstream methylation analysis? Low DNA input can lead to non-specific binding during methylated DNA enrichment, potentially skewing your results [58]. It is crucial to follow protocols specifically optimized for low DNA input amounts. Always use the manufacturer’s guidelines for minimum input requirements and consider using DNA extraction methods designed for high efficiency with sperm cells [8].
Q3: What are the most critical steps to ensure reproducibility in my methylation array workflow? The three most critical steps are:
- Consistent Bisulfite Conversion: Use pure DNA and ensure complete reaction conditions to avoid incomplete conversion, which is a major source of bias [58].
- Rigorous Quality Control (QC): Implement a thorough QC pipeline to identify mislabeled, contaminated, or technically failed samples before analysis [59].
- Proper Normalization: Apply appropriate normalization methods (e.g., Subset-Quantile Normalization) to correct for technical variation between arrays [60].
Q4: I found a significant association with SEA. How can I be sure it's not due to cell contamination or sample mix-ups? You should perform the following quality checks using your raw methylation data:
- Sex Check: Compare the sex predicted by the methylation data (e.g., using X and Y chromosome probe intensities) with the recorded sex in your metadata to catch mislabeling [59].
- Contamination Check: Use probes that query high-frequency SNPs on the array to detect outliers that may indicate sample contamination with foreign DNA [59].
- Identity Check: Use these same SNP probes as a genetic fingerprint to identify sample duplicates or mix-ups [59].
Troubleshooting Guide
This guide addresses common problems encountered during the sperm methylation analysis workflow.
Table 1: Troubleshooting Common Experimental Issues
| Problem | Potential Cause | Solution | Preventive Measures |
|---|---|---|---|
| Poor amplification of bisulfite-converted DNA | Primers not optimally designed for converted template; DNA strand breaks from harsh bisulfite treatment; Uracil in template inhibiting polymerase [58]. | -Redesign primers to be 24-32 nt, with ≤3 mixed bases, and avoid mixed bases at the 3' end.-Use a hot-start Taq polymerase (not proof-reading).-Keep amplicon size around 200 bp [58]. | Use a well-established DNA extraction protocol that yields high-molecular-weight DNA and ensure bisulfite conversion reagents are fresh [8]. |
| Very little or no methylated DNA enriched | Low DNA input causing non-specific binding of MBD protein [58]. | Follow the low-DNA-input protocol as specified in the product manual [58]. | Quantify DNA accurately and use the recommended input range for your enrichment kit. |
| Incomplete bisulfite conversion | Particulate matter in DNA sample; impurities in DNA inhibiting reaction [58]. | Centrifuge DNA sample at high speed and use only the clear supernatant for conversion [58]. | Ensure DNA used for conversion is pure. Use quality assessment (e.g., Nanodrop, Qubit) before proceeding. |
| High failure rate or poor data quality from methylation arrays | Low-quality starting DNA; incomplete bisulfite conversion; failure of experimental steps in the Infinium assay [59]. | Evaluate 17 control metrics from the array's control probes to diagnose the specific failed step (e.g., staining, extension) [59]. | Implement pre-array QC to ensure DNA quality and complete bisulfite conversion. |
Detailed Experimental Protocols
Sperm DNA Extraction Protocol (Reducing Agent Method)
This protocol is optimized for sperm cells, which package DNA primarily with protamines instead of histones [8].
Key Reagents:
- Lysis Buffer: Contains guanidine thiocyanate.
- Reducing Agent: 50 mM Tris(2-carboxyethyl)phosphine (TCEP), stable at room temperature.
- Homogenization: 0.2 mm steel beads.
- Purification: Silica-based spin columns.
Procedure:
- Homogenize and Lyse: Homogenize sperm with steel beads in a lysis buffer containing guanidine thiocyanate and TCEP at room temperature for 5 minutes. The reducing agent is critical for breaking protamine disulfide bonds.
- Purify DNA: Transfer the lysate to a silica-based spin column and follow the manufacturer's protocol for DNA binding, washing, and elution. This method avoids lengthy proteinase K digestions and can be performed at room temperature, consistently yielding over 90% high-quality DNA [8].
Bisulfite Conversion and Methylation Array QC Protocol
Bisulfite Conversion:
- Use pure DNA for conversion. If particulate matter is present after adding conversion reagent, centrifuge at high speed and use only the clear supernatant.
- Ensure all liquid is at the bottom of the tube before placing it in the thermal cycler [58].
Post-Array Quality Control: A comprehensive QC workflow should be applied to the raw data (.idat files) before any downstream analysis [59]:
- Control Metric Check: Evaluate the 17 control metrics defined by Illumina to identify samples that failed technical steps.
- Sex Check: Infer sex from X and Y chromosome probe intensities and compare with metadata to find mislabeled samples.
- Contamination & Identity Check: Use the 65 SNP probes on the array to:
- Detect contaminated samples (outliers in SNP probe intensities).
- Generate a genetic fingerprint for each sample to identify duplicates or mix-ups (samples with matching fingerprints should have the same donor ID).
Workflow Visualization
The following diagram illustrates the complete integrated workflow for sperm epigenetic clock analysis, from sample collection to biological insight, highlighting key quality control checkpoints.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials and Reagents for Sperm Methylation Analysis
| Item | Function/Description | Example/Note |
|---|---|---|
| Tris(2-carboxyethyl)phosphine (TCEP) | A stable, room-temperature reducing agent critical for breaking protamine disulfide bonds in sperm DNA during extraction [8]. | More stable alternative to dithiothreitol (DTT). |
| Silica-based Spin Columns | For purifying DNA after lysis and reduction in the extraction protocol [8]. | Compatible with the rapid, room-temperature extraction method. |
| Bisulfite Conversion Kit | Chemically converts unmethylated cytosines to uracils, while methylated cytosines remain unchanged, enabling methylation detection [58]. | Ensure kit is validated for array input. |
| Infinium Methylation BeadChip | High-throughput microarray for quantifying DNA methylation at hundreds of thousands of CpG sites [2] [8]. | Illumina EPIC array is common. |
| Hot-Start Taq Polymerase | Recommended for PCR amplification of bisulfite-converted DNA, as it can read through uracil residues [58]. | Proof-reading polymerases are not recommended. |
| Bioinformatics Software (R/Bioconductor) | Packages for quality control, normalization, and analysis of methylation array data (e.g., minfi, ChAMP, ewastools) [60] [59]. |
ewastools is specifically highlighted for QC checks [59]. |
Establishing Clinical Relevance: Validation Frameworks and Comparative Biomarker Analysis
For researchers and drug development professionals in reproductive medicine, validating Sperm Epigenetic Aging (SEA) against meaningful clinical endpoints is a critical step in transitioning from basic research to clinical application. SEA refers to the biological age of sperm cells, estimated using DNA methylation (DNAm) patterns at specific genomic loci, which can diverge from chronological age [2]. This technical support document provides a structured framework for designing and troubleshooting prospective cohort studies that aim to link SEA with live birth rates (LBR), a gold-standard endpoint in fertility research.
The rationale for this approach is strong: chronological age is a suboptimal predictor of individual fertility outcomes. Evidence suggests that a man's biological age, as captured by SEA, may provide a more accurate reflection of his reproductive potential and the likelihood of achieving a live birth [2] [6]. Successfully validating this link is a prerequisite for developing SEA into a robust biomarker that can improve patient stratification, prognostication, and ultimately, the development of novel therapeutic interventions.
Frequently Asked Questions (FAQs) on Cohort Validation
Q1: What is the core hypothesis we are testing in a prospective cohort validation study? The core hypothesis is that advanced sperm epigenetic age acceleration (EAA)—where biological age exceeds chronological age—is independently associated with a reduced probability of achieving a live birth, after controlling for relevant confounders such as female partner's age and standard semen parameters [2] [61].
Q2: Why is a prospective cohort design preferred for this type of validation? Prospective cohorts are ideal because they enable the optimal measurement of exposures (like SEA) before the outcome (live birth) occurs [62]. This temporal sequence strengthens causal inference, minimizes recall bias, and allows for standardized collection of biospecimens and clinical data at baseline.
Q3: Our study found a statistically significant association between SEA and live birth, but the effect size is small. Is this clinically meaningful? A small effect size can still be clinically significant, especially in the context of a multifactorial outcome like live birth. The utility of SEA may lie in its integration into multivariable prediction models. For example, a study validating a live birth prediction model over multiple IVF cycles achieved reasonable discrimination (c-statistic: 0.67-0.75) by combining multiple factors [63]. The incremental value of SEA over existing models (e.g., those based on female age, ovarian reserve) must be assessed.
Q4: We are encountering high variability in SEA measurements within our cohort. What could be the cause? Beyond technical noise, true biological variability is expected. SEA is influenced by a range of factors confirmed in systematic reviews, including environmental exposures such as air pollution, cigarette smoke, and certain chemicals [64]. Failing to account for these in your cohort's inclusion criteria or questionnaire data can introduce uncontrolled heterogeneity. Furthermore, the specific laboratory protocols for sperm processing and DNA methylation analysis must be rigorously standardized.
Q5: How do we handle the confounding effect of the female partner's fertility status? This is a critical design challenge. The most straightforward approach is to restrict the cohort to couples where the female partner has no diagnosed infertility factors. Alternatively, you must meticulously collect and adjust for key female factors in your statistical models, most importantly chronological age and biomarkers of ovarian reserve like Anti-Müllerian Hormone (AMH) and Antral Follicle Count (AFC) [61].
Troubleshooting Common Experimental Challenges
Challenge 1: Inconsistent Correlation Between SEA and Chronological Age
- Problem: The sperm epigenetic clock performs poorly in predicting the chronological age of donors in your cohort (e.g., low correlation coefficient).
- Potential Causes & Solutions:
- Cause: The chosen epigenetic clock model is not optimized for your population or the specific technology platform.
- Solution: Select a clock that has been validated in a population and with a technology (e.g., EPIC array, RRBS) similar to yours [2] [6]. Consider developing a custom clock if existing models are insufficient.
- Cause: Poor quality or contaminated sperm DNA.
- Solution: Implement strict semen processing protocols to isolate pure sperm fractions and ensure high-quality DNA extraction with bisulfite conversion efficiency checks [61] [6].
Challenge 2: Weak or Absent Association with Live Birth
- Problem: Despite a well-performing clock, your study finds no significant link between SEA and LBR.
- Potential Causes & Solutions:
- Cause: Insufficient statistical power due to a small cohort size or a low overall live birth rate.
- Solution: Conduct an a priori sample size calculation. One study aiming to detect a 1.2-year epigenetic age difference recruited 400 women (and their male partners) to achieve 80% power [61]. Ensure your study is similarly powered.
- Cause: Inadequate control for female factors, which are dominant drivers of IVF success.
- Solution: Re-evaluate your statistical model. Use multivariate approaches like discrete-time proportional hazards models to adjust for female age, BMI, and ovarian reserve markers [2].
- Cause: The selected clinical endpoint is inappropriate. "Live birth per cycle" is noisy; "cumulative live birth per woman over multiple complete cycles" is a more stable and clinically meaningful endpoint [63].
Challenge 3: Failure to Replicate Published AgeDMRs
- Problem: Your genome-wide analysis does not identify the age-related differentially methylated regions (ageDMRs) reported in prior literature.
- Potential Causes & Solutions:
- Cause: Low overlap between ageDMRs from different studies is common due to differences in technology, statistical power, and cohort characteristics [6].
- Solution: Focus on replicated genomic regions. A meta-analysis identified 241 genes with ageDMRs replicated across studies, enriched in biological processes related to development and the nervous system [6]. Target these genes for validation.
- Cause: Differences in data processing and statistical modeling (e.g., normalization, FDR correction).
- Solution: Closely replicate the bioinformatic pipelines of the original studies where possible.
Essential Research Reagents & Solutions
Table 1: Key Research Reagents for SEA Studies
| Reagent / Material | Function / Application | Considerations for Use |
|---|---|---|
| Semen Sample | Source of sperm DNA for epigenetic analysis. | Standardize collection (abstinence time), processing (somatic cell removal), and cryopreservation protocols [2]. |
| Bisulfite Conversion Kit | Converts unmethylated cytosines to uracils, allowing methylation quantification. | Conversion efficiency is critical; use kits with high conversion rates and include controls. |
| DNA Methylation Platform | Profiling methylation. | Infinium Methylation EPIC array offers a broad, cost-effective solution [2]. RRBS/WGBS provides base-resolution, genome-wide data [6]. |
| Sperm-Specific Epigenetic Clock | Algorithm to predict biological age from sperm DNAm data. | Choose a validated model. Some clocks use CpG sites [2], while others use differentially methylated regions (DMRs) [2]. Ensure compatibility with your data. |
| Bioinformatic Pipelines | For processing raw methylation data, normalization, and clock calculation. | Use established packages (e.g., minfi in R) and consistently apply the same preprocessing steps to all samples. |
Standardized Experimental Protocol for SEA Validation
Phase 1: Cohort Design and Participant Recruitment
- Define Inclusion/Exclusion Criteria: Ideally, recruit couples with unexplained infertility or those with only a male factor. Exclude couples with severe female factor infertility (e.g., diminished ovarian reserve, uterine anomalies) to reduce confounding [61].
- Collect Informed Consent and Baseline Data: Obtain consent for long-term follow-up. Collect data on both partners: demographics, medical/reproductive history, lifestyle (smoking, alcohol), and environmental exposures [64].
- Specify Clinical Endpoint: Clearly define the primary endpoint, preferably as "cumulative live birth resulting from all embryo transfers within a predefined treatment period (e.g., one year)" [63].
Phase 2: Biospecimen Collection and Processing
- Semen Collection: Collect semen sample after a recommended period of sexual abstinence (2-7 days). Use a standardized collection kit.
- Sperm DNA Extraction: Isolate sperm cells using density gradient centrifugation to minimize somatic cell contamination [6]. Extract high-molecular-weight DNA using a commercially available kit (e.g., DNeasy Blood & Tissue Kit) [61].
- DNA Quality Control: Assess DNA concentration, purity (A260/280), and integrity (e.g., via gel electrophoresis) before proceeding.
Phase 3: DNA Methylation Analysis and SEA Calculation
- Bisulfite Conversion: Treat DNA with bisulfite using a reliable kit. Include unmethylated and methylated DNA controls to monitor conversion efficiency.
- Methylation Profiling:
- Option A (Targeted): Use pyrosequencing or MS-SNuPE for clocks based on a few CpGs (e.g., 5-8 sites). This is cost-effective for large cohorts [65] [61].
- Option B (Genome-wide): Use the Illumina EPIC array or sequencing-based methods (RRBS/WGBS) for discovery or to apply more complex clocks [2] [6].
- Data Preprocessing: Process raw data for background correction, normalization, and probe filtering. Address technical batch effects.
- Calculate SEA and EAA: Apply the chosen sperm epigenetic clock algorithm to estimate SEA. Calculate Epigenetic Age Acceleration (EAA) as the residual from a regression model of epigenetic age on chronological age [61].
Phase 4: Statistical Analysis and Validation
- Data Integration: Merge SEA/EAA data with clinical outcomes and covariate information.
- Primary Analysis: Use a discrete-time proportional hazards model or logistic regression to assess the association between SEA/EAA and time-to-pregnancy (TTP) or live birth, adjusting for female age, BMI, and other relevant confounders [2].
- Model Performance: Evaluate the predictive performance of SEA by calculating the Area Under the Curve (AUC) and assess its incremental value over established predictors.
Table 2: Summary of Key Findings from Relevant Studies on Epigenetic Aging and Reproduction
| Study (Year) | Cohort & Design | Epigenetic Metric | Key Finding Related to Live Birth / Pregnancy | Effect Size / Statistical Result |
|---|---|---|---|---|
| LIFE Study (2022) [2] | Prospective cohort of 379 couples from the general population. | Sperm Epigenetic Clock (SEACpG) | SEA was negatively associated with pregnancy success. | FOR=0.83; 95% CI: 0.76, 0.90 per year increase in SEA. |
| IVF Cohort Study (2025) [61] | Prospective observational study of 379 women undergoing IVF. | Blood Epigenetic Age in Women | Lower epigenetic age in women was associated with a higher live birth rate (LBR). | LBR: 54% in epigenetically younger vs. others. Adjusted OR = 0.91 per year. |
| Sperm ageDMRs (2023) [6] | Analysis of 73 sperm samples from an IVF/ICSI cohort. | Age-related DMRs (ageDMRs) | No significant correlation found between ageDMRs and pregnancy outcome in this specific analysis. | Reported no significant association. |
| HFEA Model Validation (2023) [63] | External validation of a live birth prediction model (n=91,035 women). | Clinical Prediction Model (Not epigenetic) | Highlights the standard for predictive performance in IVF. | Validated model c-statistic: 0.67 (pre-treatment) to 0.75 (post-treatment). |
Abbreviations: FOR: Fecundability Odds Ratio (FOR < 1 indicates longer time to pregnancy); OR: Odds Ratio; CI: Confidence Interval; LBR: Live Birth Rate.
FAQ: Understanding Performance Metrics
What are the key performance metrics for a sperm epigenetic clock, and what values are considered good?
For sperm epigenetic clocks, the primary metrics are Mean Absolute Error (MAE) and the correlation coefficient (r) between predicted epigenetic age and chronological age. MAE represents the average absolute difference between predicted and actual chronological age, while r indicates the strength of the linear relationship.
The table below summarizes performance metrics from key studies:
| Study / Clock | Cohort Size | Tissue | Key Performance Metrics | Notes |
|---|---|---|---|---|
| LIFE Study Clock [2] | 379 | Sperm | MAE: Not specified; Correlation (r) with age: 0.91 | Fecundability Odds Ratio (FOR) = 0.83 for time-to-pregnancy. |
| VISAGE Consortium Clock [15] | 54 (Test Set) | Semen | MAE: 5.1 years | Model based on 6 CpGs from genes like SH2B2 and FOLH1B. |
| Lee et al. 3-CpG Model [15] | N/A | Semen | MAE: ~5 years | A minimal model for forensic applications. |
| Horvath Pan-Tissue Clock [66] | 3,931 (Training) | 51 Tissues | Median Absolute Deviation: 3.6 years | A widely used first-generation "pan-tissue" clock. |
How is generalizability evaluated, and why is it a major challenge?
Generalizability is assessed by applying a clock trained on one cohort to an entirely independent cohort. A significant drop in performance on the external cohort indicates poor generalizability. Challenges include:
- Cohort Demographics: Clocks trained on predominantly Caucasian populations may not perform well in other ethnicities [2].
- Cell Composition: Sperm samples can vary in purity. Changes in somatic cell contamination act as a confounding variable, similar to how shifts in naive T-cell proportions affect blood-based clocks [16].
- Technical Variation: Differences in DNA methylation measurement platforms (e.g., different Illumina array versions or targeted sequencing) can introduce significant inaccuracies. One study found that mismatches between a clock's CpG sites and those represented on newer DNA chips could lead to errors averaging 3 years and up to 25 years in some cases [67].
Our sperm epigenetic clock performs well on the training data but poorly on an external validation cohort. What are the primary sources of error we should investigate?
This is a classic sign of overfitting or cohort-specific bias. Your troubleshooting should focus on:
- Biological Variance: Check for differences in the age distribution, ethnicity, health status, or environmental exposures (e.g., smoking rates) between your training and validation cohorts. Current smoking has been associated with advanced sperm epigenetic age [2].
- Technical Variance: Ensure consistent sample processing, DNA extraction, and methylation measurement platforms across cohorts. Inaccurate age data in the training set can propagate error; one study suggests that more than 22% error in training data ages leads to a significant increase in prediction error [68].
- Cell Type Specificity: Confirm that your clock is based on markers truly specific to sperm cells. Sperm has a very different age-related methylation pattern compared to somatic tissues [15]. Contamination from somatic cells, whose methylation profiles change with age differently, can skew predictions if not accounted for.
Troubleshooting Guides
Guide: Improving Accuracy and Precision
Problem: High Mean Absolute Error (MAE) in age prediction.
| Symptom | Potential Cause | Solution |
|---|---|---|
| Consistent bias (all predictions are too high/low) | Batch effects or technical drift during processing. | Implement a rigorous calibration protocol using control samples across batches. |
| High variance (predictions are scattered) | The model is overfitted or the training set is too small/homogeneous. | Employ machine learning algorithms with built-in regularization (e.g., elastic net regression). Increase training sample size and diversity [2]. |
| Good performance in training, poor in validation | The model has learned cohort-specific artifacts, not true biological aging. | Use a hybrid approach: train on a large, public dataset and fine-tune on a smaller, targeted sperm dataset. Apply the clock to an independent cohort as a first validation step [25]. |
Experimental Protocol for Rigorous Validation:
- Cohort Design: Recruit a sufficient number of participants (n > 200 is ideal) [68] covering a wide and evenly distributed age range (e.g., 20-60 years).
- Sample Collection: Standardize semen collection and processing protocols to minimize pre-analytical variation. Include a step to purify sperm cells if possible to reduce somatic cell contamination.
- DNA Methylation Analysis: Use a consistent, high-resolution platform (e.g., Illumina MethylationEPIC array). Replicate measurements can help quantify technical noise.
- Model Training: Use an ensemble machine learning algorithm or elastic net regression, which automatically selects the most predictive CpG sites [2].
- Validation: Split data into training (e.g., 70%) and testing (e.g., 30%) sets. Crucially, validate the final model on a completely independent cohort from a different geographical location or ethnicity [2].
Guide: Ensuring Generalizability Across Populations
Problem: The epigenetic clock fails to maintain accuracy when applied to a new population.
Checklist for Assessing Generalizability:
- Population Representation: Does your training cohort adequately represent the genetic ancestry, lifestyle, and environmental exposures of the target population for your clock?
- Marker Selection: Have you selected CpG sites that are robustly age-associated across multiple independent datasets? Avoid sites that are highly variable due to non-age-related factors.
- Confounding Factors: Have you recorded and can you adjust for confounding variables such as smoking status, BMI, and fertility diagnoses in your model? [2]
- Technical Reproducibility: Is your measurement technology (e.g., a targeted SNaPshot assay) reproducible across different laboratories? One study showed a strong correlation (r = 0.97) between a targeted assay and the gold-standard Illumina array, which is promising for transferability [69].
The following workflow outlines a systematic approach to develop and validate a generalizable sperm epigenetic clock:
The Scientist's Toolkit: Research Reagent Solutions
This table details key materials and their functions for developing and validating sperm epigenetic clocks.
| Research Reagent | Function in Sperm Epigenetic Clock Research |
|---|---|
| Infinium MethylationEPIC BeadChip | Genome-wide DNA methylation screening tool for discovery phase; analyzes over 850,000 CpG sites [15]. |
| Targeted Bisulfite MPS (Massively Parallel Sequencing) | Validation and application technology for focused analysis of specific age-related CpGs; more suitable for forensic or clinical settings [15]. |
| Multiplex Methylation SNaPshot Assay | A targeted, cost-effective method for analyzing a small panel of key age-related CpG sites (e.g., in ELOVL2, FHL2); highly reproducible across labs [69]. |
| Bisulfite Conversion Reagents | Critical for pre-treating DNA before methylation analysis; converts unmethylated cytosines to uracils, allowing methylation status to be determined. |
| Elastic Net Regression | A machine learning algorithm used for model training; performs variable selection and regularization to prevent overfitting and identify the most predictive CpG sites [2] [66]. |
| Purified Sperm Cell Fractions | Samples processed to minimize somatic cell (e.g., leukocyte) contamination; crucial for ensuring the clock measures sperm-specific aging, not a mixed signal [15] [16]. |
The logical pathway from raw sample to a validated age prediction involves coordinated use of these reagents, as shown below:
For decades, the standard semen analysis—evaluating parameters like sperm concentration, motility, and morphology—has been the cornerstone of male fertility assessment, guided by World Health Organization (WHO) manuals [70] [8]. However, a significant limitation persists: these standard semen parameters are relatively poor predictors of actual reproductive success and fecundability (the probability of achieving pregnancy within a given menstrual cycle) [2] [8]. This diagnostic gap has driven the search for more robust biomarkers, leading to the emergence of Sperm Epigenetic Age (SEA) as a novel and promising metric [2] [10].
SEA refers to the biological age of sperm, estimated from specific patterns of DNA methylation, which can differ significantly from the donor's chronological age [2] [8]. This technical support article provides a comparative evaluation of SEA and traditional semen analysis, offering troubleshooting guides and detailed protocols to assist researchers in integrating this advanced biomarker into their studies on fecundability.
Comparative Analysis: SEA vs. Traditional Semen Parameters
The table below summarizes the core differences between SEA and traditional semen analysis based on current literature.
Table 1: Comparative analysis of SEA and traditional semen parameters
| Feature | Sperm Epigenetic Age (SEA) | Traditional Semen Analysis |
|---|---|---|
| Core Principle | Biological age of sperm based on DNA methylation patterns [2] | Physical and microscopic evaluation of semen quality (count, motility, morphology) [70] |
| Primary Output | Quantitative metric (Age in years); Epigenetic Age Acceleration (difference from chronological age) [2] [15] | Quantitative metrics (e.g., million/mL, %, %) and qualitative descriptions [70] |
| Association with Fecundability | Strong, independent association with longer Time-to-Pregnancy (TTP) and lower pregnancy probability [2] [10] | Weak and inconsistent predictor of pregnancy success in couples [2] [8] |
| Key Supporting Data | 17% lower cumulative pregnancy probability after 12 months for couples with older SEA; Fecundability Odds Ratio (FOR)=0.83 per unit increase in SEA [2] [10] | Poor correlation with reproductive outcomes in clinical and population-based cohorts [8] |
| Relation to Chronological Age | Correlates with but is distinct from chronological age (r=0.91 in one clock model) [2] | Parameters can decline with age, but not a direct measure of biological aging [70] |
| Influence of Lifestyle | Associated with modifiable factors, e.g., advanced SEA observed in smokers [2] [10] | Influenced by health and lifestyle, but not as a direct, quantifiable biomarker of biological aging |
Experimental Protocols for SEA Analysis
Core Workflow for Sperm Epigenetic Clock Construction and Application
The following diagram outlines the generalized workflow for developing and applying a sperm epigenetic clock, from sample collection to age prediction.
Detailed Methodology: Key Steps and Reagents
This section details the critical wet-lab and computational procedures based on published studies.
1. Semen Sample Collection and Sperm DNA Isolation
- Protocol: Collect semen samples after a recommended 2-3 days of ejaculatory abstinence. For DNA isolation, treat sperm with a reducing agent like Tris(2-carboxyethyl)phosphine (TCEP) to break down protamine-based packaging, followed by lysis with guanidine thiocyanate and purification using silica-based spin columns [8].
- Troubleshooting: For home-collected samples shipped overnight, motility analysis may be unreliable; focus on DNA integrity for epigenetic work. The use of TCEP is critical for efficient sperm lysis [8].
2. Bisulfite Conversion and Methylation Profiling
- Principle: Bisulfite treatment converts unmethylated cytosine to uracil, while methylated cytosine remains unchanged, allowing for the quantification of methylation levels.
- Profiling Platforms:
- Infinium MethylationEPIC BeadChip Array: Covers over 850,000 CpG sites. Ideal for discovery phases due to broad coverage [2] [15] [8].
- Bisulfite Amplicon Sequencing (BSAS): A targeted, high-depth sequencing method suitable for validating and applying clocks from a smaller set of CpGs. More applicable for forensic or low-DNA contexts [7].
- double-enzyme Reduced Representation Bisulfite Sequencing (dRRBS): Provides cost-effective, genome-wide coverage beyond microarray platforms, enabling discovery of novel, high-performance age-related CpG sites [7].
3. Predictive Model Building and Validation
- Feature Selection: Identify age-related CpG (AR-CpG) sites from methylation data. Studies have successfully built accurate clocks using 6 to 9 CpG sites [15] [7].
- Machine Learning: Use algorithms like Random Forest or ensemble methods for model training. One study using an ensemble machine learning algorithm achieved a high correlation (r=0.91) between predicted SEA and chronological age [2].
- Validation: Always validate the model's prediction accuracy (e.g., Mean Absolute Error - MAE) on an independent, held-out test set. MAEs of ~3.3 to 5.1 years have been reported in recent models [15] [7].
Frequently Asked Questions (FAQs) for Researchers
Q1: My research shows no correlation between standard semen parameters and SEA. Is this expected? Yes, this is a consistent finding. A 2024 study that analyzed both a clinical (SEEDS) and a non-clinical (LIFE) cohort found that SEA was not associated with standard semen characteristics like concentration or motility [8]. SEA appears to be an independent biomarker, capturing information about biological aging that is distinct from traditional quality measures.
Q2: What is the clinical relevance of SEA in predicting fecundability? Research on couples from the general population has shown that advanced SEA is significantly associated with a longer Time-to-Pregnancy (TTP). For example, a 2022 study reported a 17% lower cumulative probability of pregnancy after 12 months for couples where the male partner had an older SEA. The Fecundability Odds Ratio (FOR) was 0.83, indicating a longer TTP with advanced SEA [2] [10].
Q3: How does paternal age influence SEA and genetic risk? Chronological age is a strong driver of SEA. Furthermore, groundbreaking 2025 research using ultra-accurate sequencing (NanoSeq) revealed that as men age, harmful genetic mutations in sperm become more common—increasing from about 2% in men in their early 30s to 3-5% in middle-aged and older men [11] [27]. This is due to a process of natural selection within the testes that favors certain mutations, many linked to severe neurodevelopmental disorders and inherited cancer risk [11] [27].
Q4: Can lifestyle factors influence SEA? Yes, modifiable factors like smoking have been associated with advanced SEA. One study found that current smokers displayed significantly older SEA compared to non-smokers, suggesting that lifestyle interventions could potentially modify sperm biological age [2] [10].
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key reagents and materials for SEA research
| Item | Function/Benefit | Example/Note |
|---|---|---|
| Tris(2-carboxyethyl)phosphine (TCEP) | Reducing agent for efficient sperm cell lysis by breaking disulfide bonds in protamines. More stable than DTT at room temperature [8]. | Critical for high-quality DNA yield from sperm. |
| Infinium MethylationEPIC BeadChip | Microarray for genome-wide methylation profiling at >850,000 CpG sites. Ideal for initial clock building and discovery [2] [8]. | Standard for broad discovery. |
| Bisulfite Conversion Kit | Prepares DNA for methylation analysis by deaminating unmethylated cytosines. | Select kits optimized for low-input DNA for forensic applications. |
| Silica-Based Spin Columns | For purifying DNA after lysis and bisulfite conversion. | Compatible with the rapid sperm DNA extraction method [8]. |
| dRRBS or BSAS Reagents | For high-depth, targeted methylation sequencing. dRRBS is cost-effective for discovery; BSAS is ideal for validating and applying multi-CpG models [7]. | Enables high-accuracy models with a minimal set of CpGs. |
Troubleshooting Common Experimental Challenges
Problem: High Error in Age Prediction from Semen Stains.
- Potential Cause: Low quantity and quality of input DNA, which is further compromised by the bisulfite conversion process, leading to incomplete or biased data [15].
- Solution: Utilize targeted MPS technologies (like BSAS) that are designed for low-input and degraded DNA. Focus on models with a minimal number of highly informative CpG sites to maximize information from compromised samples [15] [7].
Problem: Inconsistent Correlation of CpG Sites Across Studies.
- Potential Cause: Differences in the studied populations (e.g., ethnicity), sample processing protocols, or statistical methods for clock construction [2] [7].
- Solution: Always validate identified AR-CpG sites in an independent cohort from your target population. When developing a new clock, use a two-stage validation process (discovery followed by confirmation) to ensure robustness [7].
Problem: Sperm Sample Contaminated with Somatic Cells.
- Potential Cause: Somatic cells (e.g., leukocytes) have drastically different methylation patterns, which can confound the sperm-specific SEA signal [7].
- Solution: Implement a strict density gradient centrifugation step during semen processing to isolate a pure sperm fraction before DNA extraction [8].
Fundamental Concepts & Troubleshooting
FAQ 1: What is the fundamental principle behind an epigenetic clock, and how is it applied to murine sperm? Epigenetic clocks are mathematical models that predict chronological or biological age based on patterns of DNA methylation (DNAm) at specific CpG sites in the genome. These age-associated methylation changes are a robust biomarker of the aging process. In murine sperm, these clocks are built by profiling DNA methylation in sperm samples from mice of different ages and using machine learning (e.g., elastic net regression) to identify a predictive set of CpG sites whose methylation levels correlate strongly with age [71]. The primary goal is to use this "epigenetic age" as a readout for studying how factors like stress, diet, or toxins affect the male germline and potentially offspring health [6] [72].
FAQ 2: My murine sperm epigenetic clock shows poor accuracy when applied to a different mouse strain. What is the likely cause and how can I address this? A primary cause is genetic background differences. Different inbred strains, such as C57BL/6 and DBA/2, exhibit distinct baseline methylation levels and rates of age-related change, leading to systematic over- or under-estimation of age [73].
- Solution: Strain-Specific Retraining. You cannot directly apply a clock trained on one strain to another. You must retrain the model using age-structured sperm methylation data from your target strain. For example, researchers developed a specific multivariate model for DBA/2 mice after finding that the slope of methylation change with age, particularly at loci like Prima1, differed from C57BL/6 mice [73].
- Actionable Protocol:
- Collect sperm samples from your target strain across a wide age range (e.g., 10-100 weeks).
- Perform DNA methylation analysis (e.g., RRBS or Mammalian Methylation Array).
- Identify age-correlated CpGs within your strain's data.
- Train a new elastic net or ridge regression model using these strain-specific CpGs to create a custom clock [71].
FAQ 3: Why do my epigenetic age predictions vary wildly between different clock models when using the same sperm samples? This is a common challenge due to the lack of standardization in epigenetic clock development. Different clocks may use different CpG sites, regression techniques (ridge vs. elastic net), and training datasets, leading to inconsistent results [74] [67].
- Solution: Use an Ensemble Approach. Instead of relying on a single clock, use a framework like EnsembleAge, which combines predictions from multiple, high-performing individual clocks. This method has been shown to outperform single clocks in detecting both pro-aging and rejuvenating interventions by reducing model-specific biases and improving robustness [74].
- Actionable Protocol: When analyzing your data, run it through several established murine epigenetic clocks and calculate the median predicted age (EnsembleAge.Dynamic) or use a pre-trained static ensemble model for a more reliable and consensus estimate [74].
FAQ 4: Can an epigenetic clock trained on blood or liver be used to estimate age from sperm samples? No. Epigenetic clocks are generally tissue-specific. While some age-related methylation changes may be consistent across tissues, the model requires retraining for each tissue type. A clock trained on blood will not provide accurate age estimates for sperm [73].
- Evidence: A study testing a blood-based 3-CpG clock on other mouse tissues (skin, kidney, liver, etc.) found that while DNAm from old mice was consistently predicted to be older than that from young mice, the absolute values were inaccurate. The authors concluded that "the model needs to be retrained to be applied for these tissues" [73].
- Guideline: Always use a clock that was specifically trained on murine sperm methylation data for sperm analysis.
Technical & Analytical Challenges
FAQ 5: How can I ensure my sperm methylation data is of high quality and free from somatic cell contamination? Sperm preparation is critical. Somatic cell contamination will severely skew methylation results, as the epigenetic profiles of other cells are vastly different.
- Solution: Validate Imprinting Control Regions (ICRs). A reliable quality control is to check the methylation status of known maternally imprinted (paternally methylated) germline DMRs.
- Actionable Protocol: In your DNA methylation data, examine specific ICRs like H19/IGF2:IG-DMR and IGF2:alt-TSS-DMR. These should be hypermethylated (e.g., >80-90% methylation) in pure sperm samples. The absence of this pattern indicates potential contamination or improper imprinting [6].
FAQ 6: My study involves an intervention (e.g., stress, diet). How can I distinguish true epigenetic aging from intervention-specific methylation changes? This is a key issue in intervention studies. The observed methylation shifts might reflect the intervention's acute effect rather than a change in the underlying aging rate.
- Solution: Include Multiple Control Groups. Your experimental design is the best defense.
- Actionable Protocol:
- Age-Matched Controls: Standard controls that match the chronological age of your treated group.
- Baseline Group: A group sacrificed at the start of the intervention to establish the baseline epigenetic age.
- Caloric Restriction Control: For certain interventions, including a caloric restriction group (a gold-standard anti-aging intervention) can help benchmark your clock's ability to detect true rejuvenation [74] [71]. By comparing the treated group's epigenetic age acceleration (EAA) against these controls, you can better attribute changes to the intervention's effect on aging.
FAQ 7: What is the best technology for profiling DNA methylation in murine sperm for clock construction? The choice involves a trade-off between cost, coverage, and consistency.
Table 1: Comparison of DNA Methylation Profiling Technologies
| Technology | Key Features | Pros | Cons | Best For |
|---|---|---|---|---|
| RRBS [71] | Selectively sequences CpG-rich regions. | Cost-effective for genome-wide coverage; avoids CpG density bias of arrays. | Inconsistent coverage across samples; can miss relevant CpGs. | Developing new clocks with broad, unbiased discovery. |
| Mammalian Methylation Array [74] | Microarray targeting evolutionarily conserved CpGs. | High reproducibility; consistent measurement of the same CpGs across all samples. | Limited to pre-defined CpG set; may miss novel, sperm-specific sites. | Large-scale studies and cross-species comparisons where consistency is key. |
| Pyrosequencing [73] | Quantifies methylation at a few specific CpGs. | Very cost-effective, simple, and highly accurate for validating individual sites. | Only tests known CpGs; not for discovery. | Validating and applying pre-existing, simple clocks (e.g., a 3-CpG model). |
Experimental Design & Advanced Applications
FAQ 8: What is a robust experimental workflow for a murine sperm epigenetic clock study? The following diagram outlines a workflow that incorporates validation and troubleshooting steps to ensure robust findings:
FAQ 9: How can I investigate the potential for paternal intergenerational epigenetic inheritance using these clocks? Sperm epigenetic clocks are a tool to measure aging-associated changes in the germline that might be transmitted to offspring.
- Experimental Workflow:
- Expose: Subject male mice (F0) to an environmental factor (e.g., long-term psychological stress, high-fat diet) [72].
- Measure: Use a sperm epigenetic clock to quantify the epigenetic age acceleration in the exposed F0 males versus controls.
- Breed: Generate offspring (F1) from these males.
- Correlate: Analyze the F1 offspring for:
- Key Insight: Studies show that while most methylation changes are erased post-fertilization, a small subset of sperm DMRs can evade reprogramming and be re-established in the offspring, potentially mediating the inheritance of paternal disorders [72].
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagents and Resources for Murine Sperm Epigenetic Clock Research
| Item / Reagent | Critical Function | Example & Notes |
|---|---|---|
| Inbred Mouse Strains | Model organism for controlled genetic studies. | C57BL/6J: Most common background. DBA/2: Used for comparative aging studies due to shorter lifespan [73]. |
| Bisulfite Conversion Kit | Chemically converts unmethylated cytosines to uracils, allowing methylation status to be read by sequencing or PCR. | EZ DNA Methylation Kit (ZymoResearch): A standard for high-conversion efficiency [75]. Critical for all downstream methylation analysis. |
| Methylation Profiling Platform | Genome-wide or targeted measurement of DNA methylation levels. | Illumina Methylation Arrays: Mammalian Methylation Array for cross-species consistency [74]. Pyrosequencing (Qiagen PyroMark): For targeted, quantitative validation of specific CpGs [73] [75]. |
| Bioinformatics Software | For statistical analysis, clock training, and age prediction. | R Packages glmnet & mlr: Essential for building penalized regression models (ridge, lasso, elastic net) for clock development [74] [71]. |
| Sperm Isolation Protocol | To obtain pure sperm cell populations free of somatic cells. | Protocol involving tissue mincing and swim-up or density gradient centrifugation. Quality must be verified by ICR analysis [6]. |
| Validated Primers & Probes | For targeted amplification and sequencing of specific CpG sites. | Primers for pyrosequencing of clock loci (e.g., Prima1, Hsf4, Kcns1 in mice) [73]. Must be designed for bisulfite-converted DNA. |
Conclusion
The path to optimized sperm epigenetic clocks hinges on a multi-faceted approach that integrates foundational biology, advanced computational methodologies, rigorous troubleshooting of confounders, and robust clinical validation. Future efforts must prioritize the development of large, diverse, and well-annotated sample cohorts to train next-generation clocks that move beyond chronological age prediction to capture biological aging processes relevant to reproductive success. Furthermore, elucidating the functional role of the blood-testis barrier and other mechanisms in mediating environmental effects on the sperm epigenome will be crucial. The ultimate goal is the translation of these precise biomarkers into clinical practice, enabling improved diagnosis of male infertility, personalized risk assessment, and the evaluation of interventions aimed at mitigating adverse reproductive and intergenerational health outcomes.
References
- 1. Associations between Sperm Epigenetic Age and Semen ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10887546/]
- 2. Sperm epigenetic clock associates with pregnancy ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9247414/]
- 3. Sperm epigenetics and aging - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6087840/]
- 4. Epigenetic age prediction in semen – marker selection and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8386575/]
- 5. Epigenetic Clocks: Beyond Biological Age, Using the Past to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12539533/]
- 6. Age-related methylation changes in the human sperm ... [https://www.aging-us.com/article/204546/text]
- 7. DNA methylation-based age estimation from semen [https://www.sciencedirect.com/science/article/abs/pii/S1872497324002114]
- 8. Associations between Sperm Epigenetic Age and Semen ... [https://www.mdpi.com/1467-3045/46/2/101]
- 9. A systematic review and meta-analysis of epigenetic age ... [https://www.sciencedirect.com/science/article/pii/S1568163724003702]
- 10. New measure of sperm age may be predictor of pregnancy ... [https://today.wayne.edu/medicine/news/2022/05/13/new-measure-of-sperm-age-may-be-predictor-of-pregnancy-success-47890]
- 11. Harmful DNA changes in the sperm found to rise sharply ... [https://www.news-medical.net/news/20251008/Harmful-DNA-changes-in-the-sperm-found-to-rise-sharply-with-age.aspx]
- 12. 精子表观遗传时钟与普通人群的妊娠结局相关 [https://toxlab.ahmu.edu.cn/2022/0716/c10752a182477/page.htm]
- 13. 强悍的生育能力,靠你的精子『生理学』年龄而非『身份证』年龄 [https://news.bioon.com/article/d8a7e3014221.html]
- 14. Investigating Aging and DNA Methylation: A Path to Improving ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12204039/]
- 15. Epigenetic age prediction in semen – marker selection and ... [https://www.aging-us.com/article/203399/text]
- 16. Development of an epigenetic clock resistant to changes in ... [https://www.nature.com/articles/s42003-024-06609-4]
- 17. Use of a sperm morphology assessment standardisation ... [https://www.nature.com/articles/s41598-025-07515-3]
- 18. An Expert Review From the French BLEFCO Group [https://pubmed.ncbi.nlm.nih.gov/41185394/]
- 19. Potential effects of environmental toxicants on sperm quality ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12133492/]
- 20. Environmental toxicants and male fertility [https://www.sciencedirect.com/science/article/pii/S1521693422001791]
- 21. Flow Cytometry in the Assessment of Sperm Function ... [https://pubmed.ncbi.nlm.nih.gov/40202643/]
- 22. Sperm chromatin: Evaluation, epigenetic signatures and ... [https://www.sciencedirect.com/science/article/pii/S0171933524000463]
- 23. Comparison of staining protocols to assess sperm viability ... [https://www.sciencedirect.com/science/article/pii/S2773093X2500011X]
- 24. Many chronological aging clocks can be found throughout the... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8590098/]
- 25. DNA methylation aging clocks : challenges and recommendations [https://genomebiology.biomedcentral.com/articles/10.1186/s13059-019-1824-y]
- 26. Sperm chromatin and DNA integrity, methyltransferase ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5897243/]
- 27. The hidden evolution making men's sperm more ... [https://www.sciencedaily.com/releases/2025/10/251019120513.htm]
- 28. The effect of age and abstinence time on semen quality [https://pmc.ncbi.nlm.nih.gov/articles/PMC8788608/]
- 29. Epigenetic clock analysis of blood samples in drug-naive first ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9843886/]
- 30. Increasing age in men is negatively associated with sperm ... [https://www.frontiersin.org/journals/aging/articles/10.3389/fragi.2025.1603916/full]
- 31. The male biological clock is ticking: a review of the literature [https://pmc.ncbi.nlm.nih.gov/articles/PMC11026002/]
- 32. Turning back the epigenetic clock: Can we reverse ageing? [https://www.ddw-online.com/turning-back-the-epigenetic-clock-can-we-reverse-ageing-38601-202511/]
- 33. Global effects of identity and aging on the human sperm ... [https://clinicalepigeneticsjournal.biomedcentral.com/articles/10.1186/s13148-023-01541-6]
- 34. Epigenetic age signatures in the forensically relevant body ... [https://www.sciencedirect.com/science/article/abs/pii/S1872497315300181]
- 35. ML Models on a Data Diet. How Training Set Size Impacts… [https://medium.com/data-from-the-trenches/ml-models-on-a-data-diet-440c2d94a0c]
- 36. Improved epigenetic age prediction models by combining ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12261677/]
- 37. What impact does increasing the training data have on ... [https://stats.stackexchange.com/questions/31249/what-impact-does-increasing-the-training-data-have-on-the-overall-system-accurac]
- 38. Impact of Dataset Size on Deep Learning Model [https://www.geeksforgeeks.org/deep-learning/impact-of-dataset-size-on-deep-learning-model/]
- 39. CUE: CpG impUtation ensemble for DNA methylation levels across the human methylation450 (HM450) and EPIC (HM850) BeadChip platforms - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8330997/]
- 40. Lab 3: Penalized regression techniques for high-dimensional ... [https://statomics.github.io/HDDA21/Lab3-Penalized-Regression.html]
- 41. GitHub - GangLiTarheel/CUE: CUE: CpG impUtation Ensemble for DNA Methylation Levels Across the HumanMethylation450 (HM450) BeadChip and HumanMethylation EPIC (HM850) BeadChip Platforms [https://github.com/GangLiTarheel/CUE]
- 42. Development of an epigenetic clock to predict visual age ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10809641/]
- 43. The ticking 'epigenetic' clock measures true biological age [https://www.stjude.org/research/progress/2024/ticking-epigenetic-clock-measures-true-biological-age.html]
- 44. Why Epigenetic Clocks May Fail to Measure Anti- ... [https://www.aging-us.com/news-room/why-epigenetic-clocks-may-fail-to-measure-anti-aging-effects]
- 45. Troubleshooting Protocols - Biological Sciences: Summer ... [https://guides.library.cmu.edu/biologicalsciencesSRI/troubleshooting]
- 46. Teaching troubleshooting skills to graduate students - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11407763/]
- 47. Batch effects in single-cell RNA sequencing data are ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6152897/]
- 48. Integrating single-cell RNA-seq datasets with substantial ... [https://bmcgenomics.biomedcentral.com/articles/10.1186/s12864-025-12126-3]
- 49. Alleviating batch effects in cell type deconvolution with ... [https://www.nature.com/articles/s41467-024-55213-x]
- 50. - June 2023: Exposures affect men’s biological... Environmental Factor [https://www.niehs.nih.gov/news/factor/2023/6/feature/3-mens-reproductive-epigenetics]
- 51. Cigarette Smoking Significantly Alters Sperm DNA Methylation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5679018/]
- 52. The Impact of Heavy Smoking on Male Infertility and Its ... [https://www.mdpi.com/2073-4425/14/8/1617]
- 53. How do lifestyle and environmental influence the factors ... sperm [https://clinicalepigeneticsjournal.biomedcentral.com/articles/10.1186/s13148-025-01815-1]
- 54. Sperm DNA methylation altered by THC and nicotine [https://www.nature.com/articles/s41598-020-72783-0]
- 55. Environmental factors accelerate epigenetic aging of ... [https://www.sciencedirect.com/science/article/pii/S2666335X25000680]
- 56. Mechanistic target of rapamycin (mTOR) pathway in Sertoli ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11405012/]
- 57. DNA methylation aging clocks - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC6876109/]
- 58. DNA Methylation Analysis Support—Troubleshooting [https://www.thermofisher.com/us/en/home/technical-resources/technical-reference-library/epigenetics-support-center/dna-methylation-analysis-support/dna-methylation-analysis-support-troubleshooting.html]
- 59. Identifying mislabeled and contaminated DNA methylation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5984806/]
- 60. How to Optimize Data Analysis from Methylation Arrays [https://www.cd-genomics.com/resource-methylation-arrays-data-analysis-tips.html]
- 61. testing an epigenetic clock to forecast in vitro fertilization ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12257822/]
- 62. Methods for Stratification and Validation Cohorts [https://pmc.ncbi.nlm.nih.gov/articles/PMC9144352/]
- 63. External validation of models for predicting cumulative live over... birth [https://pubmed.ncbi.nlm.nih.gov/37632223/]
- 64. Chemical and climatic environmental exposures ... [https://www.sciencedirect.com/science/article/pii/S0013935125005985]
- 65. Simplified Assay for Epigenetic Age Estimation in Whole ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4943959/]
- 66. Epigenetic age prediction - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8441394/]
- 67. Are 'epigenetic clocks' reliable? CNIO researchers refine ... [https://www.cnio.es/en/news/are-epigenetic-clocks-reliable-cnio-researchers-refine-the-accuracy-of-these-tests-that-measure-ageing/]
- 68. Calibrating epigenetic clocks with training data error - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10445086/]
- 69. Reproducibility and validation of a targeted and flexible ... [https://www.sciencedirect.com/science/article/pii/S0379073825000477]
- 70. Semen evaluation: methodological advancements in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8255896/]
- 71. A multi-tissue full lifespan epigenetic clock for mice [https://www.aging-us.com/article/101590/text]
- 72. Sperm epigenetic alterations contribute to inter [https://www.nature.com/articles/s41421-021-00343-5]
- 73. Epigenetic age-predictor for mice based on three CpG sites [https://pmc.ncbi.nlm.nih.gov/articles/PMC6156076/]
- 74. EnsembleAge: enhancing epigenetic age assessment with ... [https://link.springer.com/article/10.1007/s11357-025-01808-1]
- 75. Evaluation of Epigenetic Age Based on DNA Methylation ... [https://www.frontiersin.org/journals/genetics/articles/10.3389/fgene.2021.772298/full]